Jevgenij Gamper
Towards Measuring Sell Side Outcomes in Buy Side Marketplace Experiments using In-Experiment Bipartite Graph
Sep 06, 2024


Abstract:In this study, we evaluate causal inference estimators for online controlled bipartite graph experiments in a real marketplace setting. Our novel contribution is constructing a bipartite graph using in-experiment data, rather than relying on prior knowledge or historical data, the common approach in the literature published to date. We build the bipartite graph from various interactions between buyers and sellers in the marketplace, establishing a novel research direction at the intersection of bipartite experiments and mediation analysis. This approach is crucial for modern marketplaces aiming to evaluate seller-side causal effects in buyer-side experiments, or vice versa. We demonstrate our method using historical buyer-side experiments conducted at Vinted, the largest second-hand marketplace in Europe with over 80M users.
Challenges and opportunities in applying Neural Temporal Point Processes to large scale industry data
Aug 18, 2022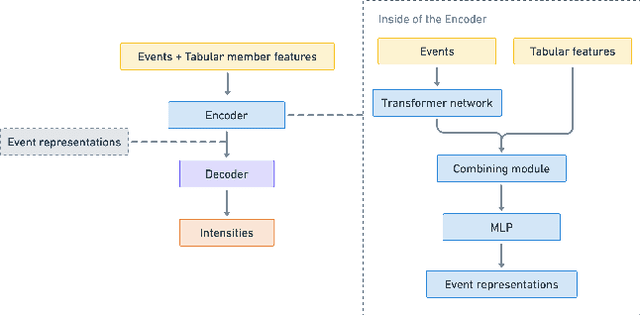
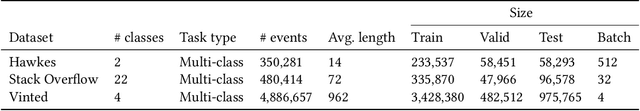
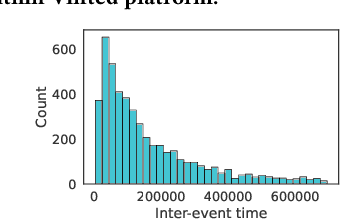
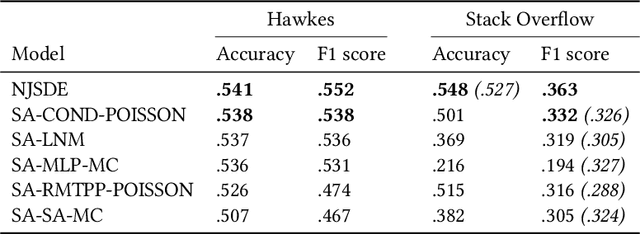
Abstract:In this work, we identify open research opportunities in applying Neural Temporal Point Process (NTPP) models to industry scale customer behavior data by carefully reproducing NTPP models published up to date on known literature benchmarks as well as applying NTPP models to a novel, real world consumer behavior dataset that is twice as large as the largest publicly available NTPP benchmark. We identify the following challenges. First, NTPP models, albeit their generative nature, remain vulnerable to dataset imbalances and cannot forecast rare events. Second, NTPP models based on stochastic differential equations, despite their theoretical appeal and leading performance on literature benchmarks, do not scale easily to large industry-scale data. The former is in light of previously made observations on deep generative models. Additionally, to combat a cold-start problem, we explore a novel addition to NTPP models - a parametrization based on static user features.
Multiple Instance Captioning: Learning Representations from Histopathology Textbooks and Articles
Mar 08, 2021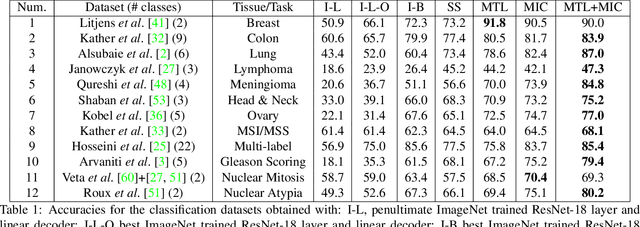
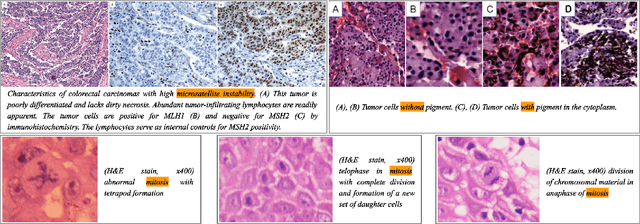
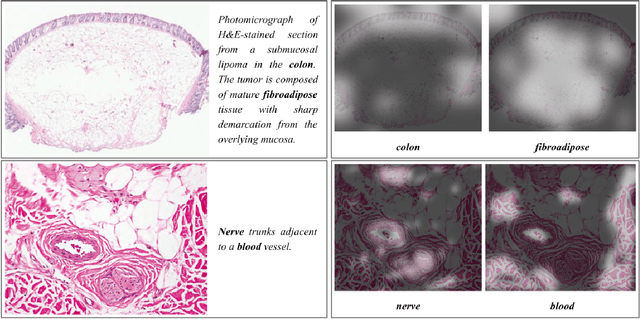
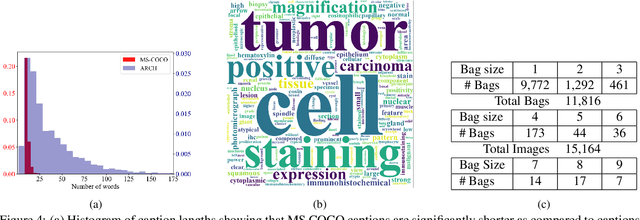
Abstract:We present ARCH, a computational pathology (CP) multiple instance captioning dataset to facilitate dense supervision of CP tasks. Existing CP datasets focus on narrow tasks; ARCH on the other hand contains dense diagnostic and morphological descriptions for a range of stains, tissue types and pathologies. Using intrinsic dimensionality estimation, we show that ARCH is the only CP dataset to (ARCH-)rival its computer vision analog MS-COCO Captions. We conjecture that an encoder pre-trained on dense image captions learns transferable representations for most CP tasks. We support the conjecture with evidence that ARCH representation transfers to a variety of pathology sub-tasks better than ImageNet features or representations obtained via self-supervised or multi-task learning on pathology images alone. We release our best model and invite other researchers to test it on their CP tasks.
Predicting Landsat Reflectance with Deep Generative Fusion
Nov 09, 2020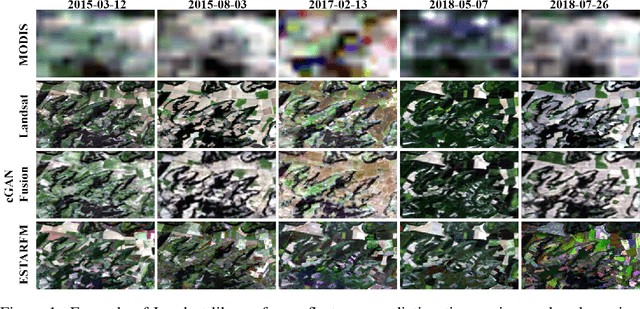
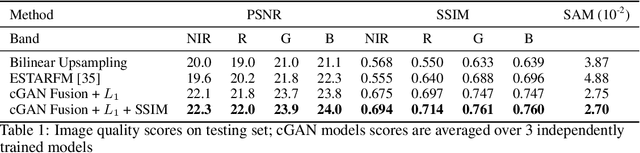
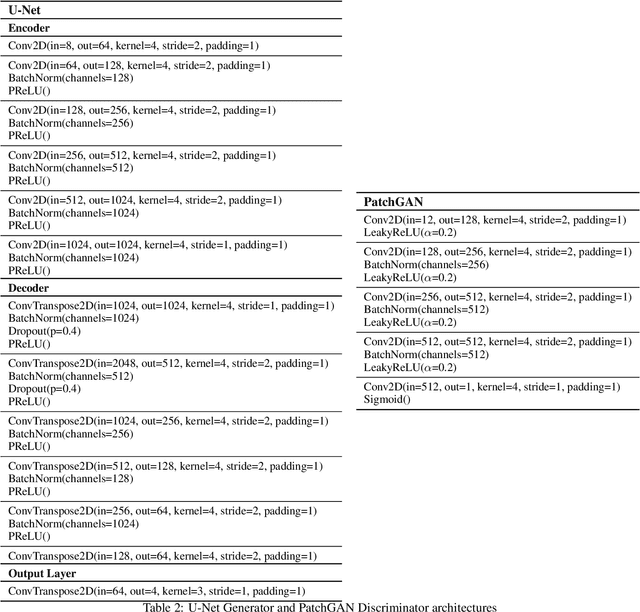
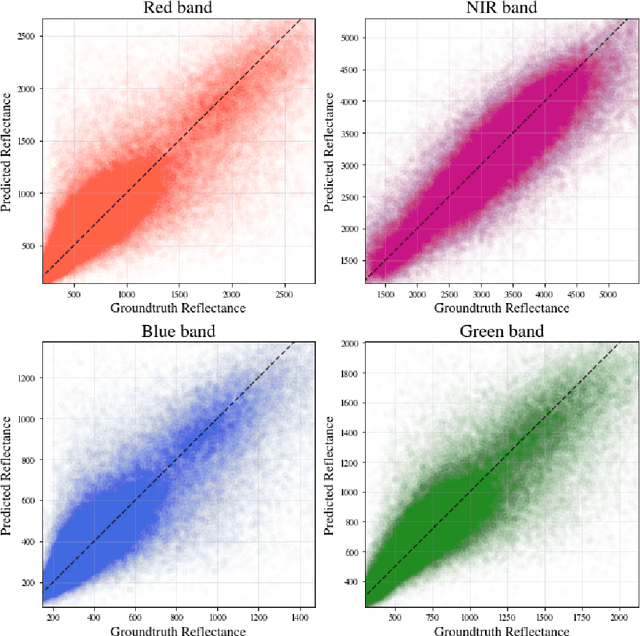
Abstract:Public satellite missions are commonly bound to a trade-off between spatial and temporal resolution as no single sensor provides fine-grained acquisitions with frequent coverage. This hinders their potential to assist vegetation monitoring or humanitarian actions, which require detecting rapid and detailed terrestrial surface changes. In this work, we probe the potential of deep generative models to produce high-resolution optical imagery by fusing products with different spatial and temporal characteristics. We introduce a dataset of co-registered Moderate Resolution Imaging Spectroradiometer (MODIS) and Landsat surface reflectance time series and demonstrate the ability of our generative model to blend coarse daily reflectance information into low-paced finer acquisitions. We benchmark our proposed model against state-of-the-art reflectance fusion algorithms.
Exoplanet Validation with Machine Learning: 50 new validated Kepler planets
Aug 24, 2020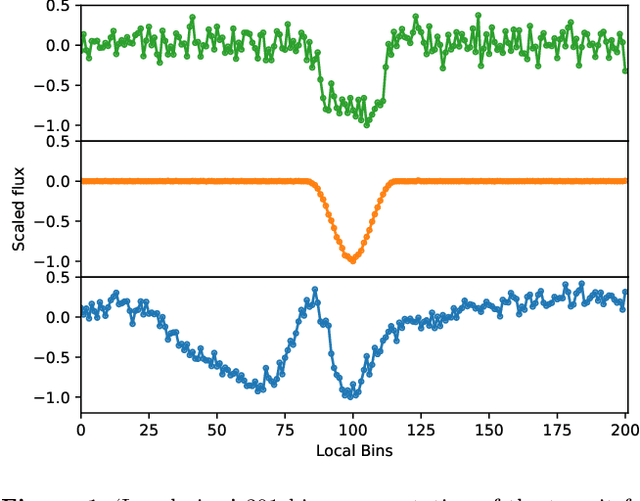
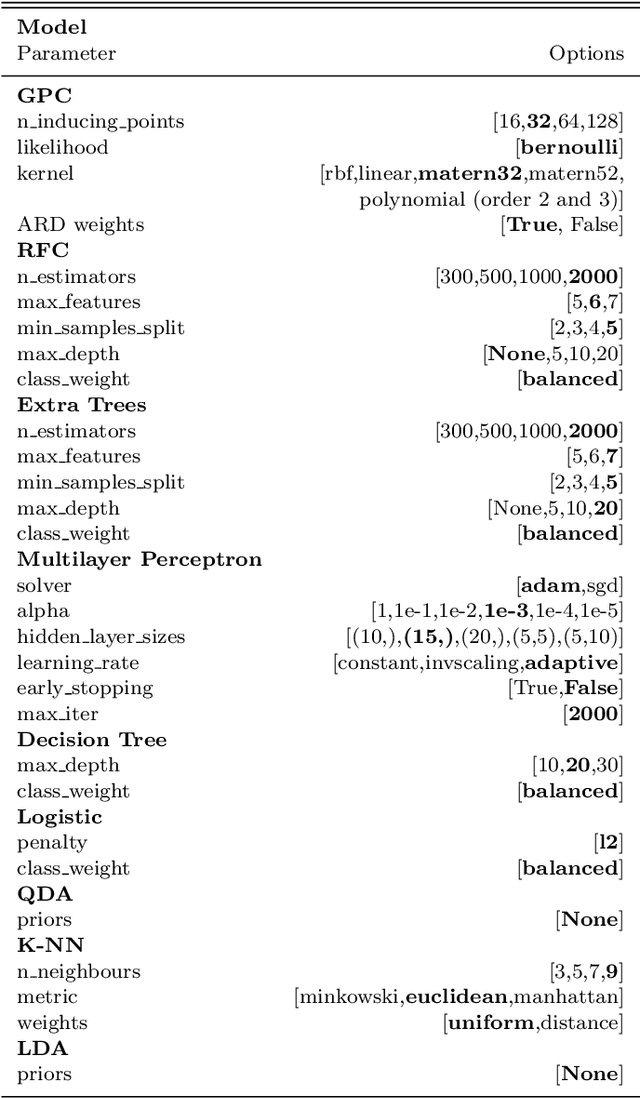
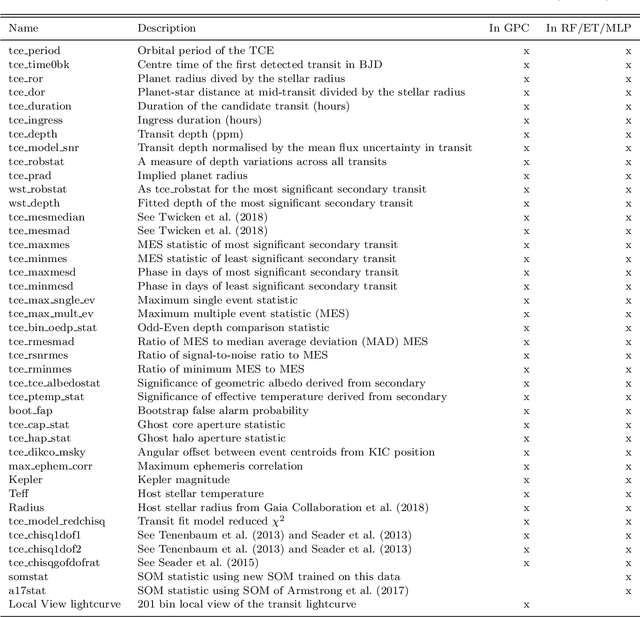
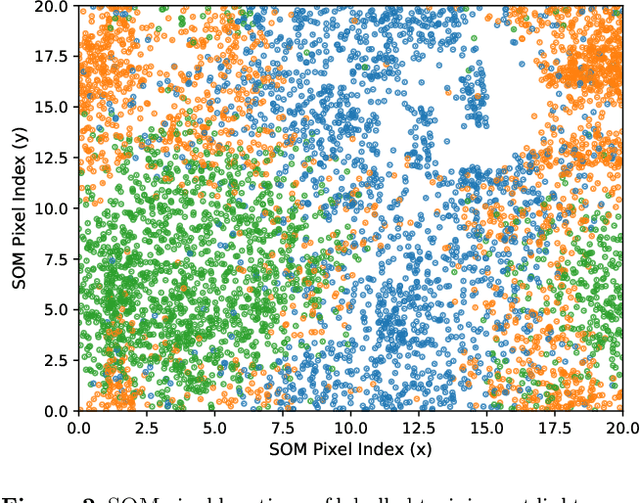

Abstract:Over 30% of the ~4000 known exoplanets to date have been discovered using 'validation', where the statistical likelihood of a transit arising from a false positive (FP), non-planetary scenario is calculated. For the large majority of these validated planets calculations were performed using the vespa algorithm (Morton et al. 2016). Regardless of the strengths and weaknesses of vespa, it is highly desirable for the catalogue of known planets not to be dependent on a single method. We demonstrate the use of machine learning algorithms, specifically a gaussian process classifier (GPC) reinforced by other models, to perform probabilistic planet validation incorporating prior probabilities for possible FP scenarios. The GPC can attain a mean log-loss per sample of 0.54 when separating confirmed planets from FPs in the Kepler threshold crossing event (TCE) catalogue. Our models can validate thousands of unseen candidates in seconds once applicable vetting metrics are calculated, and can be adapted to work with the active TESS mission, where the large number of observed targets necessitates the use of automated algorithms. We discuss the limitations and caveats of this methodology, and after accounting for possible failure modes newly validate 50 Kepler candidates as planets, sanity checking the validations by confirming them with vespa using up to date stellar information. Concerning discrepancies with vespa arise for many other candidates, which typically resolve in favour of our models. Given such issues, we caution against using single-method planet validation with either method until the discrepancies are fully understood.
Multi-Task Learning in Histo-pathology for Widely Generalizable Model
May 09, 2020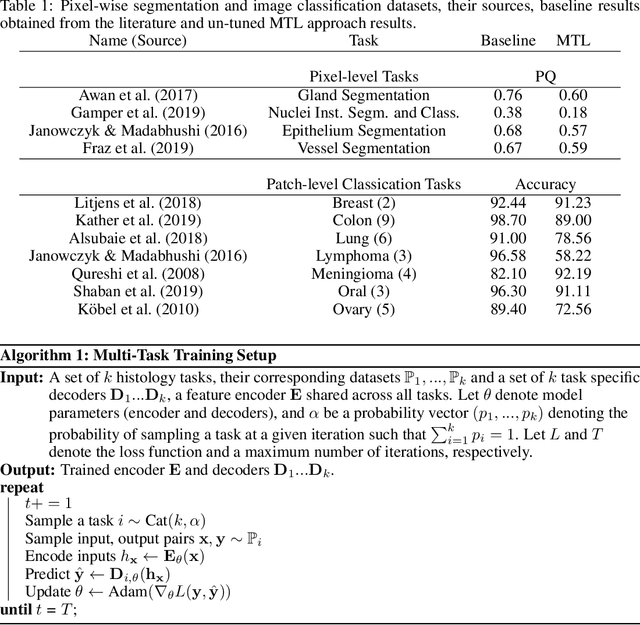
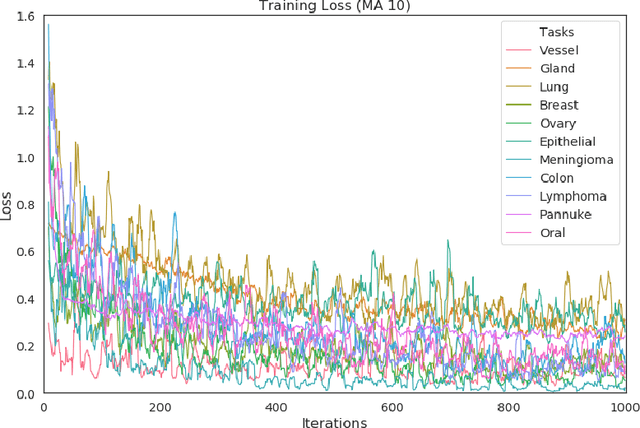
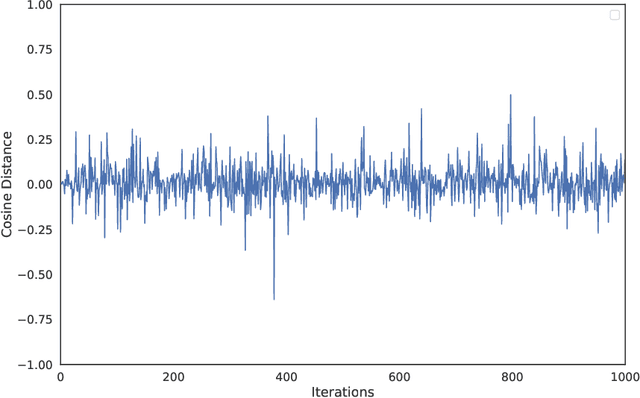
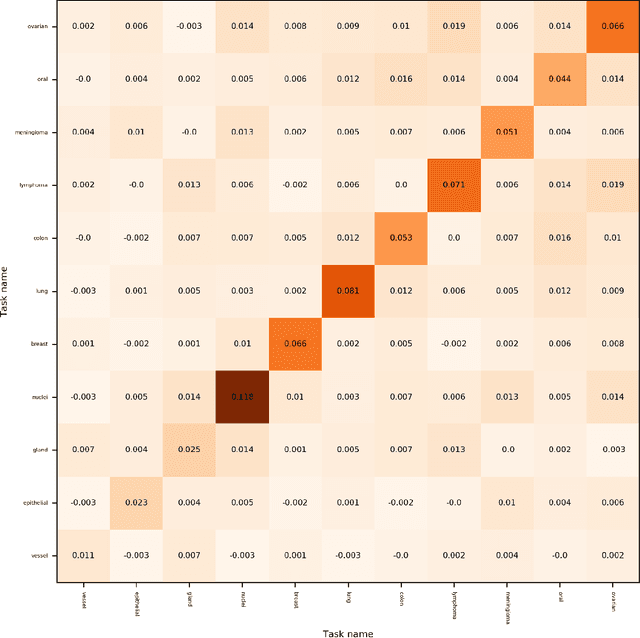
Abstract:In this work we show preliminary results of deep multi-task learning in the area of computational pathology. We combine 11 tasks ranging from patch-wise oral cancer classification, one of the most prevalent cancers in the developing world, to multi-tissue nuclei instance segmentation and classification.
PanNuke Dataset Extension, Insights and Baselines
Apr 22, 2020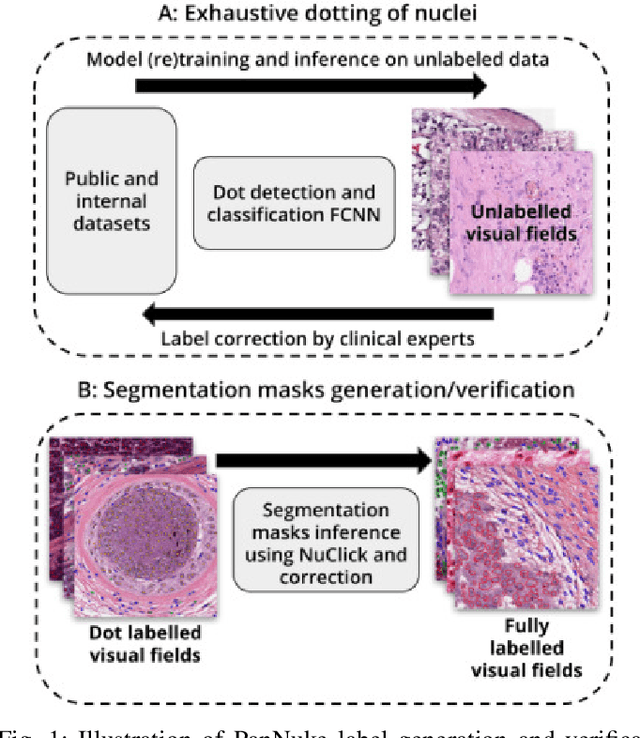
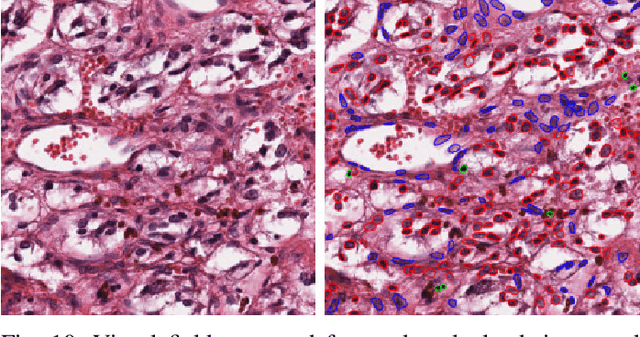
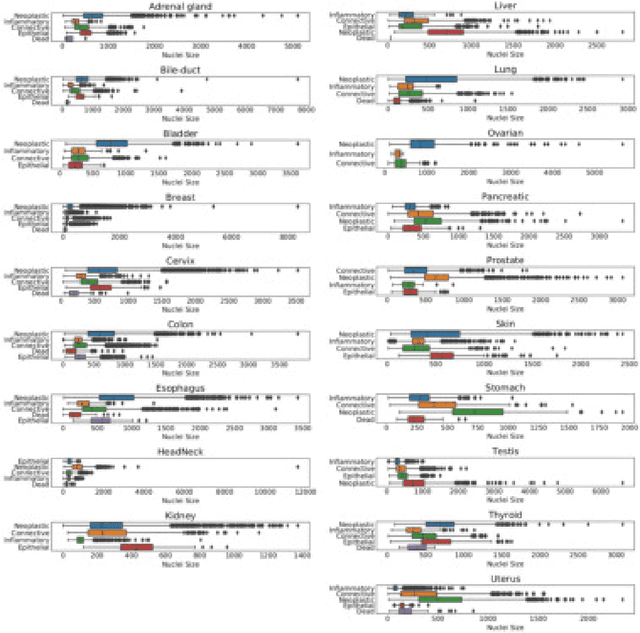
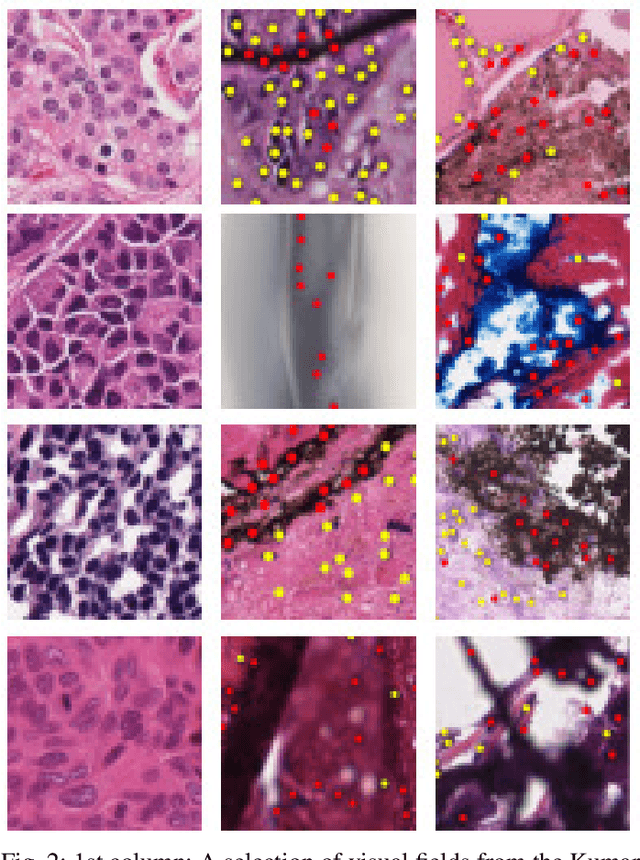
Abstract:The emerging area of computational pathology (CPath) is ripe ground for the application of deep learning (DL) methods to healthcare due to the sheer volume of raw pixel data in whole-slide images (WSIs) of cancerous tissue slides. However, it is imperative for the DL algorithms relying on nuclei-level details to be able to cope with data from `the clinical wild', which tends to be quite challenging. We study, and extend recently released PanNuke dataset consisting of ~200,000 nuclei categorized into 5 clinically important classes for the challenging tasks of segmenting and classifying nuclei in WSIs. Previous pan-cancer datasets consisted of only up to 9 different tissues and up to 21,000 unlabeled nuclei and just over 24,000 labeled nuclei with segmentation masks. PanNuke consists of 19 different tissue types that have been semi-automatically annotated and quality controlled by clinical pathologists, leading to a dataset with statistics similar to the clinical wild and with minimal selection bias. We study the performance of segmentation and classification models when applied to the proposed dataset and demonstrate the application of models trained on PanNuke to whole-slide images. We provide comprehensive statistics about the dataset and outline recommendations and research directions to address the limitations of existing DL tools when applied to real-world CPath applications.
Meta-SVDD: Probabilistic Meta-Learning for One-Class Classification in Cancer Histology Images
Mar 06, 2020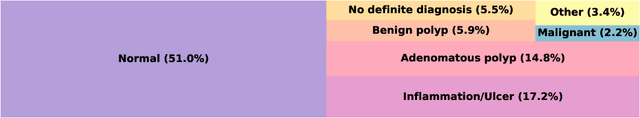
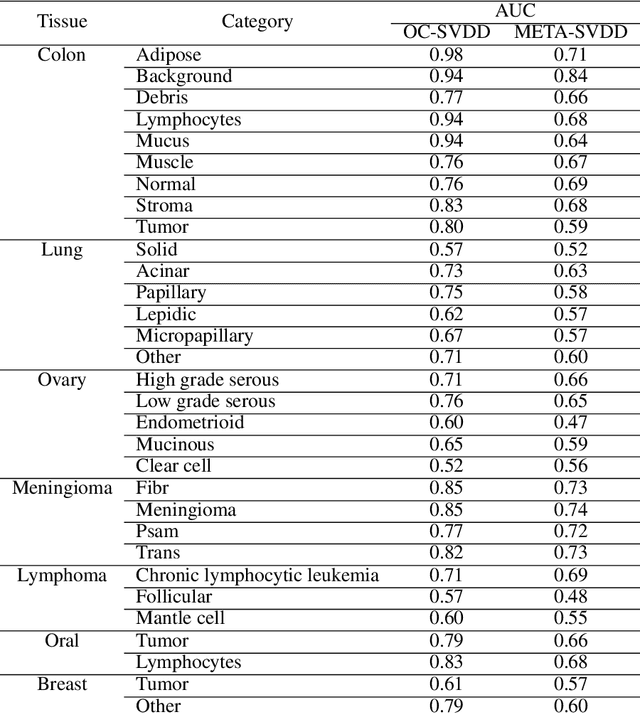
Abstract:To train a robust deep learning model, one usually needs a balanced set of categories in the training data. The data acquired in a medical domain, however, frequently contains an abundance of healthy patients, versus a small variety of positive, abnormal cases. Moreover, the annotation of a positive sample requires time consuming input from medical domain experts. This scenario would suggest a promise for one-class classification type approaches. In this work we propose a general one-class classification model for histology, that is meta-trained on multiple histology datasets simultaneously, and can be applied to new tasks without expensive re-training. This model could be easily used by pathology domain experts, and potentially be used for screening purposes.
AI Ethics for Systemic Issues: A Structural Approach
Nov 08, 2019Abstract:The debate on AI ethics largely focuses on technical improvements and stronger regulation to prevent accidents or misuse of AI, with solutions relying on holding individual actors accountable for responsible AI development. While useful and necessary, we argue that this "agency" approach disregards more indirect and complex risks resulting from AI's interaction with the socio-economic and political context. This paper calls for a "structural" approach to assessing AI's effects in order to understand and prevent such systemic risks where no individual can be held accountable for the broader negative impacts. This is particularly relevant for AI applied to systemic issues such as climate change and food security which require political solutions and global cooperation. To properly address the wide range of AI risks and ensure 'AI for social good', agency-focused policies must be complemented by policies informed by a structural approach.
Conditional Denoising of Remote Sensing Imagery Using Cycle-Consistent Deep Generative Models
Oct 31, 2019
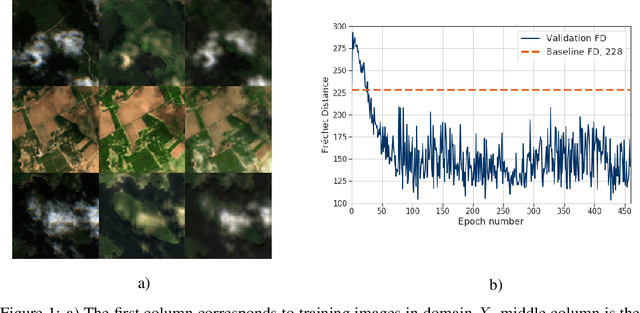

Abstract:The potential of using remote sensing imagery for environmental modelling and for providing real time support to humanitarian operations such as hurricane relief efforts is well established. These applications are substantially affected by missing data due to non-structural noise such as clouds, shadows and other atmospheric effects. In this work we probe the potential of applying a cycle-consistent latent variable deep generative model (DGM) for denoising cloudy Sentinel-2 observations conditioned on the information in cloud penetrating bands. We adapt the recently proposed Fr\'{e}chet Distance metric to remote sensing images for evaluating performance of the generator, demonstrate the potential of DGMs for conditional denoising, and discuss future directions as well as the limitations of DGMs in Earth science and humanitarian applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge