Nasir Rajpoot
*: shared first/last authors
A QuadTree Image Representation for Computational Pathology
Aug 24, 2021
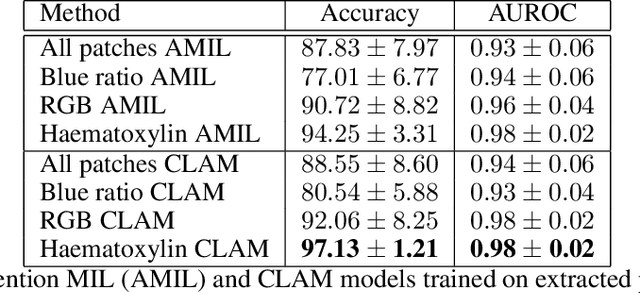

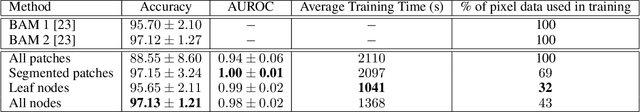
Abstract:The field of computational pathology presents many challenges for computer vision algorithms due to the sheer size of pathology images. Histopathology images are large and need to be split up into image tiles or patches so modern convolutional neural networks (CNNs) can process them. In this work, we present a method to generate an interpretable image representation of computational pathology images using quadtrees and a pipeline to use these representations for highly accurate downstream classification. To the best of our knowledge, this is the first attempt to use quadtrees for pathology image data. We show it is highly accurate, able to achieve as good results as the currently widely adopted tissue mask patch extraction methods all while using over 38% less data.
Cells are Actors: Social Network Analysis with Classical ML for SOTA Histology Image Classification
Jul 10, 2021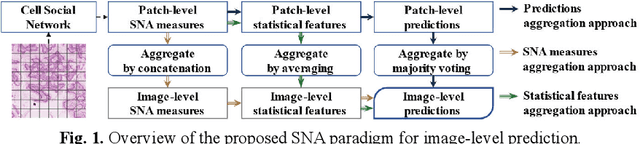
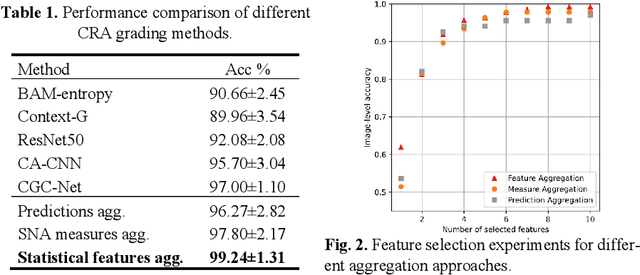
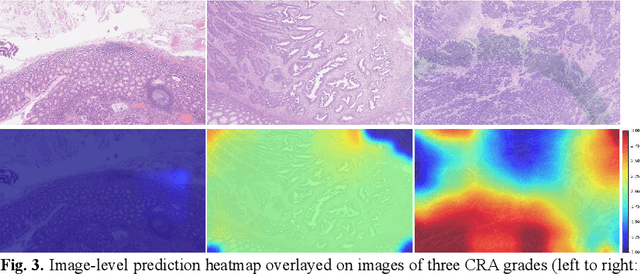
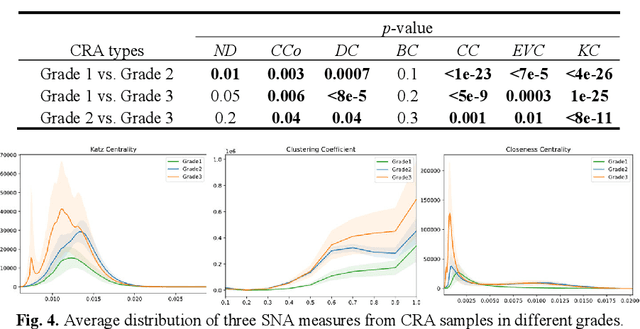
Abstract:Digitization of histology images and the advent of new computational methods, like deep learning, have helped the automatic grading of colorectal adenocarcinoma cancer (CRA). Present automated CRA grading methods, however, usually use tiny image patches and thus fail to integrate the entire tissue micro-architecture for grading purposes. To tackle these challenges, we propose to use a statistical network analysis method to describe the complex structure of the tissue micro-environment by modelling nuclei and their connections as a network. We show that by analyzing only the interactions between the cells in a network, we can extract highly discriminative statistical features for CRA grading. Unlike other deep learning or convolutional graph-based approaches, our method is highly scalable (can be used for cell networks consist of millions of nodes), completely explainable, and computationally inexpensive. We create cell networks on a broad CRC histology image dataset, experiment with our method, and report state-of-the-art performance for the prediction of three-class CRA grading.
Semantic annotation for computational pathology: Multidisciplinary experience and best practice recommendations
Jun 25, 2021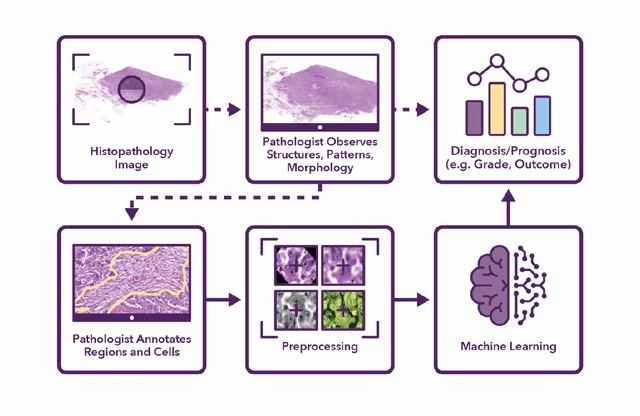
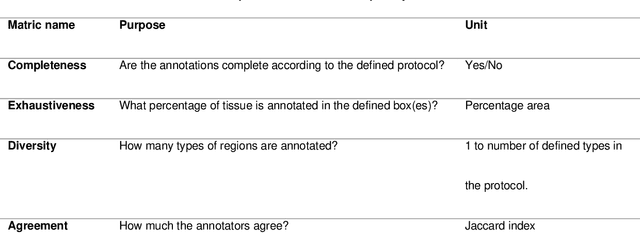

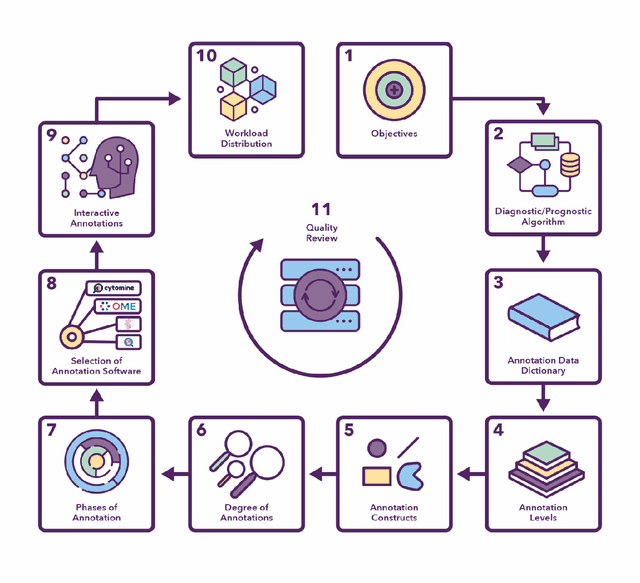
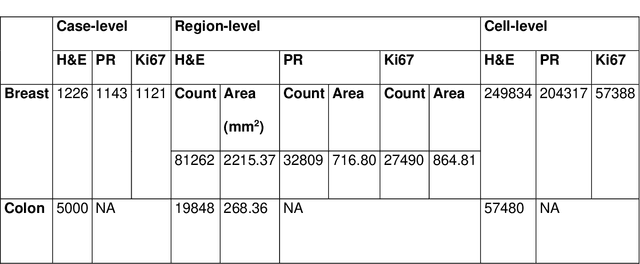
Abstract:Recent advances in whole slide imaging (WSI) technology have led to the development of a myriad of computer vision and artificial intelligence (AI) based diagnostic, prognostic, and predictive algorithms. Computational Pathology (CPath) offers an integrated solution to utilize information embedded in pathology WSIs beyond what we obtain through visual assessment. For automated analysis of WSIs and validation of machine learning (ML) models, annotations at the slide, tissue and cellular levels are required. The annotation of important visual constructs in pathology images is an important component of CPath projects. Improper annotations can result in algorithms which are hard to interpret and can potentially produce inaccurate and inconsistent results. Despite the crucial role of annotations in CPath projects, there are no well-defined guidelines or best practices on how annotations should be carried out. In this paper, we address this shortcoming by presenting the experience and best practices acquired during the execution of a large-scale annotation exercise involving a multidisciplinary team of pathologists, ML experts and researchers as part of the Pathology image data Lake for Analytics, Knowledge and Education (PathLAKE) consortium. We present a real-world case study along with examples of different types of annotations, diagnostic algorithm, annotation data dictionary and annotation constructs. The analyses reported in this work highlight best practice recommendations that can be used as annotation guidelines over the lifecycle of a CPath project.
Now You See It, Now You Dont: Adversarial Vulnerabilities in Computational Pathology
Jun 16, 2021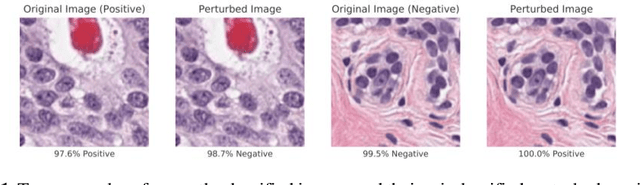

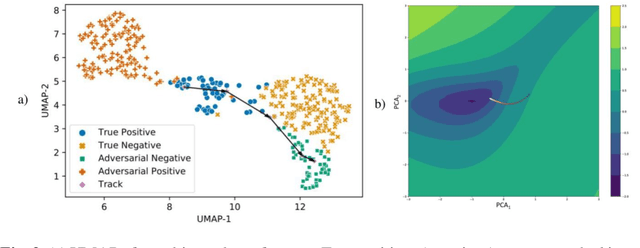
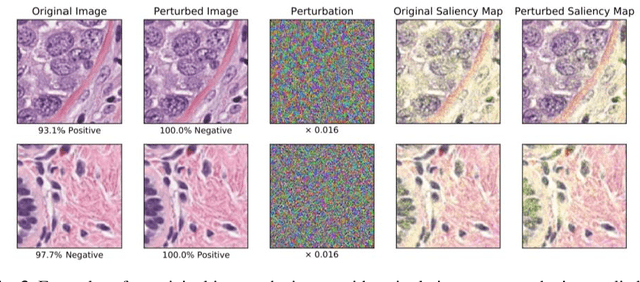
Abstract:Deep learning models are routinely employed in computational pathology (CPath) for solving problems of diagnostic and prognostic significance. Typically, the generalization performance of CPath models is analyzed using evaluation protocols such as cross-validation and testing on multi-centric cohorts. However, to ensure that such CPath solutions are robust and safe for use in a clinical setting, a critical analysis of their predictive performance and vulnerability to adversarial attacks is required, which is the focus of this paper. Specifically, we show that a highly accurate model for classification of tumour patches in pathology images (AUC > 0.95) can easily be attacked with minimal perturbations which are imperceptible to lay humans and trained pathologists alike. Our analytical results show that it is possible to generate single-instance white-box attacks on specific input images with high success rate and low perturbation energy. Furthermore, we have also generated a single universal perturbation matrix using the training dataset only which, when added to unseen test images, results in forcing the trained neural network to flip its prediction labels with high confidence at a success rate of > 84%. We systematically analyze the relationship between perturbation energy of an adversarial attack, its impact on morphological constructs of clinical significance, their perceptibility by a trained pathologist and saliency maps obtained using deep learning models. Based on our analysis, we strongly recommend that computational pathology models be critically analyzed using the proposed adversarial validation strategy prior to clinical adoption.
A digital score of tumour-associated stroma infiltrating lymphocytes predicts survival in head and neck squamous cell carcinoma
Apr 16, 2021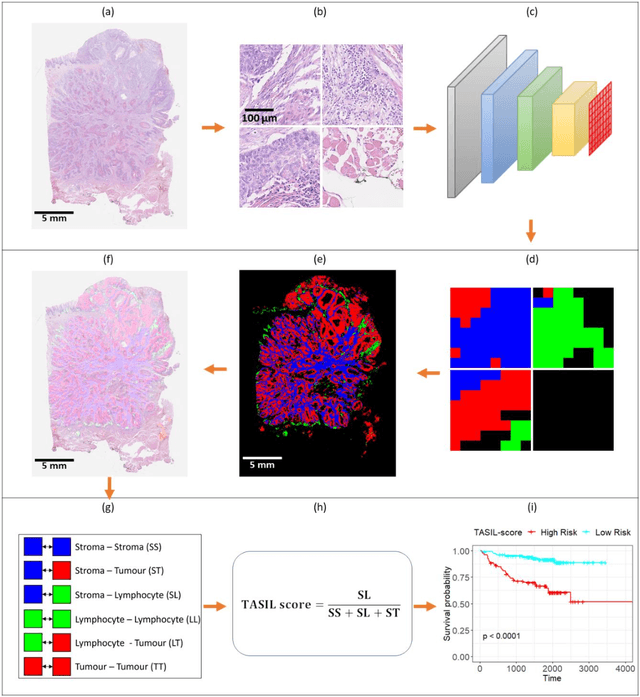
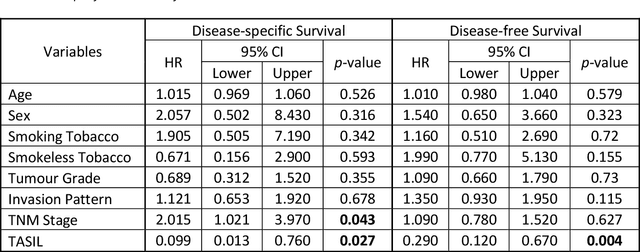
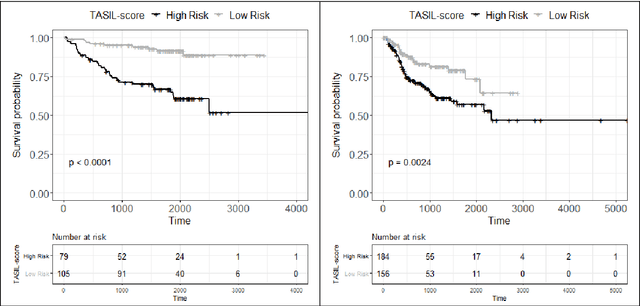
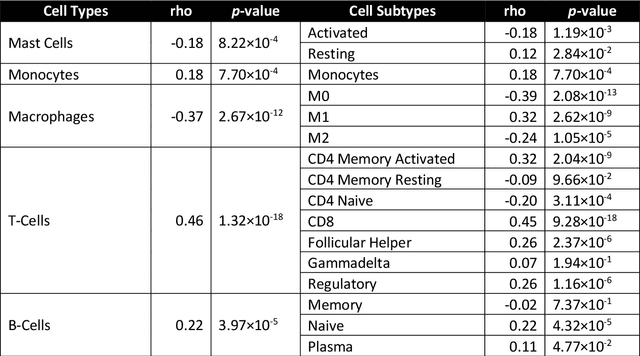
Abstract:The infiltration of T-lymphocytes in the stroma and tumour is an indication of an effective immune response against the tumour, resulting in better survival. In this study, our aim is to explore the prognostic significance of tumour-associated stroma infiltrating lymphocytes (TASILs) in head and neck squamous cell carcinoma (HNSCC) through an AI based automated method. A deep learning based automated method was employed to segment tumour, stroma and lymphocytes in digitally scanned whole slide images of HNSCC tissue slides. The spatial patterns of lymphocytes and tumour-associated stroma were digitally quantified to compute the TASIL-score. Finally, prognostic significance of the TASIL-score for disease-specific and disease-free survival was investigated with the Cox proportional hazard analysis. Three different cohorts of Haematoxylin & Eosin (H&E) stained tissue slides of HNSCC cases (n=537 in total) were studied, including publicly available TCGA head and neck cancer cases. The TASIL-score carries prognostic significance (p=0.002) for disease-specific survival of HNSCC patients. The TASIL-score also shows a better separation between low- and high-risk patients as compared to the manual TIL scoring by pathologists for both disease-specific and disease-free survival. A positive correlation of TASIL-score with molecular estimates of CD8+ T cells was also found, which is in line with existing findings. To the best of our knowledge, this is the first study to automate the quantification of TASIL from routine H&E slides of head and neck cancer. Our TASIL-score based findings are aligned with the clinical knowledge with the added advantages of objectivity, reproducibility and strong prognostic value. A comprehensive evaluation on large multicentric cohorts is required before the proposed digital score can be adopted in clinical practice.
Deep Multi-Resolution Dictionary Learning for Histopathology Image Analysis
Apr 01, 2021
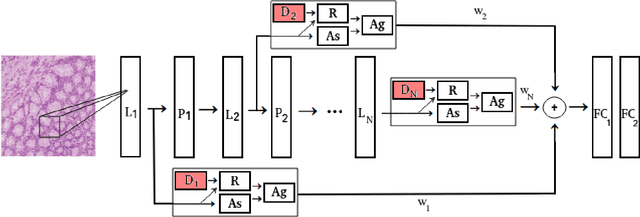
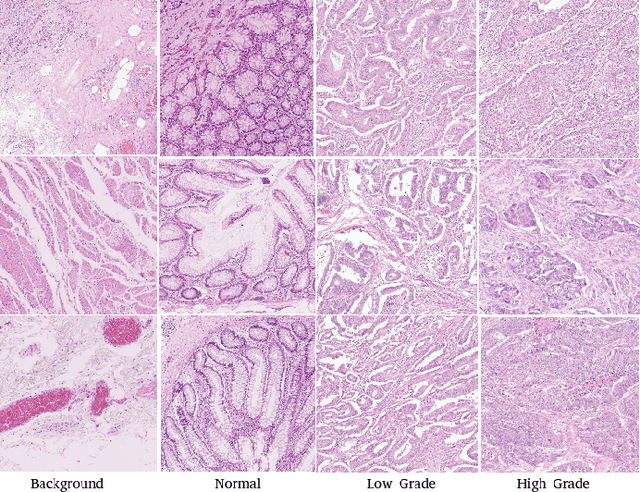
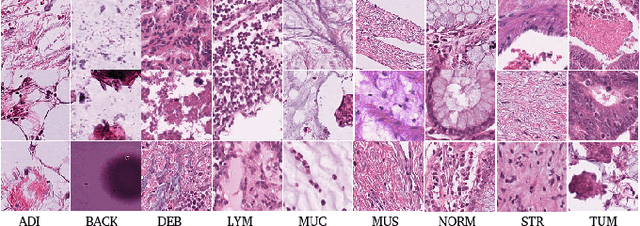
Abstract:The problem of recognizing various types of tissues present in multi-gigapixel histology images is an important fundamental pre-requisite for downstream analysis of the tumor microenvironment in a bottom-up analysis paradigm for computational pathology. In this paper, we propose a deep dictionary learning approach to solve the problem of tissue phenotyping in histology images. We propose deep Multi-Resolution Dictionary Learning (deepMRDL) in order to benefit from deep texture descriptors at multiple different spatial resolutions. We show the efficacy of the proposed approach through extensive experiments on four benchmark histology image datasets from different organs (colorectal cancer, breast cancer and breast lymphnodes) and tasks (namely, cancer grading, tissue phenotyping, tumor detection and tissue type classification). We also show that the proposed framework can employ most off-the-shelf CNNs models to generate effective deep texture descriptors.
Multiple Instance Captioning: Learning Representations from Histopathology Textbooks and Articles
Mar 08, 2021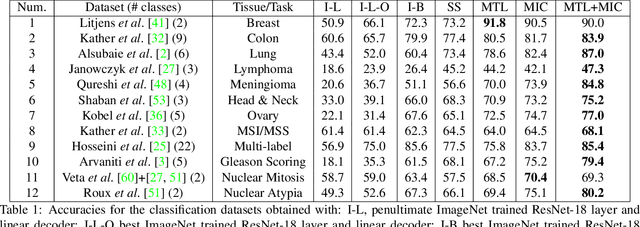
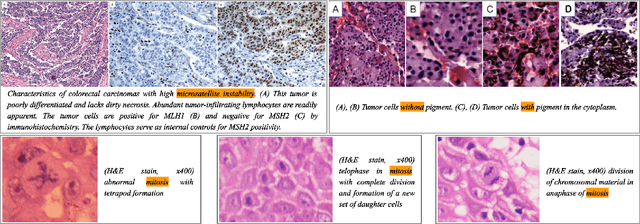
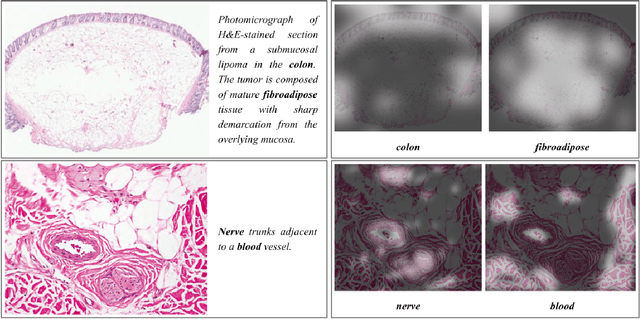
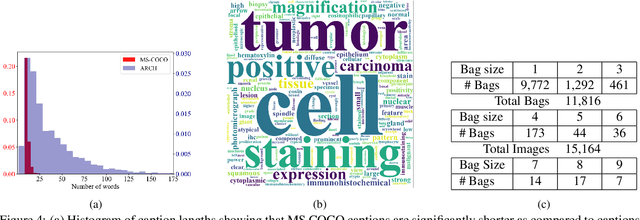
Abstract:We present ARCH, a computational pathology (CP) multiple instance captioning dataset to facilitate dense supervision of CP tasks. Existing CP datasets focus on narrow tasks; ARCH on the other hand contains dense diagnostic and morphological descriptions for a range of stains, tissue types and pathologies. Using intrinsic dimensionality estimation, we show that ARCH is the only CP dataset to (ARCH-)rival its computer vision analog MS-COCO Captions. We conjecture that an encoder pre-trained on dense image captions learns transferable representations for most CP tasks. We support the conjecture with evidence that ARCH representation transfers to a variety of pathology sub-tasks better than ImageNet features or representations obtained via self-supervised or multi-task learning on pathology images alone. We release our best model and invite other researchers to test it on their CP tasks.
Classification of COVID-19 via Homology of CT-SCAN
Feb 21, 2021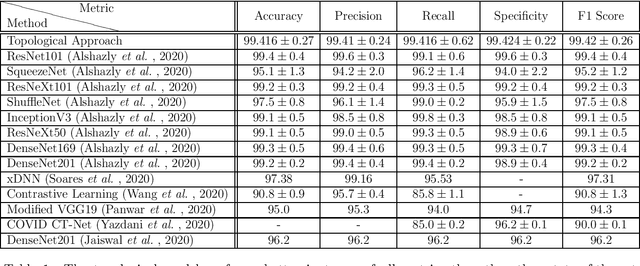
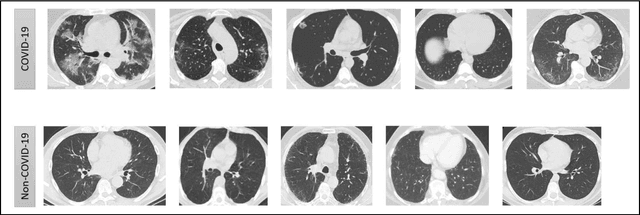
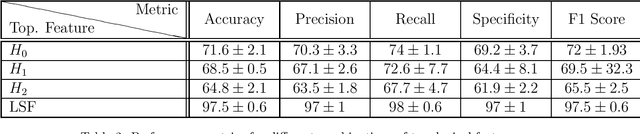
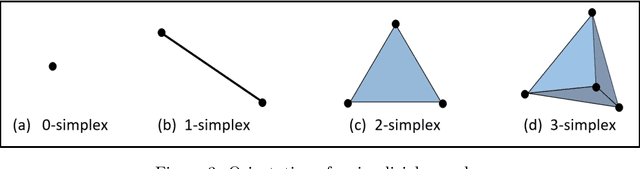
Abstract:In this worldwide spread of SARS-CoV-2 (COVID-19) infection, it is of utmost importance to detect the disease at an early stage especially in the hot spots of this epidemic. There are more than 110 Million infected cases on the globe, sofar. Due to its promptness and effective results computed tomography (CT)-scan image is preferred to the reverse-transcription polymerase chain reaction (RT-PCR). Early detection and isolation of the patient is the only possible way of controlling the spread of the disease. Automated analysis of CT-Scans can provide enormous support in this process. In this article, We propose a novel approach to detect SARS-CoV-2 using CT-scan images. Our method is based on a very intuitive and natural idea of analyzing shapes, an attempt to mimic a professional medic. We mainly trace SARS-CoV-2 features by quantifying their topological properties. We primarily use a tool called persistent homology, from Topological Data Analysis (TDA), to compute these topological properties. We train and test our model on the "SARS-CoV-2 CT-scan dataset" \citep{soares2020sars}, an open-source dataset, containing 2,481 CT-scans of normal and COVID-19 patients. Our model yielded an overall benchmark F1 score of $99.42\% $, accuracy $99.416\%$, precision $99.41\%$, and recall $99.42\%$. The TDA techniques have great potential that can be utilized for efficient and prompt detection of COVID-19. The immense potential of TDA may be exploited in clinics for rapid and safe detection of COVID-19 globally, in particular in the low and middle-income countries where RT-PCR labs and/or kits are in a serious crisis.
Self-Path: Self-supervision for Classification of Pathology Images with Limited Annotations
Aug 12, 2020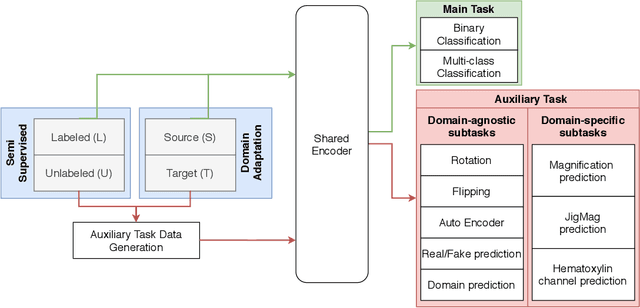
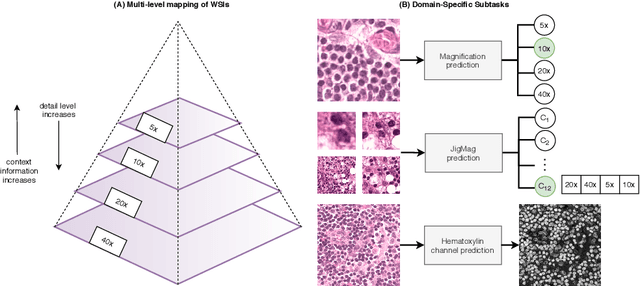
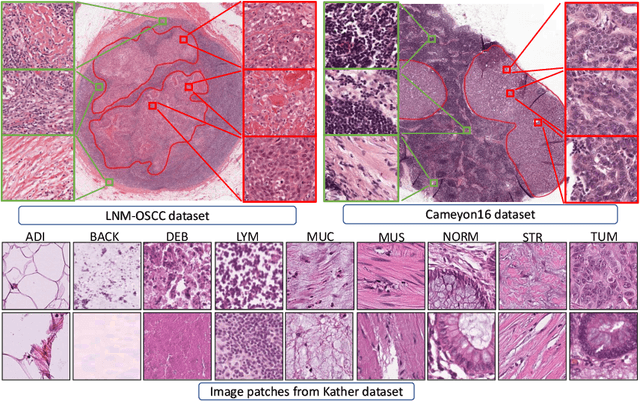
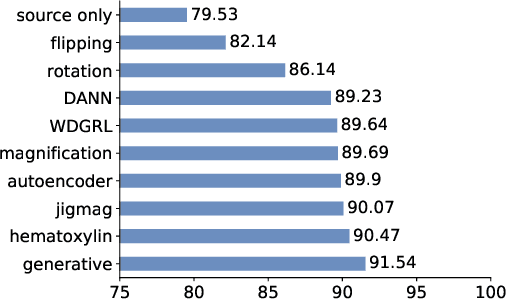
Abstract:While high-resolution pathology images lend themselves well to `data hungry' deep learning algorithms, obtaining exhaustive annotations on these images is a major challenge. In this paper, we propose a self-supervised CNN approach to leverage unlabeled data for learning generalizable and domain invariant representations in pathology images. The proposed approach, which we term as Self-Path, is a multi-task learning approach where the main task is tissue classification and pretext tasks are a variety of self-supervised tasks with labels inherent to the input data. We introduce novel domain specific self-supervision tasks that leverage contextual, multi-resolution and semantic features in pathology images for semi-supervised learning and domain adaptation. We investigate the effectiveness of Self-Path on 3 different pathology datasets. Our results show that Self-Path with the domain-specific pretext tasks achieves state-of-the-art performance for semi-supervised learning when small amounts of labeled data are available. Further, we show that Self-Path improves domain adaptation for classification of histology image patches when there is no labeled data available for the target domain. This approach can potentially be employed for other applications in computational pathology, where annotation budget is often limited or large amount of unlabeled image data is available.
SAFRON: Stitching Across the Frontier for Generating Colorectal Cancer Histology Images
Aug 11, 2020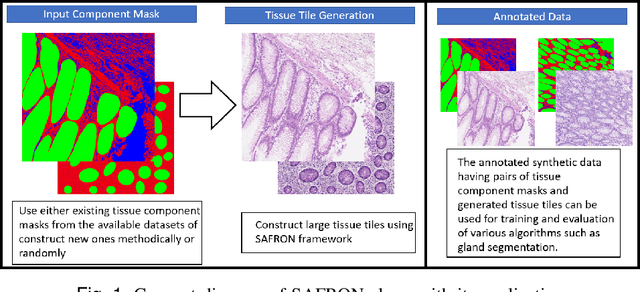
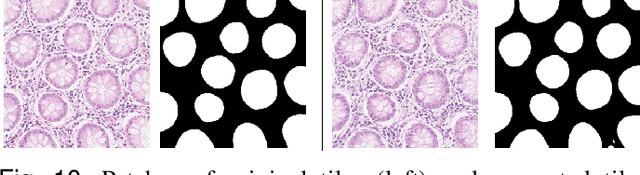
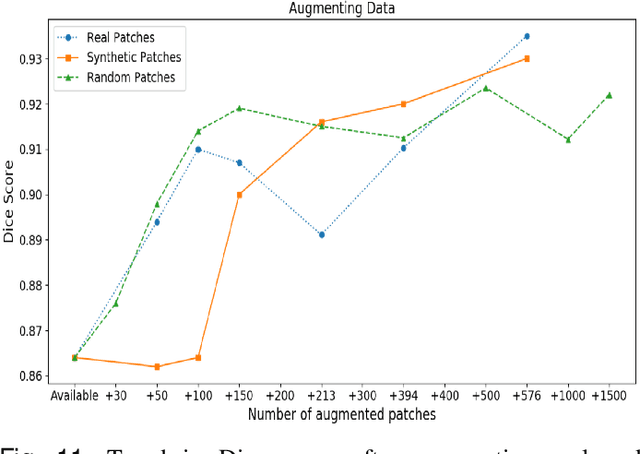

Abstract:Synthetic images can be used for the development and evaluation of deep learning algorithms in the context of limited availability of annotations. In the field of computational pathology where histology images are large and visual context is crucial, synthesis of large tissue images via generative modeling is a challenging task due to memory and computing constraints hindering the generation of large images. To address this challenge, we propose a novel framework named as SAFRON to construct realistic large tissue image tiles from ground truth annotations while preserving morphological features and with minimal boundary artifacts at the seams. To this end, we train the proposed SAFRON framework based on conditional generative adversarial networks on large tissue image tiles from the Colorectal Adenocarcinoma Gland (CRAG) and DigestPath datasets. We demonstrate that our model can generate high quality and realistic image tiles of arbitrary large size after training it on relatively small image patches. We also show that training on synthetic data generated by SAFRON can significantly boost the performance of a standard algorithm for gland segmentation of colorectal cancer tissue images. Sample high resolution images generated using SAFRON are available at the URL:https://warwick.ac.uk/TIALab/SAFRON
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge