Nicolas Padoy
IHU Strasbourg, UNISTRA
On-the-Fly Point Annotation for Fast Medical Video Labeling
Apr 22, 2024Abstract:Purpose: In medical research, deep learning models rely on high-quality annotated data, a process often laborious and timeconsuming. This is particularly true for detection tasks where bounding box annotations are required. The need to adjust two corners makes the process inherently frame-by-frame. Given the scarcity of experts' time, efficient annotation methods suitable for clinicians are needed. Methods: We propose an on-the-fly method for live video annotation to enhance the annotation efficiency. In this approach, a continuous single-point annotation is maintained by keeping the cursor on the object in a live video, mitigating the need for tedious pausing and repetitive navigation inherent in traditional annotation methods. This novel annotation paradigm inherits the point annotation's ability to generate pseudo-labels using a point-to-box teacher model. We empirically evaluate this approach by developing a dataset and comparing on-the-fly annotation time against traditional annotation method. Results: Using our method, annotation speed was 3.2x faster than the traditional annotation technique. We achieved a mean improvement of 6.51 +- 0.98 AP@50 over conventional method at equivalent annotation budgets on the developed dataset. Conclusion: Without bells and whistles, our approach offers a significant speed-up in annotation tasks. It can be easily implemented on any annotation platform to accelerate the integration of deep learning in video-based medical research.
SelfPose3d: Self-Supervised Multi-Person Multi-View 3d Pose Estimation
Apr 02, 2024



Abstract:We present a new self-supervised approach, SelfPose3d, for estimating 3d poses of multiple persons from multiple camera views. Unlike current state-of-the-art fully-supervised methods, our approach does not require any 2d or 3d ground-truth poses and uses only the multi-view input images from a calibrated camera setup and 2d pseudo poses generated from an off-the-shelf 2d human pose estimator. We propose two self-supervised learning objectives: self-supervised person localization in 3d space and self-supervised 3d pose estimation. We achieve self-supervised 3d person localization by training the model on synthetically generated 3d points, serving as 3d person root positions, and on the projected root-heatmaps in all the views. We then model the 3d poses of all the localized persons with a bottleneck representation, map them onto all views obtaining 2d joints, and render them using 2d Gaussian heatmaps in an end-to-end differentiable manner. Afterwards, we use the corresponding 2d joints and heatmaps from the pseudo 2d poses for learning. To alleviate the intrinsic inaccuracy of the pseudo labels, we propose an adaptive supervision attention mechanism to guide the self-supervision. Our experiments and analysis on three public benchmark datasets, including Panoptic, Shelf, and Campus, show the effectiveness of our approach, which is comparable to fully-supervised methods. Code is available at \url{https://github.com/CAMMA-public/SelfPose3D}
Enhancing Gait Video Analysis in Neurodegenerative Diseases by Knowledge Augmentation in Vision Language Model
Mar 20, 2024



Abstract:We present a knowledge augmentation strategy for assessing the diagnostic groups and gait impairment from monocular gait videos. Based on a large-scale pre-trained Vision Language Model (VLM), our model learns and improves visual, textual, and numerical representations of patient gait videos, through a collective learning across three distinct modalities: gait videos, class-specific descriptions, and numerical gait parameters. Our specific contributions are two-fold: First, we adopt a knowledge-aware prompt tuning strategy to utilize the class-specific medical description in guiding the text prompt learning. Second, we integrate the paired gait parameters in the form of numerical texts to enhance the numeracy of the textual representation. Results demonstrate that our model not only significantly outperforms state-of-the-art (SOTA) in video-based classification tasks but also adeptly decodes the learned class-specific text features into natural language descriptions using the vocabulary of quantitative gait parameters. The code and the model will be made available at our project page.
Optimizing Latent Graph Representations of Surgical Scenes for Zero-Shot Domain Transfer
Mar 11, 2024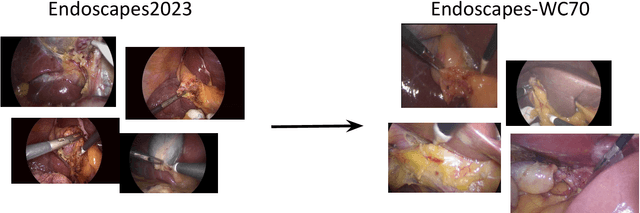
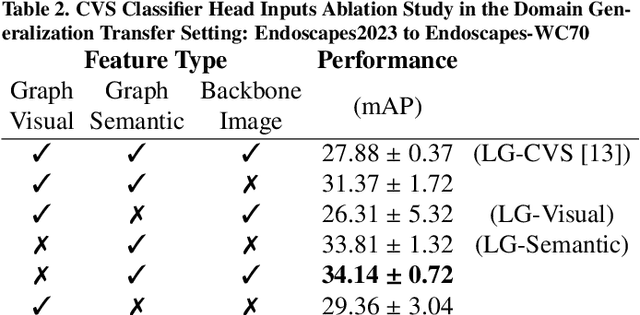
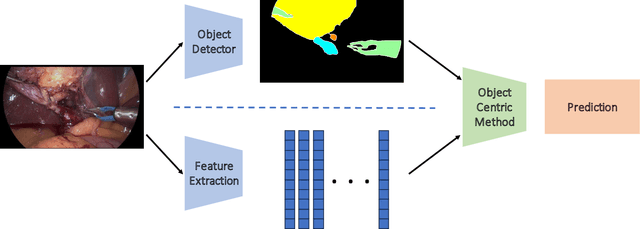

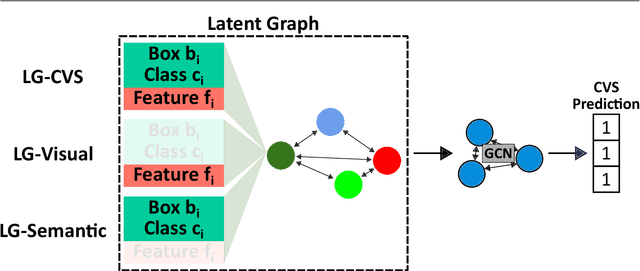
Abstract:Purpose: Advances in deep learning have resulted in effective models for surgical video analysis; however, these models often fail to generalize across medical centers due to domain shift caused by variations in surgical workflow, camera setups, and patient demographics. Recently, object-centric learning has emerged as a promising approach for improved surgical scene understanding, capturing and disentangling visual and semantic properties of surgical tools and anatomy to improve downstream task performance. In this work, we conduct a multi-centric performance benchmark of object-centric approaches, focusing on Critical View of Safety assessment in laparoscopic cholecystectomy, then propose an improved approach for unseen domain generalization. Methods: We evaluate four object-centric approaches for domain generalization, establishing baseline performance. Next, leveraging the disentangled nature of object-centric representations, we dissect one of these methods through a series of ablations (e.g. ignoring either visual or semantic features for downstream classification). Finally, based on the results of these ablations, we develop an optimized method specifically tailored for domain generalization, LG-DG, that includes a novel disentanglement loss function. Results: Our optimized approach, LG-DG, achieves an improvement of 9.28% over the best baseline approach. More broadly, we show that object-centric approaches are highly effective for domain generalization thanks to their modular approach to representation learning. Conclusion: We investigate the use of object-centric methods for unseen domain generalization, identify method-agnostic factors critical for performance, and present an optimized approach that substantially outperforms existing methods.
Overcoming Dimensional Collapse in Self-supervised Contrastive Learning for Medical Image Segmentation
Feb 27, 2024



Abstract:Self-supervised learning (SSL) approaches have achieved great success when the amount of labeled data is limited. Within SSL, models learn robust feature representations by solving pretext tasks. One such pretext task is contrastive learning, which involves forming pairs of similar and dissimilar input samples, guiding the model to distinguish between them. In this work, we investigate the application of contrastive learning to the domain of medical image analysis. Our findings reveal that MoCo v2, a state-of-the-art contrastive learning method, encounters dimensional collapse when applied to medical images. This is attributed to the high degree of inter-image similarity shared between the medical images. To address this, we propose two key contributions: local feature learning and feature decorrelation. Local feature learning improves the ability of the model to focus on the local regions of the image, while feature decorrelation removes the linear dependence among the features. Our experimental findings demonstrate that our contributions significantly enhance the model's performance in the downstream task of medical segmentation, both in the linear evaluation and full fine-tuning settings. This work illustrates the importance of effectively adapting SSL techniques to the characteristics of medical imaging tasks. The source code will be made publicly available at: https://github.com/CAMMA-public/med-moco
The Endoscapes Dataset for Surgical Scene Segmentation, Object Detection, and Critical View of Safety Assessment: Official Splits and Benchmark
Dec 19, 2023



Abstract:This technical report provides a detailed overview of Endoscapes, a dataset of laparoscopic cholecystectomy (LC) videos with highly intricate annotations targeted at automated assessment of the Critical View of Safety (CVS). Endoscapes comprises 201 LC videos with frames annotated sparsely but regularly with segmentation masks, bounding boxes, and CVS assessment by three different clinical experts. Altogether, there are 11090 frames annotated with CVS and 1933 frames annotated with tool and anatomy bounding boxes from the 201 videos, as well as an additional 422 frames from 50 of the 201 videos annotated with tool and anatomy segmentation masks. In this report, we provide detailed dataset statistics (size, class distribution, dataset splits, etc.) and a comprehensive performance benchmark for instance segmentation, object detection, and CVS prediction. The dataset and model checkpoints are publically available at https://github.com/CAMMA-public/Endoscapes.
ST(OR)2: Spatio-Temporal Object Level Reasoning for Activity Recognition in the Operating Room
Dec 19, 2023



Abstract:Surgical robotics holds much promise for improving patient safety and clinician experience in the Operating Room (OR). However, it also comes with new challenges, requiring strong team coordination and effective OR management. Automatic detection of surgical activities is a key requirement for developing AI-based intelligent tools to tackle these challenges. The current state-of-the-art surgical activity recognition methods however operate on image-based representations and depend on large-scale labeled datasets whose collection is time-consuming and resource-expensive. This work proposes a new sample-efficient and object-based approach for surgical activity recognition in the OR. Our method focuses on the geometric arrangements between clinicians and surgical devices, thus utilizing the significant object interaction dynamics in the OR. We conduct experiments in a low-data regime study for long video activity recognition. We also benchmark our method againstother object-centric approaches on clip-level action classification and show superior performance.
Challenges in Multi-centric Generalization: Phase and Step Recognition in Roux-en-Y Gastric Bypass Surgery
Dec 18, 2023Abstract:Most studies on surgical activity recognition utilizing Artificial intelligence (AI) have focused mainly on recognizing one type of activity from small and mono-centric surgical video datasets. It remains speculative whether those models would generalize to other centers. In this work, we introduce a large multi-centric multi-activity dataset consisting of 140 videos (MultiBypass140) of laparoscopic Roux-en-Y gastric bypass (LRYGB) surgeries performed at two medical centers: the University Hospital of Strasbourg (StrasBypass70) and Inselspital, Bern University Hospital (BernBypass70). The dataset has been fully annotated with phases and steps. Furthermore, we assess the generalizability and benchmark different deep learning models in 7 experimental studies: 1) Training and evaluation on BernBypass70; 2) Training and evaluation on StrasBypass70; 3) Training and evaluation on the MultiBypass140; 4) Training on BernBypass70, evaluation on StrasBypass70; 5) Training on StrasBypass70, evaluation on BernBypass70; Training on MultiBypass140, evaluation 6) on BernBypass70 and 7) on StrasBypass70. The model's performance is markedly influenced by the training data. The worst results were obtained in experiments 4) and 5) confirming the limited generalization capabilities of models trained on mono-centric data. The use of multi-centric training data, experiments 6) and 7), improves the generalization capabilities of the models, bringing them beyond the level of independent mono-centric training and validation (experiments 1) and 2)). MultiBypass140 shows considerable variation in surgical technique and workflow of LRYGB procedures between centers. Therefore, generalization experiments demonstrate a remarkable difference in model performance. These results highlight the importance of multi-centric datasets for AI model generalization to account for variance in surgical technique and workflows.
Advancing Surgical VQA with Scene Graph Knowledge
Dec 15, 2023Abstract:Modern operating room is becoming increasingly complex, requiring innovative intra-operative support systems. While the focus of surgical data science has largely been on video analysis, integrating surgical computer vision with language capabilities is emerging as a necessity. Our work aims to advance Visual Question Answering (VQA) in the surgical context with scene graph knowledge, addressing two main challenges in the current surgical VQA systems: removing question-condition bias in the surgical VQA dataset and incorporating scene-aware reasoning in the surgical VQA model design. First, we propose a Surgical Scene Graph-based dataset, SSG-QA, generated by employing segmentation and detection models on publicly available datasets. We build surgical scene graphs using spatial and action information of instruments and anatomies. These graphs are fed into a question engine, generating diverse QA pairs. Our SSG-QA dataset provides a more complex, diverse, geometrically grounded, unbiased, and surgical action-oriented dataset compared to existing surgical VQA datasets. We then propose SSG-QA-Net, a novel surgical VQA model incorporating a lightweight Scene-embedded Interaction Module (SIM), which integrates geometric scene knowledge in the VQA model design by employing cross-attention between the textual and the scene features. Our comprehensive analysis of the SSG-QA dataset shows that SSG-QA-Net outperforms existing methods across different question types and complexities. We highlight that the primary limitation in the current surgical VQA systems is the lack of scene knowledge to answer complex queries. We present a novel surgical VQA dataset and model and show that results can be significantly improved by incorporating geometric scene features in the VQA model design. The source code and the dataset will be made publicly available at: https://github.com/CAMMA-public/SSG-QA
MOSaiC: a Web-based Platform for Collaborative Medical Video Assessment and Annotation
Dec 14, 2023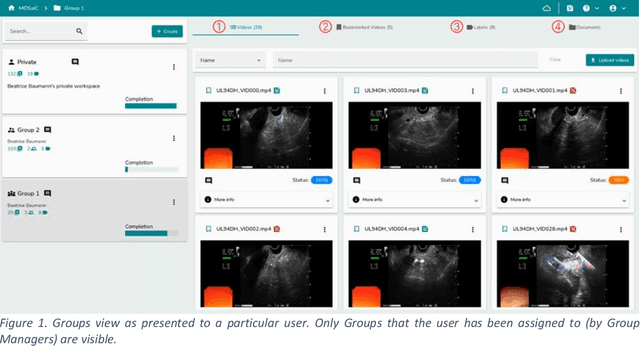
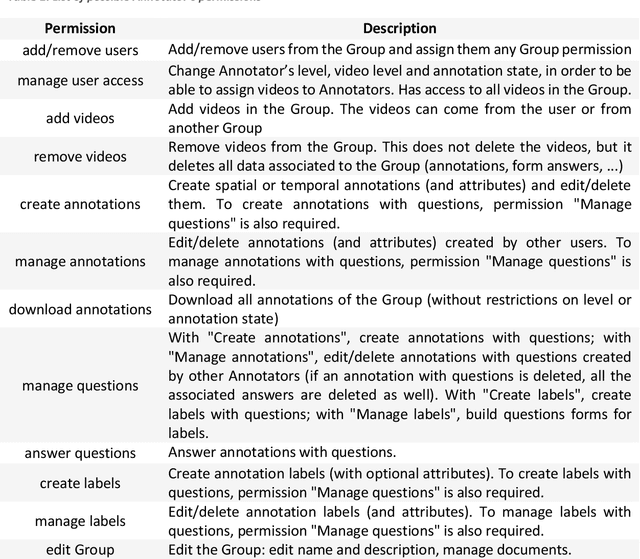
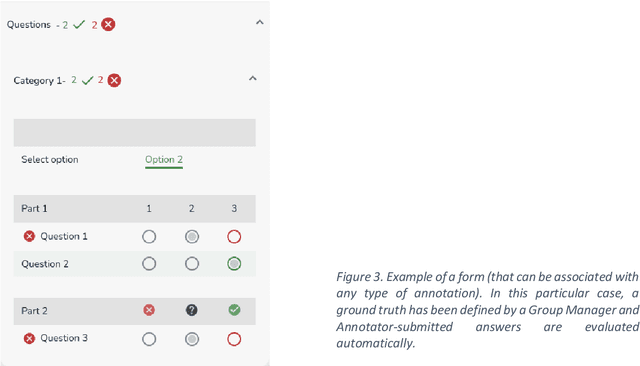
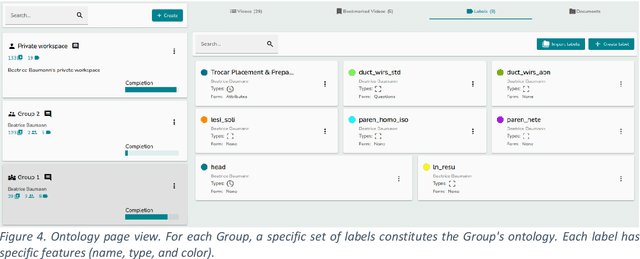
Abstract:This technical report presents MOSaiC 3.6.2, a web-based collaborative platform designed for the annotation and evaluation of medical videos. MOSaiC is engineered to facilitate video-based assessment and accelerate surgical data science projects. We provide an overview of MOSaiC's key functionalities, encompassing group and video management, annotation tools, ontologies, assessment capabilities, and user administration. Finally, we briefly describe several medical data science studies where MOSaiC has been instrumental in the dataset development.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge