Qinmu Peng
BGM: Building a Dynamic Guidance Map without Visual Images for Trajectory Prediction
Oct 08, 2020
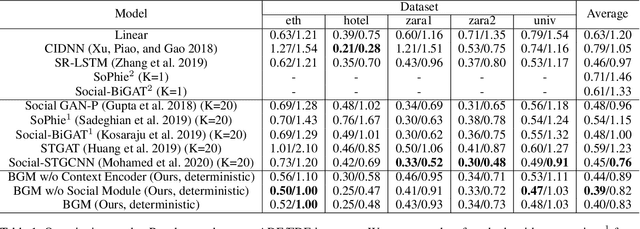
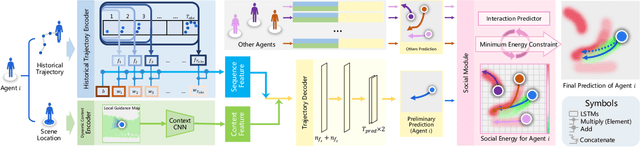
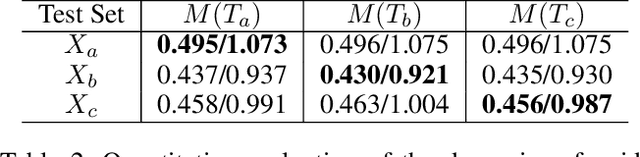
Abstract:Visual images usually contain the informative context of the environment, thereby helping to predict agents' behaviors. However, they hardly impose the dynamic effects on agents' actual behaviors due to the respectively fixed semantics. To solve this problem, we propose a deterministic model named BGM to construct a guidance map to represent the dynamic semantics, which circumvents to use visual images for each agent to reflect the difference of activities in different periods. We first record all agents' activities in the scene within a period close to the current to construct a guidance map and then feed it to a Context CNN to obtain their context features. We adopt a Historical Trajectory Encoder to extract the trajectory features and then combine them with the context feature as the input of the social energy based trajectory decoder, thus obtaining the prediction that meets the social rules. Experiments demonstrate that BGM achieves state-of-the-art prediction accuracy on the two widely used ETH and UCY datasets and handles more complex scenarios.
Modal Regression based Structured Low-rank Matrix Recovery for Multi-view Learning
Mar 22, 2020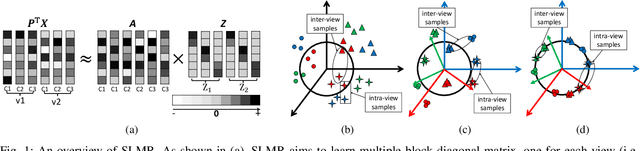
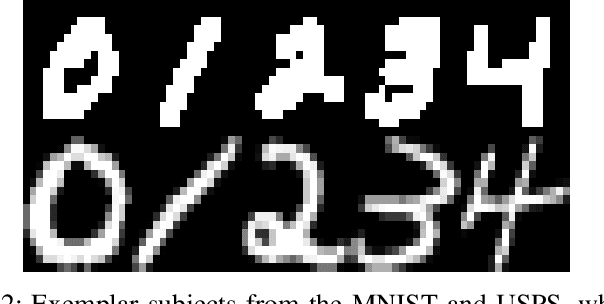
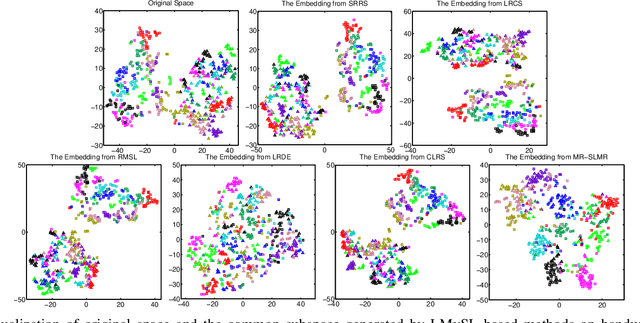

Abstract:Low-rank Multi-view Subspace Learning (LMvSL) has shown great potential in cross-view classification in recent years. Despite their empirical success, existing LMvSL based methods are incapable of well handling view discrepancy and discriminancy simultaneously, which thus leads to the performance degradation when there is a large discrepancy among multi-view data. To circumvent this drawback, motivated by the block-diagonal representation learning, we propose Structured Low-rank Matrix Recovery (SLMR), a unique method of effectively removing view discrepancy and improving discriminancy through the recovery of structured low-rank matrix. Furthermore, recent low-rank modeling provides a satisfactory solution to address data contaminated by predefined assumptions of noise distribution, such as Gaussian or Laplacian distribution. However, these models are not practical since complicated noise in practice may violate those assumptions and the distribution is generally unknown in advance. To alleviate such limitation, modal regression is elegantly incorporated into the framework of SLMR (term it MR-SLMR). Different from previous LMvSL based methods, our MR-SLMR can handle any zero-mode noise variable that contains a wide range of noise, such as Gaussian noise, random noise and outliers. The alternating direction method of multipliers (ADMM) framework and half-quadratic theory are used to efficiently optimize MR-SLMR. Experimental results on four public databases demonstrate the superiority of MR-SLMR and its robustness to complicated noise.
A Spatial-Temporal Attentive Network with Spatial Continuity for Trajectory Prediction
Mar 16, 2020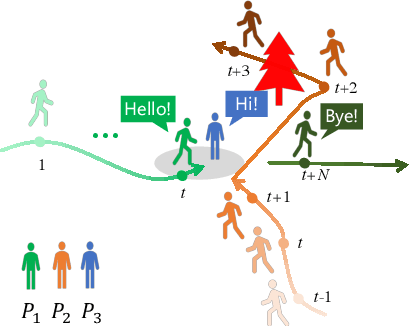
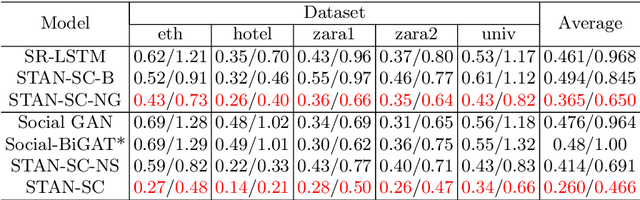
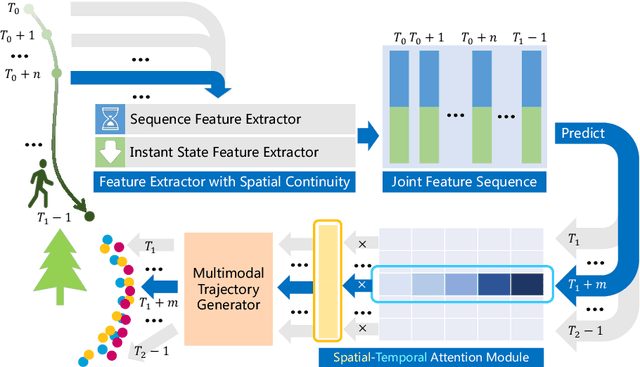
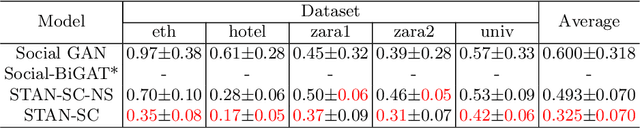
Abstract:It remains challenging to automatically predict the multi-agent trajectory due to multiple interactions including agent to agent interaction and scene to agent interaction. Although recent methods have achieved promising performance, most of them just consider spatial influence of the interactions and ignore the fact that temporal influence always accompanies spatial influence. Moreover, those methods based on scene information always require extra segmented scene images to generate multiple socially acceptable trajectories. To solve these limitations, we propose a novel model named spatial-temporal attentive network with spatial continuity (STAN-SC). First, spatial-temporal attention mechanism is presented to explore the most useful and important information. Second, we conduct a joint feature sequence based on the sequence and instant state information to make the generative trajectories keep spatial continuity. Experiments are performed on the two widely used ETH-UCY datasets and demonstrate that the proposed model achieves state-of-the-art prediction accuracy and handles more complex scenarios.
Similarity-DT: Kernel Similarity Embedding for Dynamic Texture Synthesis
Dec 11, 2019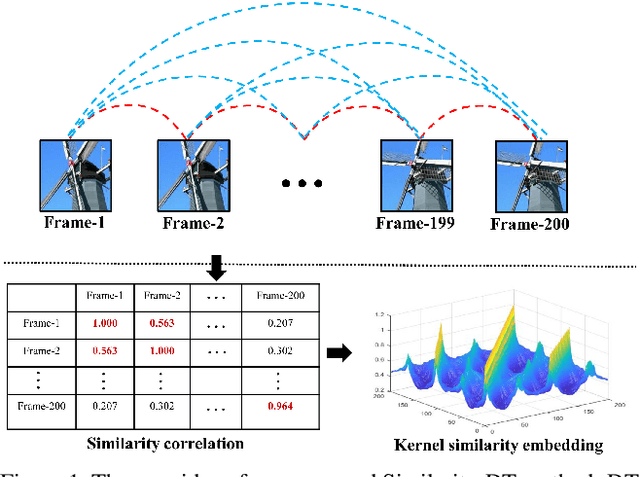
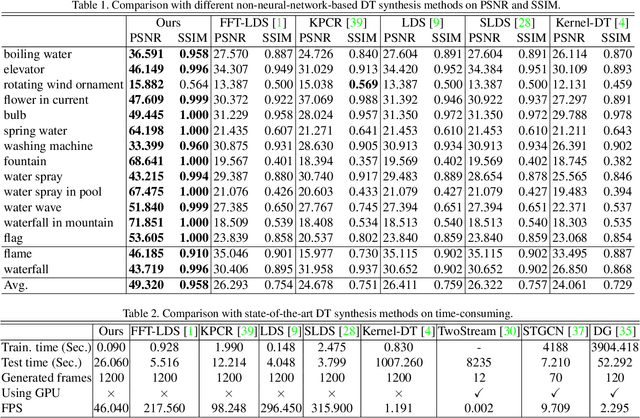
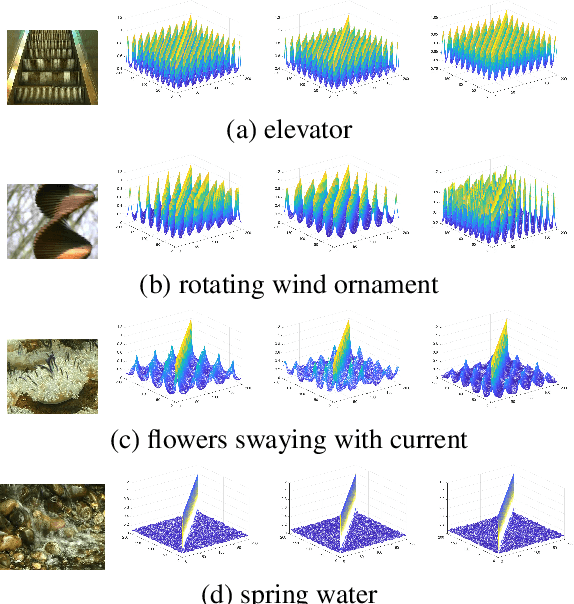
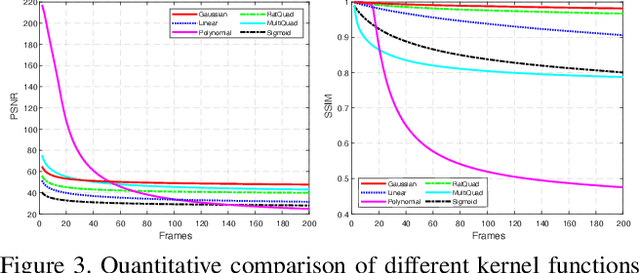
Abstract:Dynamic texture (DT) exhibits statistical stationarity in the spatial domain and stochastic repetitiveness in the temporal dimension, indicating that different frames of DT possess high similarity correlation. However, there are no DT synthesis methods to consider the similarity prior for representing DT instead, which can explicitly capture the homogeneous and heterogeneous correlation between different frames of DT. In this paper, we propose a novel DT synthesis method (named Similarity-DT), which embeds the similarity prior into the representation of DT. Specifically, we first raise two hypotheses: the content of texture video frames varies over time-to-time, while the more closed frames should be more similar; the transition between frame-to-frame could be modeled as a linear or nonlinear function to capture the similarity correlation. Then, our proposed Similarity-DT integrates kernel learning and extreme learning machine (ELM) into a powerful unified synthesis model to learn kernel similarity embedding to represent the spatial-temporal transition among frame-to-frame of DTs. Extensive experiments on DT videos collected from internet and two benchmark datasets, i.e., Gatech Graphcut Textures and Dyntex, demonstrate that the learned kernel similarity embedding effectively exhibits the discriminative representation for DTs. Hence our method is capable of preserving long-term temporal continuity of the synthesized DT sequences with excellent sustainability and generalization. We also show that our method effectively generates realistic DT videos with fast speed and low computation, compared with the state-of-the-art approaches.
Closed-Loop Adaptation for Weakly-Supervised Semantic Segmentation
May 29, 2019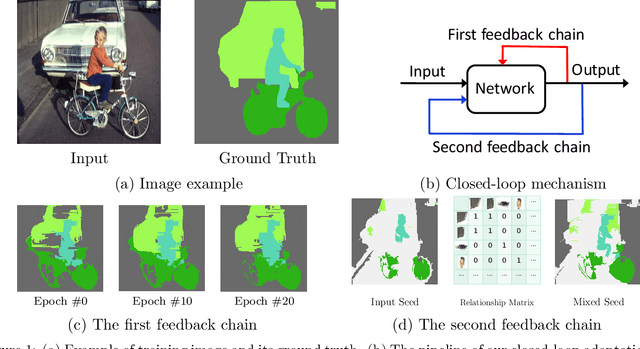
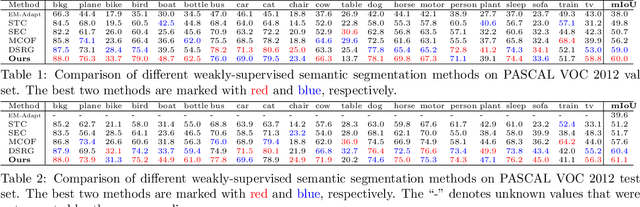
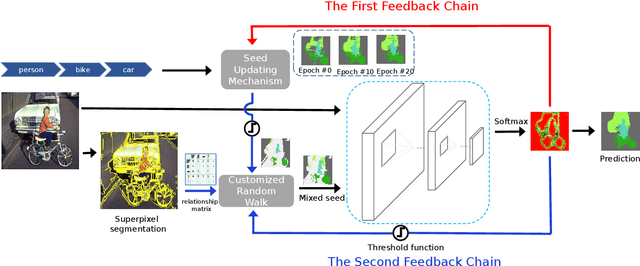

Abstract:Weakly-supervised semantic segmentation aims to assign each pixel a semantic category under weak supervisions, such as image-level tags. Most of existing weakly-supervised semantic segmentation methods do not use any feedback from segmentation output and can be considered as open-loop systems. They are prone to accumulated errors because of the static seeds and the sensitive structure information. In this paper, we propose a generic self-adaptation mechanism for existing weakly-supervised semantic segmentation methods by introducing two feedback chains, thus constituting a closed-loop system. Specifically, the first chain iteratively produces dynamic seeds by incorporating cross-image structure information, whereas the second chain further expands seed regions by a customized random walk process to reconcile inner-image structure information characterized by superpixels. Experiments on PASCAL VOC 2012 suggest that our network outperforms state-of-the-art methods with significantly less computational and memory burden.
Fast and accurate reconstruction of HARDI using a 1D encoder-decoder convolutional network
Mar 21, 2019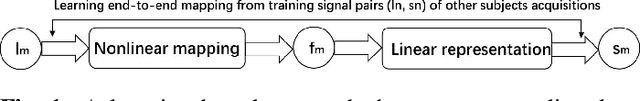
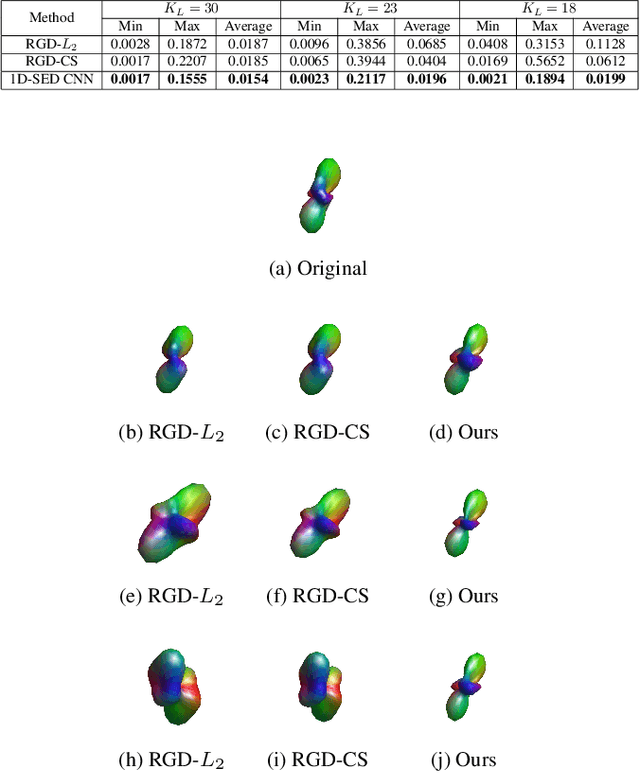
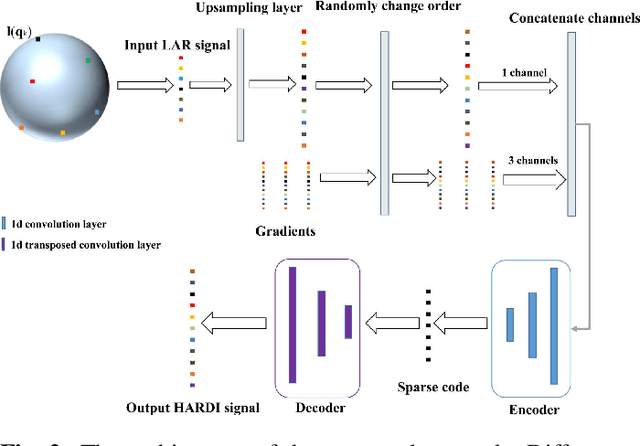
Abstract:High angular resolution diffusion imaging (HARDI) demands a lager amount of data measurements compared to diffusion tensor imaging, restricting its use in practice. In this work, we explore a learning-based approach to reconstruct HARDI from a smaller number of measurements in q-space. The approach aims to directly learn the mapping relationship between the measured and HARDI signals from the collecting HARDI acquisitions of other subjects. Specifically, the mapping is represented as a 1D encoder-decoder convolutional neural network under the guidance of the compressed sensing (CS) theory for HARDI reconstruction. The proposed network architecture mainly consists of two parts: an encoder network produces the sparse coefficients and a decoder network yields a reconstruction result. Experiment results demonstrate we can robustly reconstruct HARDI signals with the accurate results and fast speed.
Fully-automatic segmentation of kidneys in clinical ultrasound images using a boundary distance regression network
Jan 05, 2019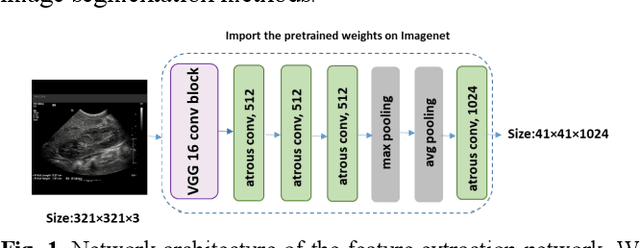
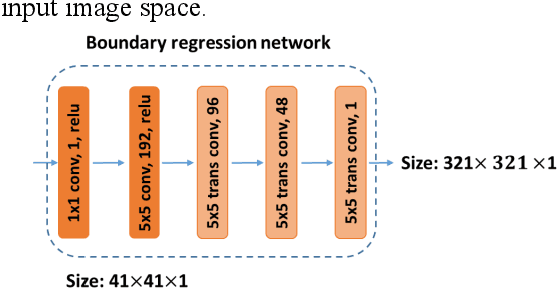
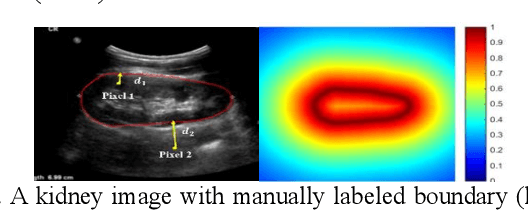
Abstract:It remains challenging to automatically segment kidneys in clinical ultrasound images due to the kidneys' varied shapes and image intensity distributions, although semi-automatic methods have achieved promising performance. In this study, we developed a novel boundary distance regression deep neural network to segment the kidneys, informed by the fact that the kidney boundaries are relatively consistent across images in terms of their appearance. Particularly, we first use deep neural networks pre-trained for classification of natural images to extract high-level image features from ultrasound images, then these feature maps are used as input to learn kidney boundary distance maps using a boundary distance regression network, and finally the predicted boundary distance maps are classified as kidney pixels or non-kidney pixels using a pixel classification network in an end-to-end learning fashion. Experimental results have demonstrated that our method could effectively improve the performance of automatic kidney segmentation, significantly better than deep learning based pixel classification networks.
Subsequent Boundary Distance Regression and Pixelwise Classification Networks for Automatic Kidney Segmentation in Ultrasound Images
Nov 12, 2018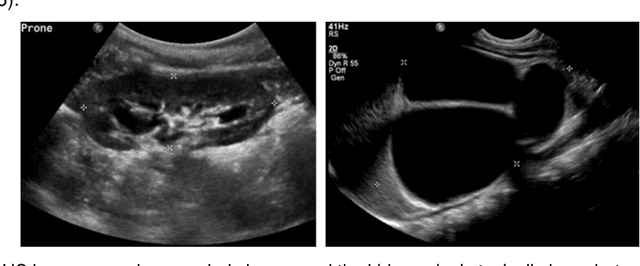
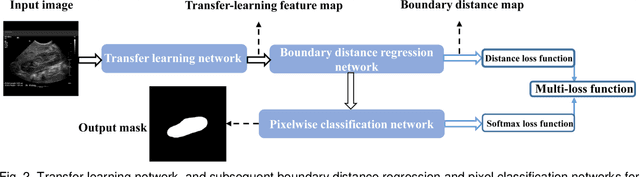
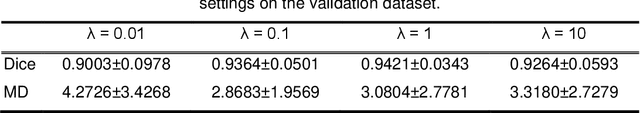
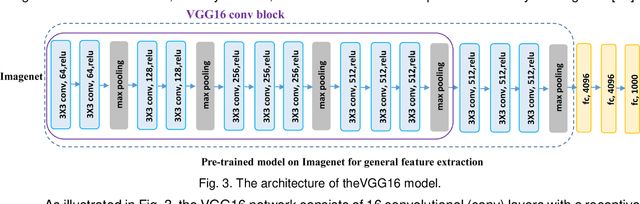
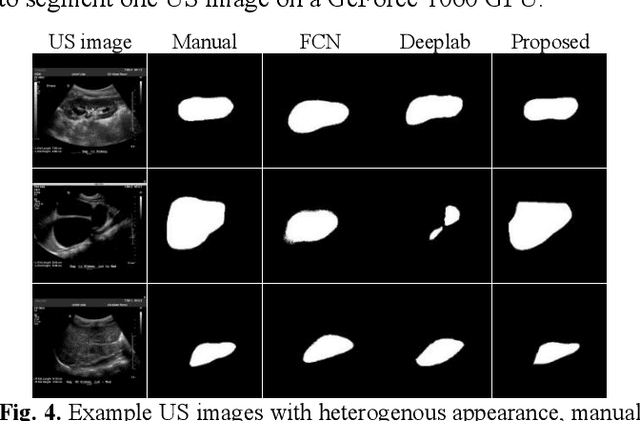
Abstract:It remains challenging to automatically segment kidneys in clinical ultrasound (US) images due to the kidneys' varied shapes and image intensity distributions, although semi-automatic methods have achieved promising performance. In this study, we propose subsequent boundary distance regression and pixel classification networks to segment the kidneys, informed by the fact that the kidney boundaries have relatively homogenous texture patterns across images. Particularly, we first use deep neural networks pre-trained for classification of natural images to extract high-level image features from US images, then these features are used as input to learn kidney boundary distance maps using a boundary distance regression network, and finally the predicted boundary distance maps are classified as kidney pixels or non-kidney pixels using a pixel classification network in an end-to-end learning fashion. We also proposed a novel data-augmentation method based on kidney shape registration to generate enriched training data from a small number of US images with manually segmented kidney labels. Experimental results have demonstrated that our method could effectively improve the performance of automatic kidney segmentation, significantly better than deep learning based pixel classification networks.
Coarse-to-Fine Salient Object Detection with Low-Rank Matrix Recovery
Oct 02, 2018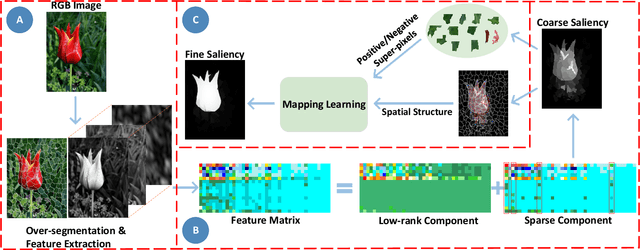
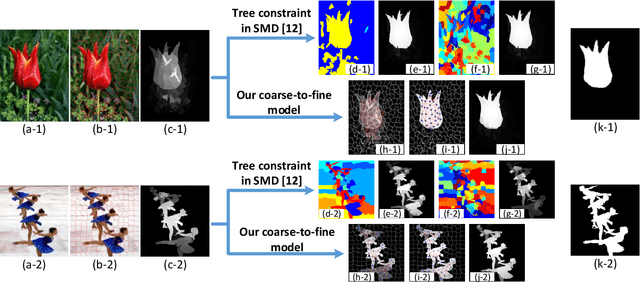
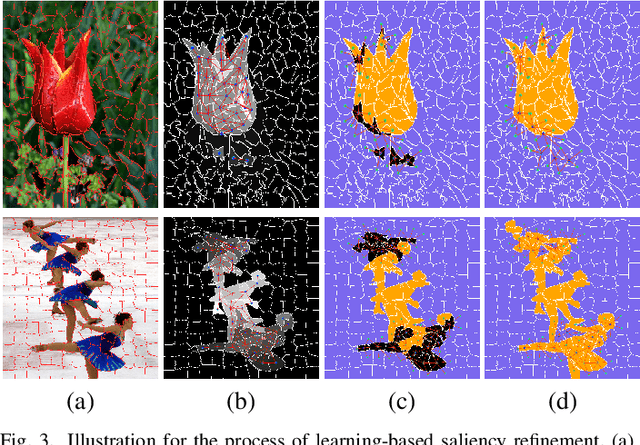
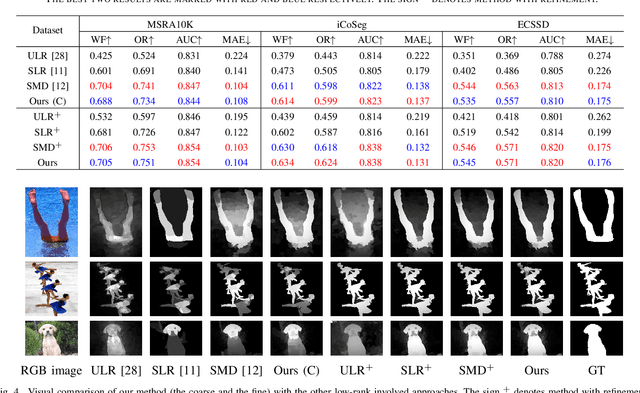
Abstract:Low-Rank Matrix Recovery (LRMR) has recently been applied to saliency detection by decomposing image features into a low-rank component associated with background and a sparse component associated with visual salient regions. Despite its great potential, existing LRMR-based saliency detection methods seldom consider the inter-relationship among elements within these two components, thus are prone to generating scattered or incomplete saliency maps. In this paper, we introduce a novel and efficient LRMR-based saliency detection model under a coarse-to-fine framework to circumvent this limitation. First, we roughly measure the saliency of image regions with a baseline LRMR model that integrates a $\ell_1$-norm sparsity constraint and a Laplacian regularization smooth term. Given samples from the coarse saliency map, we then learn a projection that maps image features to refined saliency values, to significantly sharpen the object boundaries and to preserve the object entirety. We evaluate our framework against existing LRMR based methods on three benchmark datasets. Experimental results validate the superiority of our method as well as the effectiveness of our suggested coarse-to-fine framework, especially for images containing multiple objects.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge