S. Kevin Zhou
Transforming medical imaging with Transformers? A comparative review of key properties, current progresses, and future perspectives
Jun 03, 2022
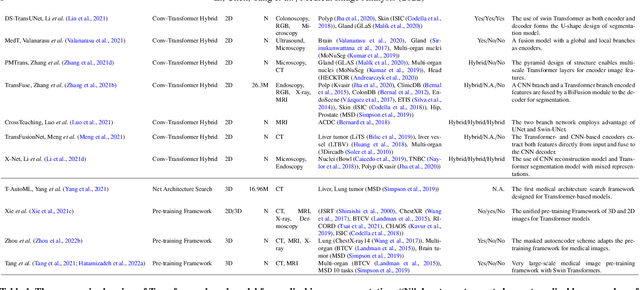
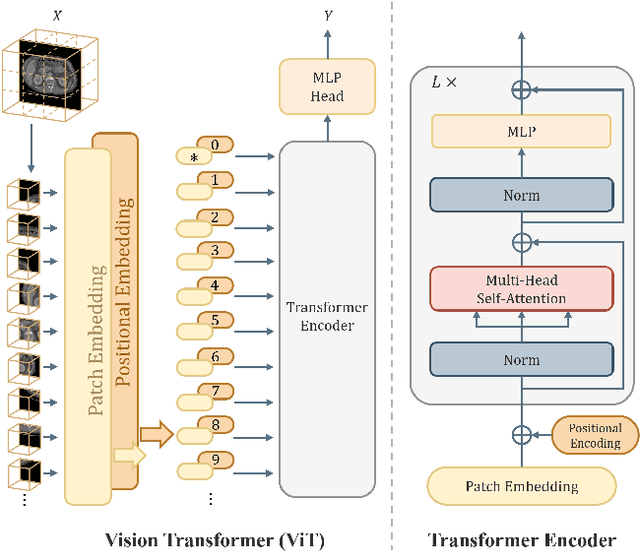
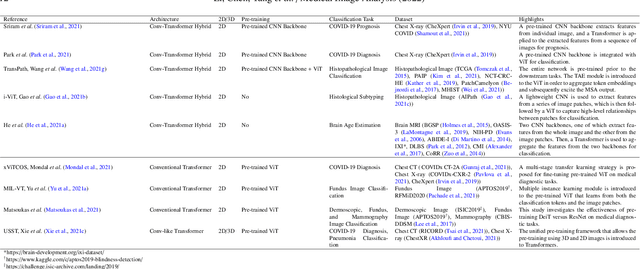
Abstract:Transformer, the latest technological advance of deep learning, has gained prevalence in natural language processing or computer vision. Since medical imaging bear some resemblance to computer vision, it is natural to inquire about the status quo of Transformers in medical imaging and ask the question: can the Transformer models transform medical imaging? In this paper, we attempt to make a response to the inquiry. After a brief introduction of the fundamentals of Transformers, especially in comparison with convolutional neural networks (CNNs), and highlighting key defining properties that characterize the Transformers, we offer a comprehensive review of the state-of-the-art Transformer-based approaches for medical imaging and exhibit current research progresses made in the areas of medical image segmentation, recognition, detection, registration, reconstruction, enhancement, etc. In particular, what distinguishes our review lies in its organization based on the Transformer's key defining properties, which are mostly derived from comparing the Transformer and CNN, and its type of architecture, which specifies the manner in which the Transformer and CNN are combined, all helping the readers to best understand the rationale behind the reviewed approaches. We conclude with discussions of future perspectives.
DFTR: Depth-supervised Fusion Transformer for Salient Object Detection
Apr 11, 2022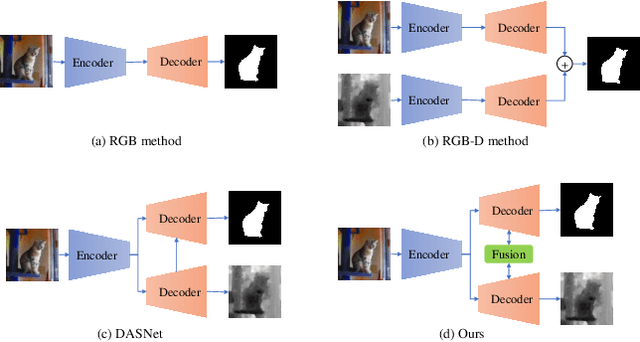
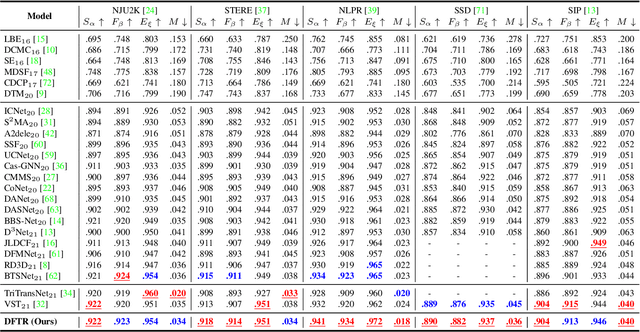
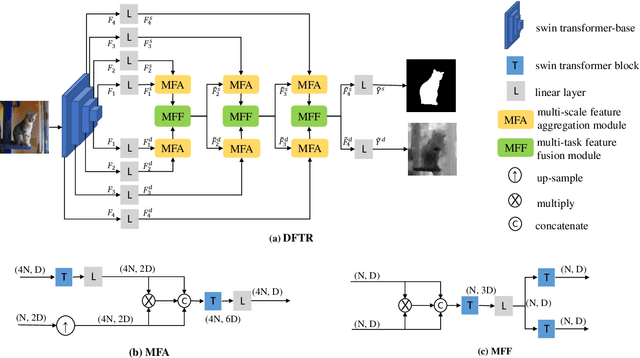
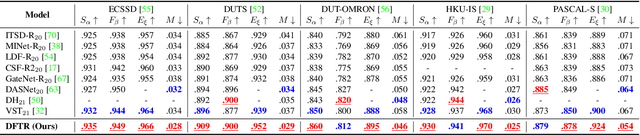
Abstract:Automated salient object detection (SOD) plays an increasingly crucial role in many computer vision applications. By reformulating the depth information as supervision rather than as input, depth-supervised convolutional neural networks (CNN) have achieved promising results on both RGB and RGB-D SOD scenarios with the merits of no requirements for extra depth networks and depth inputs in the inference stage. This paper, for the first time, seeks to expand the applicability of depth supervision to the Transformer architecture. Specifically, we develop a Depth-supervised Fusion TRansformer (DFTR), to further improve the accuracy of both RGB and RGB-D SOD. The proposed DFTR involves three primary features: 1) DFTR, to the best of our knowledge, is the first pure Transformer-based model for depth-supervised SOD; 2) A multi-scale feature aggregation (MFA) module is proposed to fully exploit the multi-scale features encoded by the Swin Transformer in a coarse-to-fine manner; 3) To enable bidirectional information flow across different streams of features, a novel multi-stage feature fusion (MFF) module is further integrated into our DFTR with the emphasis on salient regions at different network learning stages. We extensively evaluate the proposed DFTR on ten benchmarking datasets. Experimental results show that our DFTR consistently outperforms the existing state-of-the-art methods for both RGB and RGB-D SOD tasks. The code and model will be made publicly available.
SATr: Slice Attention with Transformer for Universal Lesion Detection
Mar 13, 2022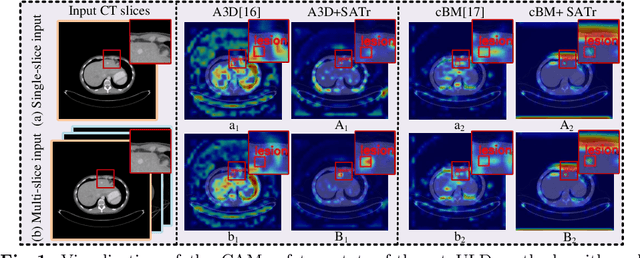
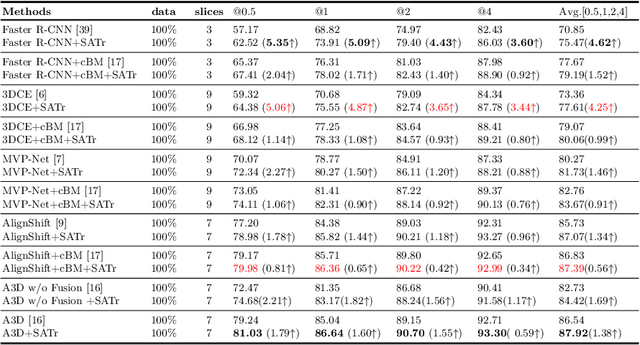
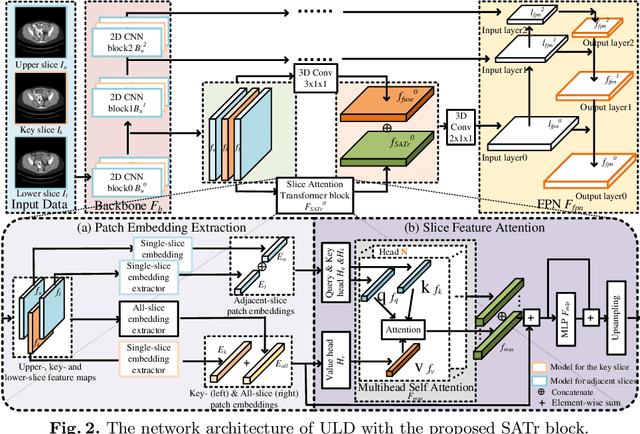
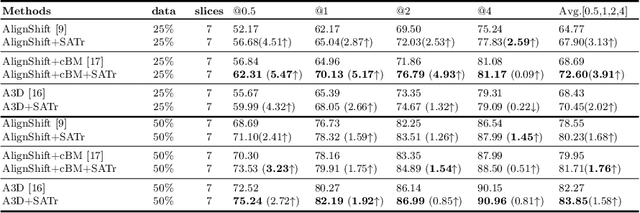
Abstract:Universal Lesion Detection (ULD) in computed tomography plays an essential role in computer-aided diagnosis. Promising ULD results have been reported by multi-slice-input detection approaches which model 3D context from multiple adjacent CT slices, but such methods still experience difficulty in obtaining a global representation among different slices and within each individual slice since they only use convolution-based fusion operations. In this paper, we propose a novel Slice Attention Transformer (SATr) block which can be easily plugged into convolution-based ULD backbones to form hybrid network structures. Such newly formed hybrid backbones can better model long-distance feature dependency via the cascaded self-attention modules in the Transformer block while still holding a strong power of modeling local features with the convolutional operations in the original backbone. Experiments with five state-of-the-art methods show that the proposed SATr block can provide an almost free boost to lesion detection accuracy without extra hyperparameters or special network designs.
DATR: Domain-adaptive transformer for multi-domain landmark detection
Mar 12, 2022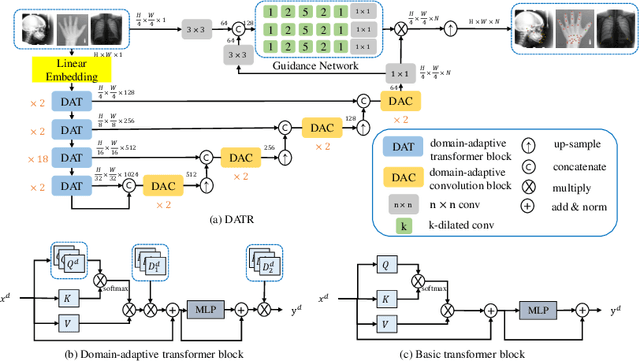
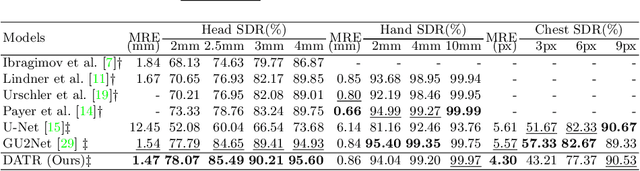
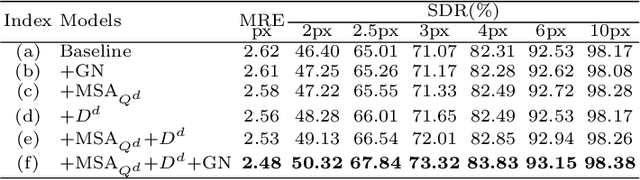
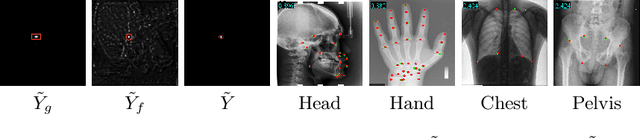
Abstract:Accurate anatomical landmark detection plays an increasingly vital role in medical image analysis. Although existing methods achieve satisfying performance, they are mostly based on CNN and specialized for a single domain say associated with a particular anatomical region. In this work, we propose a universal model for multi-domain landmark detection by taking advantage of transformer for modeling long dependencies and develop a domain-adaptive transformer model, named as DATR, which is trained on multiple mixed datasets from different anatomies and capable of detecting landmarks of any image from those anatomies. The proposed DATR exhibits three primary features: (i) It is the first universal model which introduces transformer as an encoder for multi-anatomy landmark detection; (ii) We design a domain-adaptive transformer for anatomy-aware landmark detection, which can be effectively extended to any other transformer network; (iii) Following previous studies, we employ a light-weighted guidance network, which encourages the transformer network to detect more accurate landmarks. We carry out experiments on three widely used X-ray datasets for landmark detection, which have 1,588 images and 62 landmarks in total, including three different anatomies (head, hand, and chest). Experimental results demonstrate that our proposed DATR achieves state-of-the-art performances by most metrics and behaves much better than any previous convolution-based models. The code will be released publicly.
Towards performant and reliable undersampled MR reconstruction via diffusion model sampling
Mar 11, 2022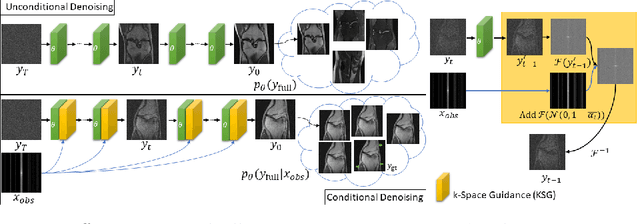
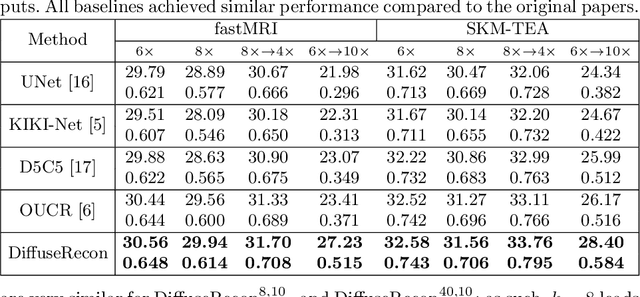
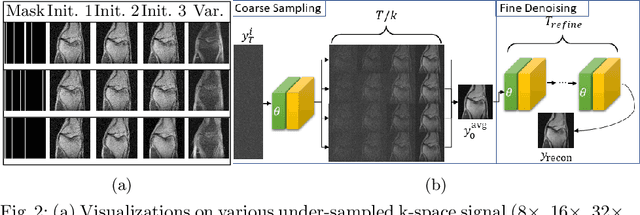
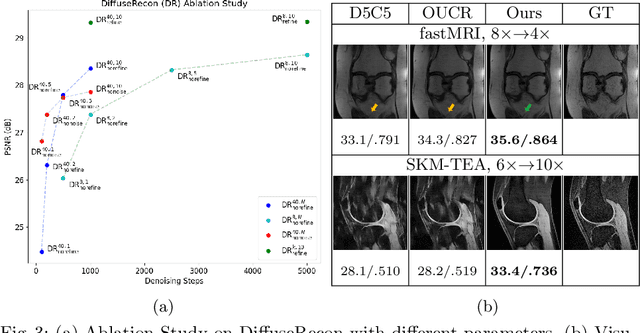
Abstract:Magnetic Resonance (MR) image reconstruction from under-sampled acquisition promises faster scanning time. To this end, current State-of-The-Art (SoTA) approaches leverage deep neural networks and supervised training to learn a recovery model. While these approaches achieve impressive performances, the learned model can be fragile on unseen degradation, e.g. when given a different acceleration factor. These methods are also generally deterministic and provide a single solution to an ill-posed problem; as such, it can be difficult for practitioners to understand the reliability of the reconstruction. We introduce DiffuseRecon, a novel diffusion model-based MR reconstruction method. DiffuseRecon guides the generation process based on the observed signals and a pre-trained diffusion model, and does not require additional training on specific acceleration factors. DiffuseRecon is stochastic in nature and generates results from a distribution of fully-sampled MR images; as such, it allows us to explicitly visualize different potential reconstruction solutions. Lastly, DiffuseRecon proposes an accelerated, coarse-to-fine Monte-Carlo sampling scheme to approximate the most likely reconstruction candidate. The proposed DiffuseRecon achieves SoTA performances reconstructing from raw acquisition signals in fastMRI and SKM-TEA. Code will be open-sourced at www.github.com/cpeng93/DiffuseRecon.
Recovering medical images from CT film photos
Mar 10, 2022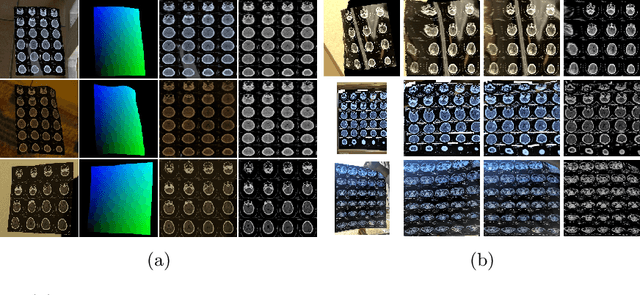
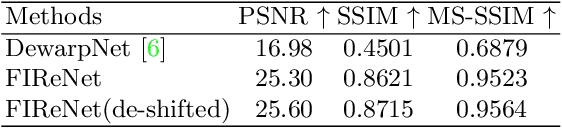
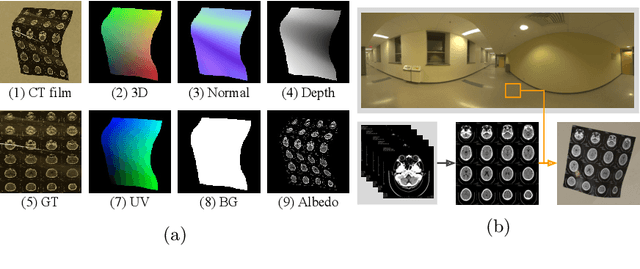


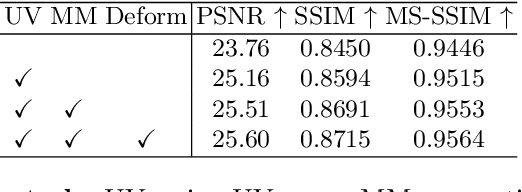
Abstract:While medical images such as computed tomography (CT) are stored in DICOM format in hospital PACS, it is still quite routine in many countries to print a film as a transferable medium for the purposes of self-storage and secondary consultation. Also, with the ubiquitousness of mobile phone cameras, it is quite common to take pictures of CT films, which unfortunately suffer from geometric deformation and illumination variation. In this work, we study the problem of recovering a CT film, which marks \textbf{the first attempt} in the literature, to the best of our knowledge. We start with building a large-scale head CT film database CTFilm20K, consisting of approximately 20,000 pictures, using the widely used computer graphics software Blender. We also record all accompanying information related to the geometric deformation (such as 3D coordinate, depth, normal, and UV maps) and illumination variation (such as albedo map). Then we propose a deep framework called \textbf{F}ilm \textbf{I}mage \textbf{Re}covery \textbf{Net}work (\textbf{FIReNet}) to tackle geometric deformation and illumination variation using the multiple maps extracted from the CT films to collaboratively guide the recovery process. Finally, we convert the dewarped images to DICOM files with our cascade model for further analysis such as radiomics feature extraction. Extensive experiments demonstrate the superiority of our approach over the previous approaches. We plan to open source the simulated images and deep models for promoting the research on CT film image analysis.
Undersampled MRI Reconstruction with Side Information-Guided Normalisation
Mar 07, 2022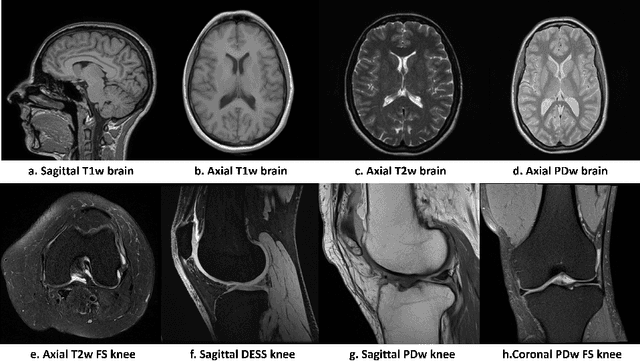
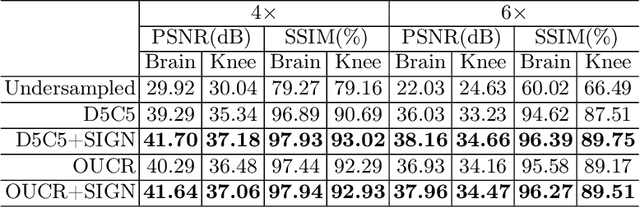
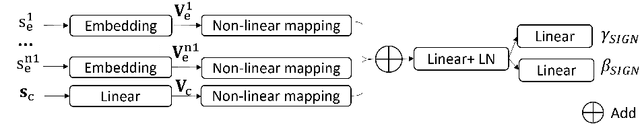
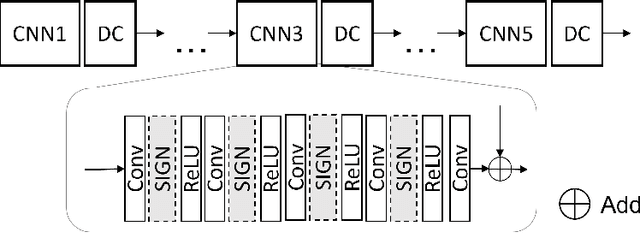
Abstract:Magnetic resonance (MR) images exhibit various contrasts and appearances based on factors such as different acquisition protocols, views, manufacturers, scanning parameters, etc. This generally accessible appearance-related side information affects deep learning-based undersampled magnetic resonance imaging (MRI) reconstruction frameworks, but has been overlooked in the majority of current works. In this paper, we investigate the use of such side information as normalisation parameters in a convolutional neural network (CNN) to improve undersampled MRI reconstruction. Specifically, a Side Information-Guided Normalisation (SIGN) module, containing only few layers, is proposed to efficiently encode the side information and output the normalisation parameters. We examine the effectiveness of such a module on two popular reconstruction architectures, D5C5 and OUCR. The experimental results on both brain and knee images under various acceleration rates demonstrate that the proposed method improves on its corresponding baseline architectures with a significant margin.
Rib Suppression in Digital Chest Tomosynthesis
Mar 05, 2022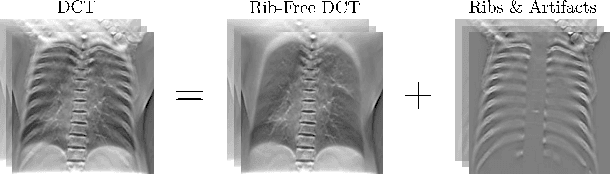
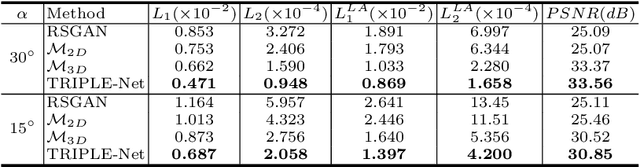
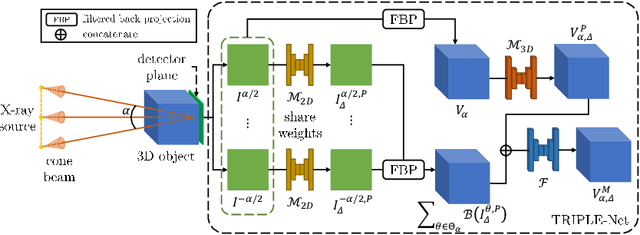
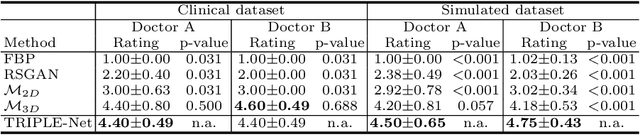
Abstract:Digital chest tomosynthesis (DCT) is a technique to produce sectional 3D images of a human chest for pulmonary disease screening, with 2D X-ray projections taken within an extremely limited range of angles. However, under the limited angle scenario, DCT contains strong artifacts caused by the presence of ribs, jamming the imaging quality of the lung area. Recently, great progress has been achieved for rib suppression in a single X-ray image, to reveal a clearer lung texture. We firstly extend the rib suppression problem to the 3D case at the software level. We propose a $\textbf{T}$omosynthesis $\textbf{RI}$b Su$\textbf{P}$pression and $\textbf{L}$ung $\textbf{E}$nhancement $\textbf{Net}$work (TRIPLE-Net) to model the 3D rib component and provide a rib-free DCT. TRIPLE-Net takes the advantages from both 2D and 3D domains, which model the ribs in DCT with the exact FBP procedure and 3D depth information, respectively. The experiments on simulated datasets and clinical data have shown the effectiveness of TRIPLE-Net to preserve lung details as well as improve the imaging quality of pulmonary diseases. Finally, an expert user study confirms our findings.
Relative distance matters for one-shot landmark detection
Mar 04, 2022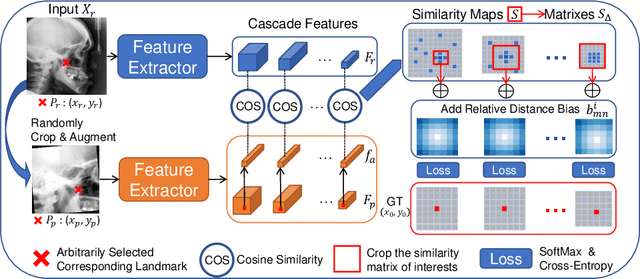
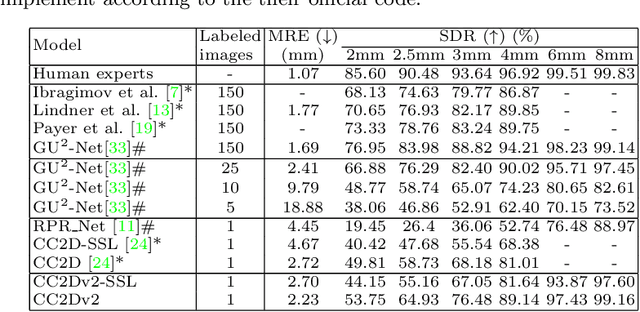
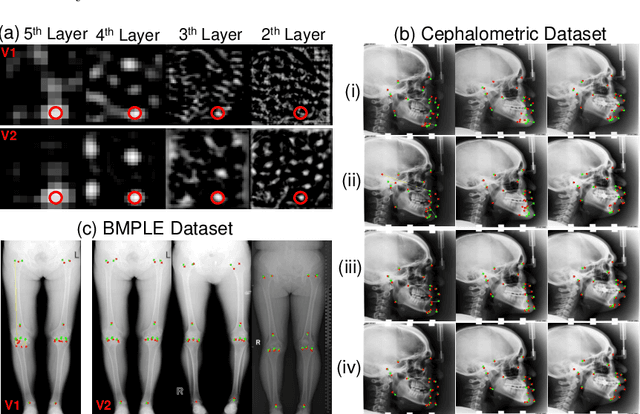
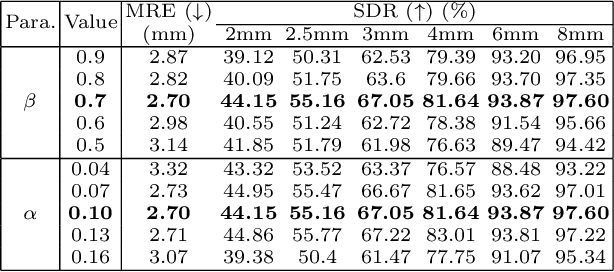
Abstract:Contrastive learning based methods such as cascade comparing to detect (CC2D) have shown great potential for one-shot medical landmark detection. However, the important cue of relative distance between landmarks is ignored in CC2D. In this paper, we upgrade CC2D to version II by incorporating a simple-yet-effective relative distance bias in the training stage, which is theoretically proved to encourage the encoder to project the relatively distant landmarks to the embeddings with low similarities. As consequence, CC2Dv2 is less possible to detect a wrong point far from the correct landmark. Furthermore, we present an open-source, landmark-labeled dataset for the measurement of biomechanical parameters of the lower extremity to alleviate the burden of orthopedic surgeons. The effectiveness of CC2Dv2 is evaluated on the public dataset from the ISBI 2015 Grand-Challenge of cephalometric radiographs and our new dataset, which greatly outperforms the state-of-the-art one-shot landmark detection approaches.
MixCL: Pixel label matters to contrastive learning
Mar 04, 2022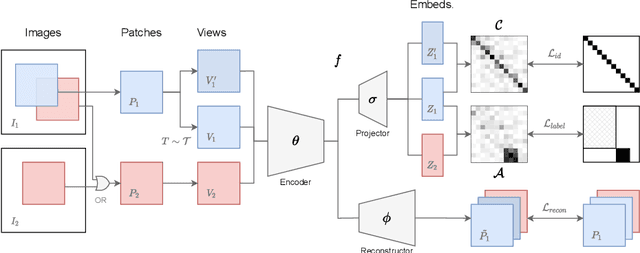
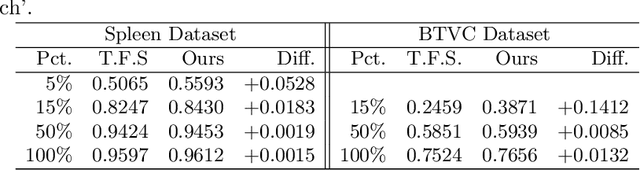
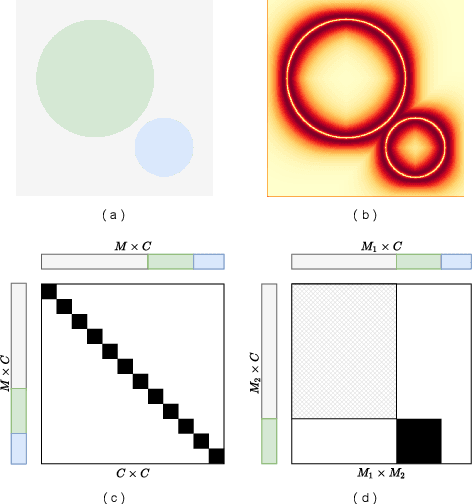
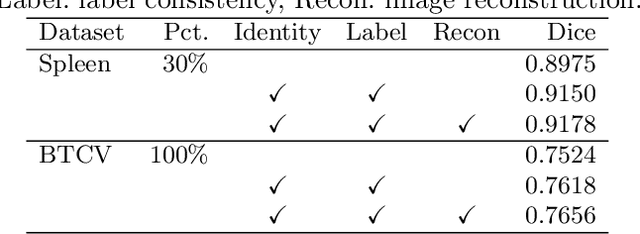
Abstract:Contrastive learning and self-supervised techniques have gained prevalence in computer vision for the past few years. It is essential for medical image analysis, which is often notorious for its lack of annotations. Most existing self-supervised methods applied in natural imaging tasks focus on designing proxy tasks for unlabeled data. For example, contrastive learning is often based on the fact that an image and its transformed version share the same identity. However, pixel annotations contain much valuable information for medical image segmentation, which is largely ignored in contrastive learning. In this work, we propose a novel pre-training framework called Mixed Contrastive Learning (MixCL) that leverages both image identities and pixel labels for better modeling by maintaining identity consistency, label consistency, and reconstruction consistency together. Consequently, thus pre-trained model has more robust representations that characterize medical images. Extensive experiments demonstrate the effectiveness of the proposed method, improving the baseline by 5.28% and 14.12% in Dice coefficient when 5% labeled data of Spleen and 15% of BTVC are used in fine-tuning, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge