Rajiv Raman
Adaptive Class Learning to Screen Diabetic Disorders in Fundus Images of Eye
Jan 21, 2025Abstract:The prevalence of ocular illnesses is growing globally, presenting a substantial public health challenge. Early detection and timely intervention are crucial for averting visual impairment and enhancing patient prognosis. This research introduces a new framework called Class Extension with Limited Data (CELD) to train a classifier to categorize retinal fundus images. The classifier is initially trained to identify relevant features concerning Healthy and Diabetic Retinopathy (DR) classes and later fine-tuned to adapt to the task of classifying the input images into three classes: Healthy, DR, and Glaucoma. This strategy allows the model to gradually enhance its classification capabilities, which is beneficial in situations where there are only a limited number of labeled datasets available. Perturbation methods are also used to identify the input image characteristics responsible for influencing the models decision-making process. We achieve an overall accuracy of 91% on publicly available datasets.
The Persistent Robot Charging Problem for Long-Duration Autonomy
Sep 01, 2024



Abstract:This paper introduces a novel formulation aimed at determining the optimal schedule for recharging a fleet of $n$ heterogeneous robots, with the primary objective of minimizing resource utilization. This study provides a foundational framework applicable to Multi-Robot Mission Planning, particularly in scenarios demanding Long-Duration Autonomy (LDA) or other contexts that necessitate periodic recharging of multiple robots. A novel Integer Linear Programming (ILP) model is proposed to calculate the optimal initial conditions (partial charge) for individual robots, leading to the minimal utilization of charging stations. This formulation was further generalized to maximize the servicing time for robots given adequate charging stations. The efficacy of the proposed formulation is evaluated through a comparative analysis, measuring its performance against the thrift price scheduling algorithm documented in the existing literature. The findings not only validate the effectiveness of the proposed approach but also underscore its potential as a valuable tool in optimizing resource allocation for a range of robotic and engineering applications.
Underspecification Presents Challenges for Credibility in Modern Machine Learning
Nov 06, 2020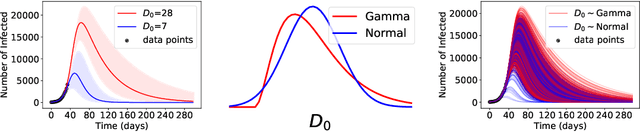

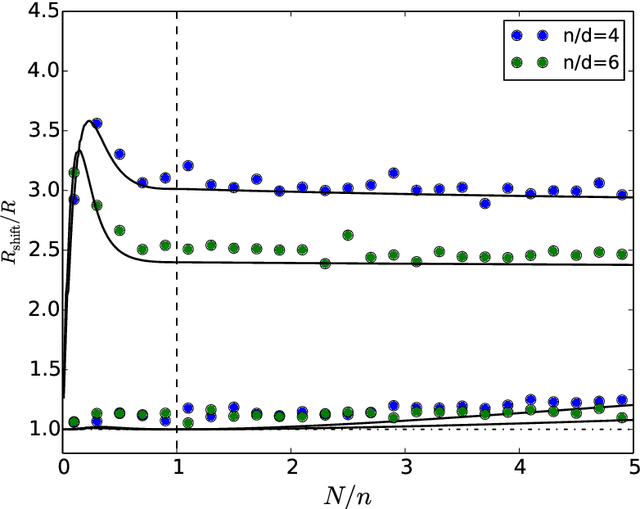

Abstract:ML models often exhibit unexpectedly poor behavior when they are deployed in real-world domains. We identify underspecification as a key reason for these failures. An ML pipeline is underspecified when it can return many predictors with equivalently strong held-out performance in the training domain. Underspecification is common in modern ML pipelines, such as those based on deep learning. Predictors returned by underspecified pipelines are often treated as equivalent based on their training domain performance, but we show here that such predictors can behave very differently in deployment domains. This ambiguity can lead to instability and poor model behavior in practice, and is a distinct failure mode from previously identified issues arising from structural mismatch between training and deployment domains. We show that this problem appears in a wide variety of practical ML pipelines, using examples from computer vision, medical imaging, natural language processing, clinical risk prediction based on electronic health records, and medical genomics. Our results show the need to explicitly account for underspecification in modeling pipelines that are intended for real-world deployment in any domain.
Quantitative and Qualitative Evaluation of Explainable Deep Learning Methods for Ophthalmic Diagnosis
Sep 26, 2020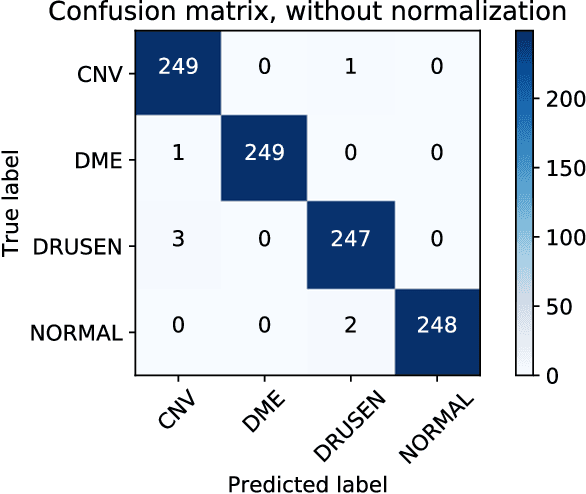
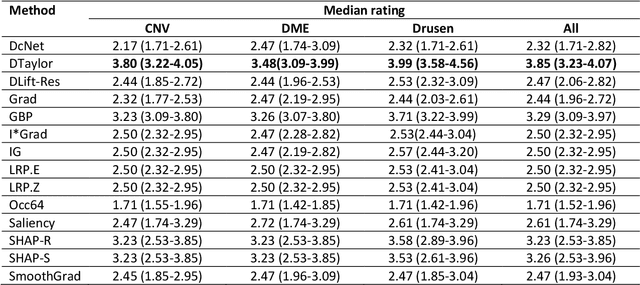
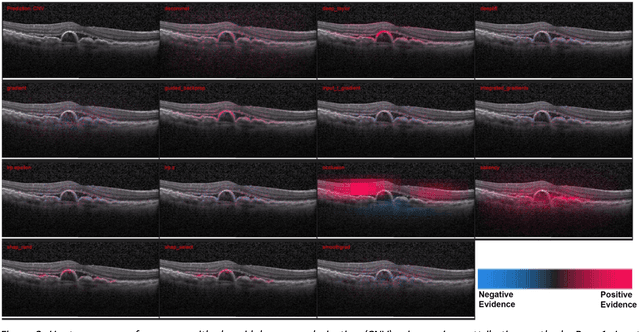
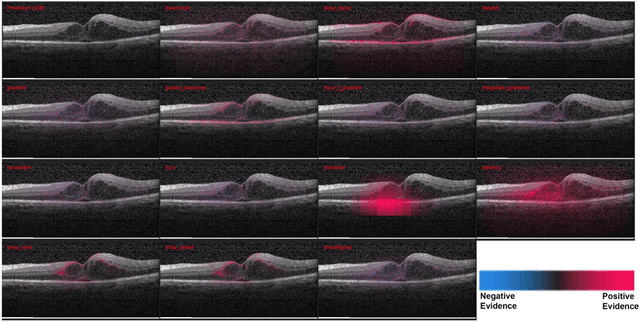
Abstract:Background: The lack of explanations for the decisions made by algorithms such as deep learning has hampered their acceptance by the clinical community despite highly accurate results on multiple problems. Recently, attribution methods have emerged for explaining deep learning models, and they have been tested on medical imaging problems. The performance of attribution methods is compared on standard machine learning datasets and not on medical images. In this study, we perform a comparative analysis to determine the most suitable explainability method for retinal OCT diagnosis. Methods: A commonly used deep learning model known as Inception v3 was trained to diagnose 3 retinal diseases - choroidal neovascularization (CNV), diabetic macular edema (DME), and drusen. The explanations from 13 different attribution methods were rated by a panel of 14 clinicians for clinical significance. Feedback was obtained from the clinicians regarding the current and future scope of such methods. Results: An attribution method based on a Taylor series expansion, called Deep Taylor was rated the highest by clinicians with a median rating of 3.85/5. It was followed by two other attribution methods, Guided backpropagation and SHAP (SHapley Additive exPlanations). Conclusion: Explanations of deep learning models can make them more transparent for clinical diagnosis. This study compared different explanations methods in the context of retinal OCT diagnosis and found that the best performing method may not be the one considered best for other deep learning tasks. Overall, there was a high degree of acceptance from the clinicians surveyed in the study. Keywords: explainable AI, deep learning, machine learning, image processing, Optical coherence tomography, retina, Diabetic macular edema, Choroidal Neovascularization, Drusen
Deep Learning vs. Human Graders for Classifying Severity Levels of Diabetic Retinopathy in a Real-World Nationwide Screening Program
Oct 18, 2018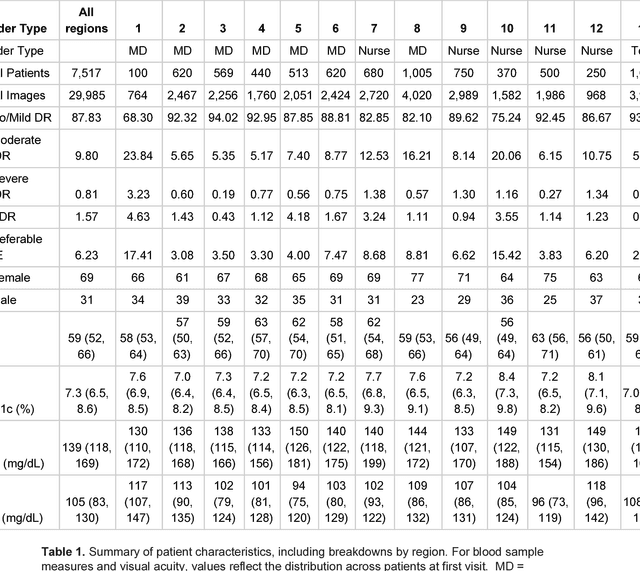
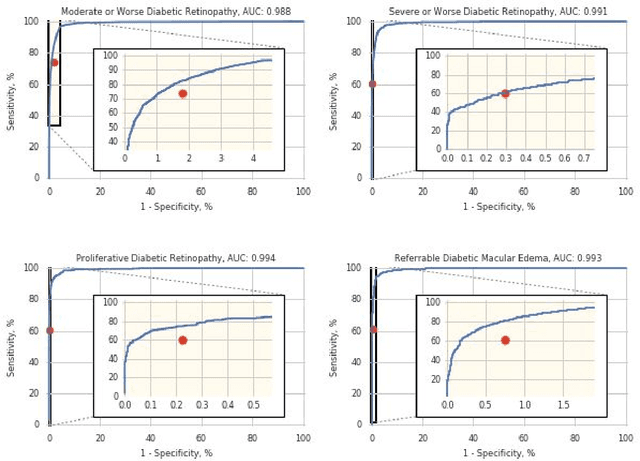
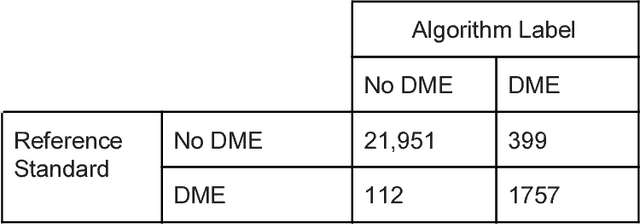
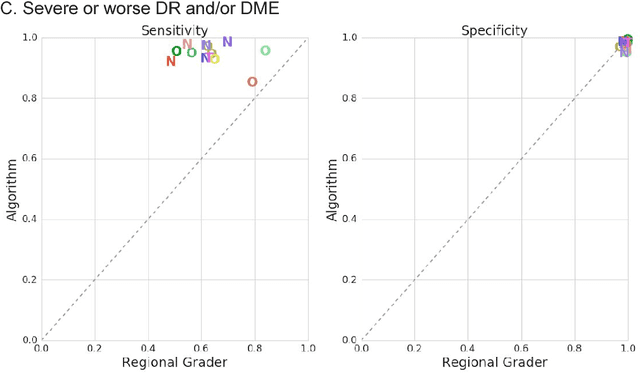
Abstract:Deep learning algorithms have been used to detect diabetic retinopathy (DR) with specialist-level accuracy. This study aims to validate one such algorithm on a large-scale clinical population, and compare the algorithm performance with that of human graders. 25,326 gradable retinal images of patients with diabetes from the community-based, nation-wide screening program of DR in Thailand were analyzed for DR severity and referable diabetic macular edema (DME). Grades adjudicated by a panel of international retinal specialists served as the reference standard. Across different severity levels of DR for determining referable disease, deep learning significantly reduced the false negative rate (by 23%) at the cost of slightly higher false positive rates (2%). Deep learning algorithms may serve as a valuable tool for DR screening.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge