Michael Cochez
Inductive Entity Representations from Text via Link Prediction
Oct 07, 2020
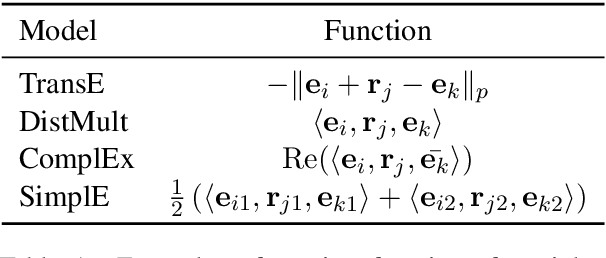
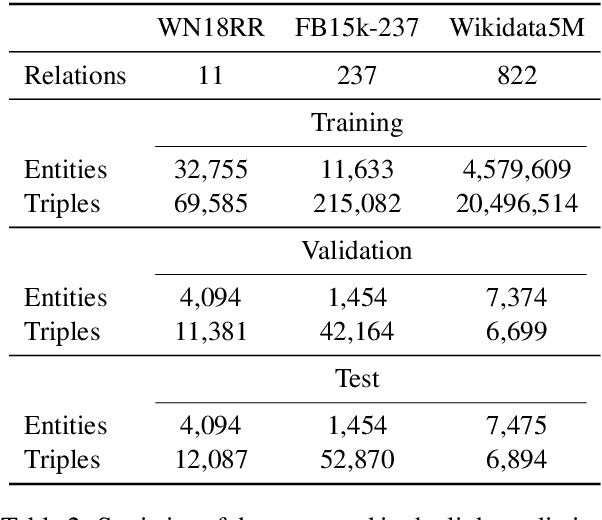
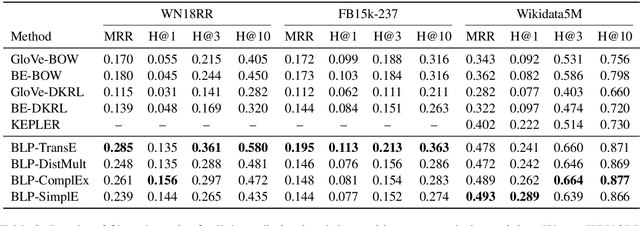
Abstract:We present a method for learning representations of entities, that uses a Transformer-based architecture as an entity encoder, and link prediction training on a knowledge graph with textual entity descriptions. We demonstrate that our approach can be applied effectively for link prediction in different inductive settings involving entities not seen during training, outperforming related state-of-the-art methods (22% MRR improvement on average). We provide evidence that the learned representations transfer to other tasks that do not require fine-tuning the entity encoder. In an entity classification task we obtain an average improvement of 16% accuracy compared with baselines that also employ pre-trained models. For an information retrieval task, significant improvements of up to 8.8% in NDCG@10 were obtained for natural language queries.
Classification Benchmarks for Under-resourced Bengali Language based on Multichannel Convolutional-LSTM Network
Apr 19, 2020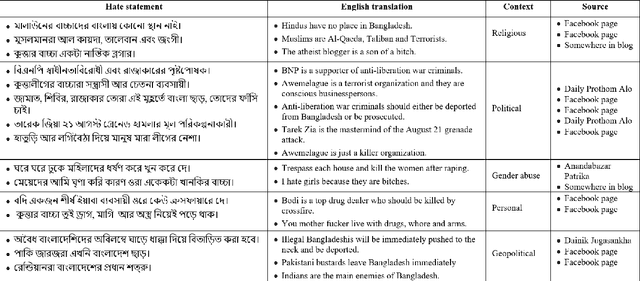
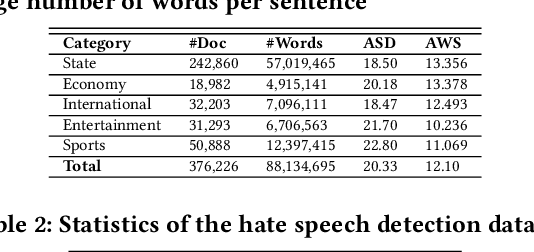
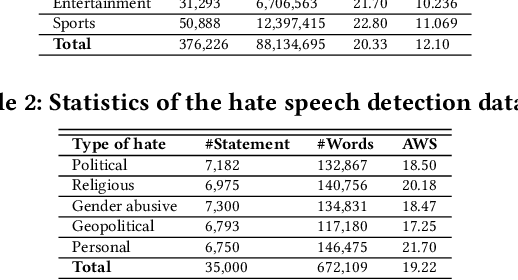
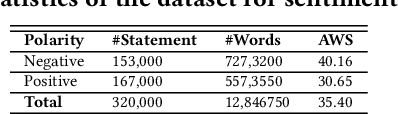
Abstract:Exponential growths of social media and micro-blogging sites not only provide platforms for empowering freedom of expressions and individual voices but also enables people to express anti-social behaviour like online harassment, cyberbullying, and hate speech. Numerous works have been proposed to utilize these data for social and anti-social behaviours analysis, document characterization, and sentiment analysis by predicting the contexts mostly for highly resourced languages such as English. However, there are languages that are under-resources, e.g., South Asian languages like Bengali, Tamil, Assamese, Telugu that lack of computational resources for the NLP tasks. In this paper, we provide several classification benchmarks for Bengali, an under-resourced language. We prepared three datasets of expressing hate, commonly used topics, and opinions for hate speech detection, document classification, and sentiment analysis, respectively. We built the largest Bengali word embedding models to date based on 250 million articles, which we call BengFastText. We perform three different experiments, covering document classification, sentiment analysis, and hate speech detection. We incorporate word embeddings into a Multichannel Convolutional-LSTM (MConv-LSTM) network for predicting different types of hate speech, document classification, and sentiment analysis. Experiments demonstrate that BengFastText can capture the semantics of words from respective contexts correctly. Evaluations against several baseline embedding models, e.g., Word2Vec and GloVe yield up to 92.30%, 82.25%, and 90.45% F1-scores in case of document classification, sentiment analysis, and hate speech detection, respectively during 5-fold cross-validation tests.
DeepCOVIDExplainer: Explainable COVID-19 Predictions Based on Chest X-ray Images
Apr 10, 2020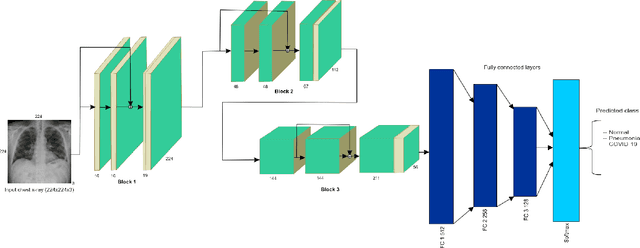
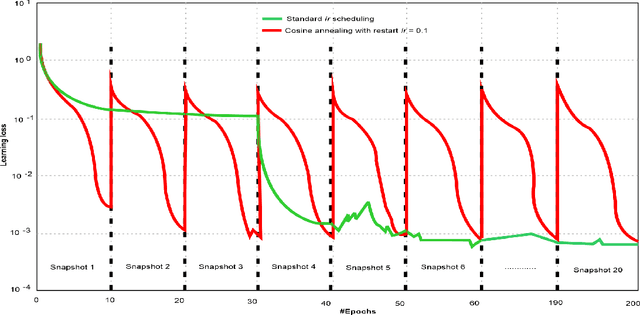
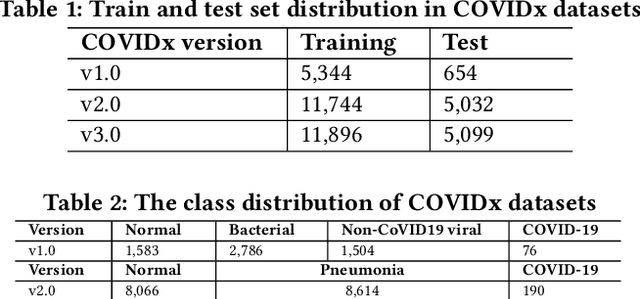
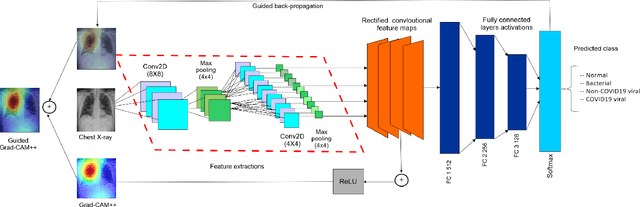
Abstract:Amid the coronavirus disease(COVID-19) pandemic, humanity experiences a rapid increase in infection numbers across the world. Challenge hospitals are faced with, in the fight against the virus, is the effective screening of incoming patients. One methodology is the assessment of chest radiography(CXR) images, which usually requires expert radiologists' knowledge. In this paper, we propose an explainable deep neural networks(DNN)-based method for automatic detection of COVID-19 symptoms from CXR images, which we call 'DeepCOVIDExplainer'. We used 16,995 CXR images across 13,808 patients, covering normal, pneumonia, and COVID-19 cases. CXR images are first comprehensively preprocessed, before being augmented and classified with a neural ensemble method, followed by highlighting class-discriminating regions using gradient-guided class activation maps(Grad-CAM++) and layer-wise relevance propagation(LRP). Further, we provide human-interpretable explanations of the predictions. Evaluation results based on hold-out data show that our approach can identify COVID-19 confidently with a positive predictive value(PPV) of 89.61% and recall of 83%, improving over recent comparable approaches. We hope that our findings will be a useful contribution to the fight against COVID-19 and, in more general, towards an increasing acceptance and adoption of AI-assisted applications in the clinical practice.
Knowledge Graphs
Mar 28, 2020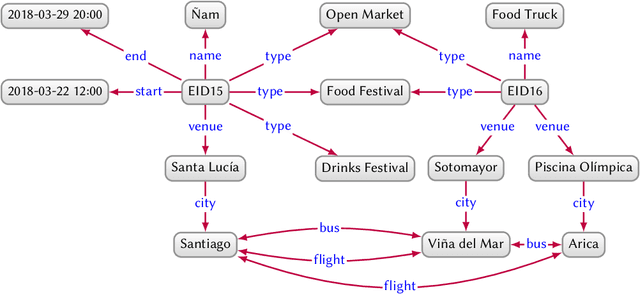
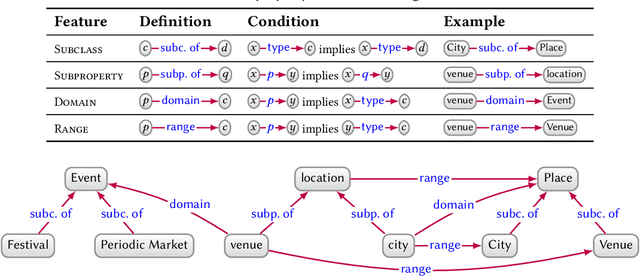
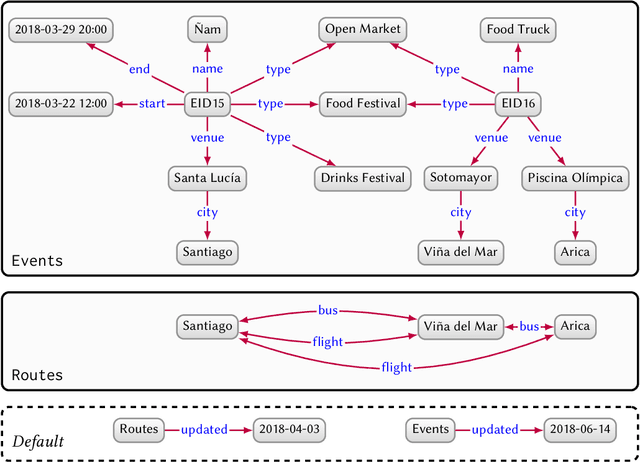
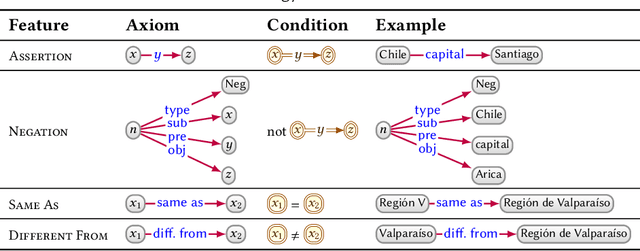
Abstract:In this paper we provide a comprehensive introduction to knowledge graphs, which have recently garnered significant attention from both industry and academia in scenarios that require exploiting diverse, dynamic, large-scale collections of data. After a general introduction, we motivate and contrast various graph-based data models and query languages that are used for knowledge graphs. We discuss the roles of schema, identity, and context in knowledge graphs. We explain how knowledge can be represented and extracted using a combination of deductive and inductive techniques. We summarise methods for the creation, enrichment, quality assessment, refinement, and publication of knowledge graphs. We provide an overview of prominent open knowledge graphs and enterprise knowledge graphs, their applications, and how they use the aforementioned techniques. We conclude with high-level future research directions for knowledge graphs.
Message Passing for Query Answering over Knowledge Graphs
Feb 06, 2020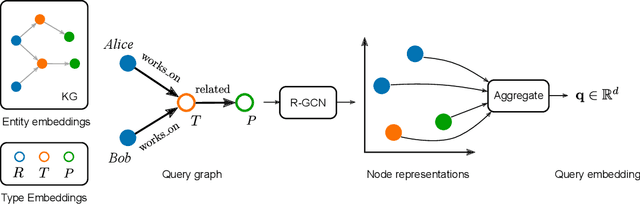
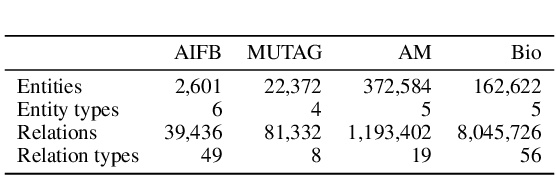
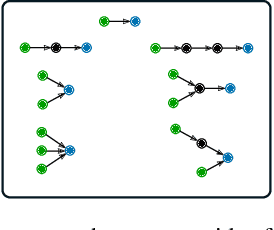
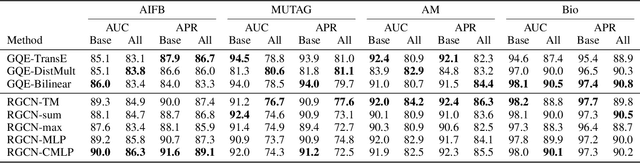
Abstract:Logic-based systems for query answering over knowledge graphs return only answers that rely on information explicitly represented in the graph. To improve recall, recent works have proposed the use of embeddings to predict additional information like missing links, or labels. These embeddings enable scoring entities in the graph as the answer a query, without being fully dependent on the graph structure. In its simplest case, answering a query in such a setting requires predicting a link between two entities. However, link prediction is not sufficient to address complex queries that involve multiple entities and variables. To solve this task, we propose to apply a message passing mechanism to a graph representation of the query, where nodes correspond to variables and entities. This results in an embedding of the query, such that answering entities are close to it in the embedding space. The general formulation of our method allows it to encode a more diverse set of query types in comparison to previous work. We evaluate our method by answering queries that rely on edges not seen during training, obtaining competitive performance. In contrast with previous work, we show that our method can generalize from training for the single-hop, link prediction task, to answering queries with more complex structures. A qualitative analysis reveals that the learned embeddings successfully capture the notion of different entity types.
Privacy Attacks on Network Embeddings
Dec 23, 2019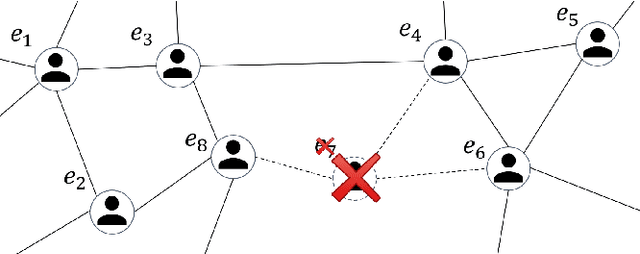
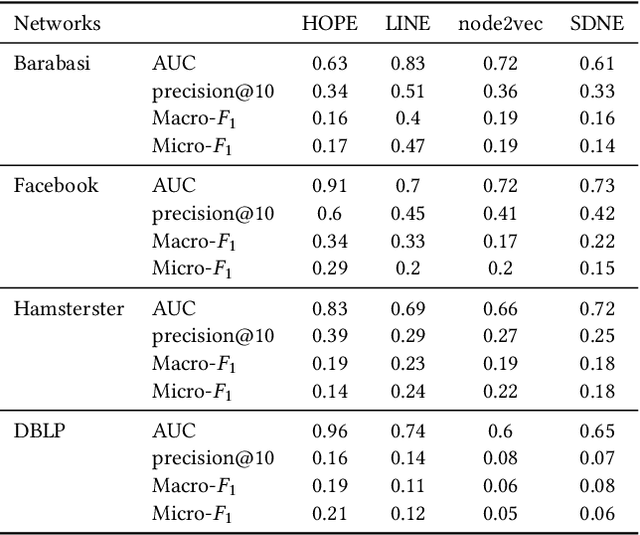
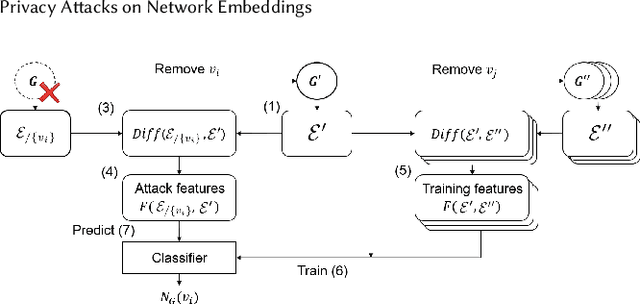
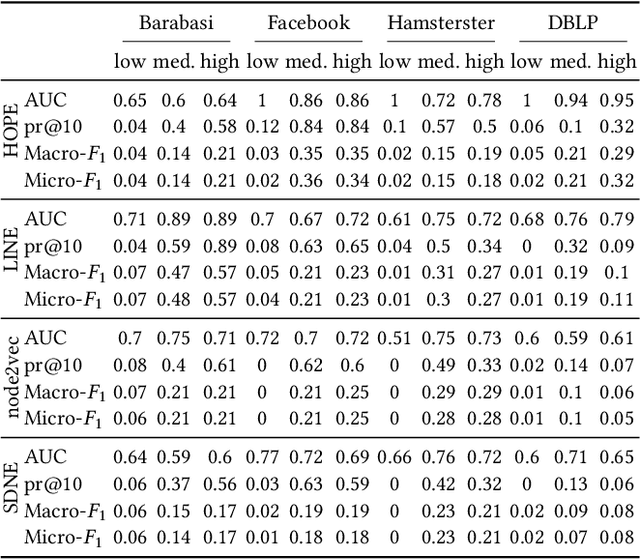
Abstract:Data ownership and data protection are increasingly important topics with ethical and legal implications, e.g., with the right to erasure established in the European General Data Protection Regulation (GDPR). In this light, we investigate network embeddings, i.e., the representation of network nodes as low-dimensional vectors. We consider a typical social network scenario with nodes representing users and edges relationships between them. We assume that a network embedding of the nodes has been trained. After that, a user demands the removal of his data, requiring the full deletion of the corresponding network information, in particular the corresponding node and incident edges. In that setting, we analyze whether after the removal of the node from the network and the deletion of the vector representation of the respective node in the embedding significant information about the link structure of the removed node is still encoded in the embedding vectors of the remaining nodes. This would require a (potentially computationally expensive) retraining of the embedding. For that purpose, we deploy an attack that leverages information from the remaining network and embedding to recover information about the neighbors of the removed node. The attack is based on (i) measuring distance changes in network embeddings and (ii) a machine learning classifier that is trained on networks that are constructed by removing additional nodes. Our experiments demonstrate that substantial information about the edges of a removed node/user can be retrieved across many different datasets. This implies that to fully protect the privacy of users, node deletion requires complete retraining - or at least a significant modification - of original network embeddings. Our results suggest that deleting the corresponding vector representation from network embeddings alone is not sufficient from a privacy perspective.
OncoNetExplainer: Explainable Predictions of Cancer Types Based on Gene Expression Data
Sep 09, 2019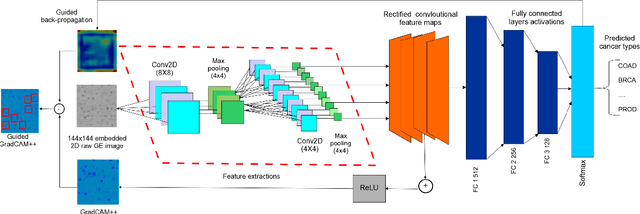
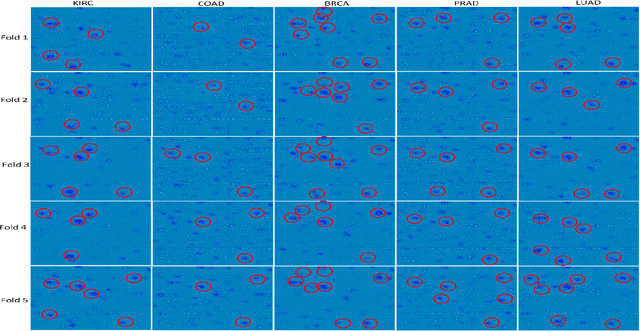
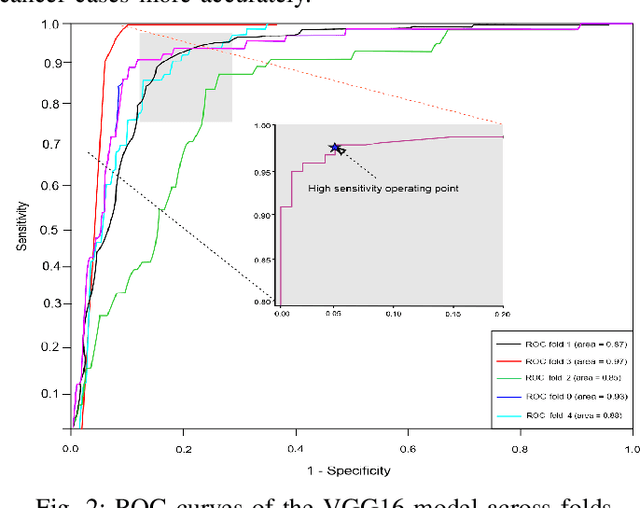

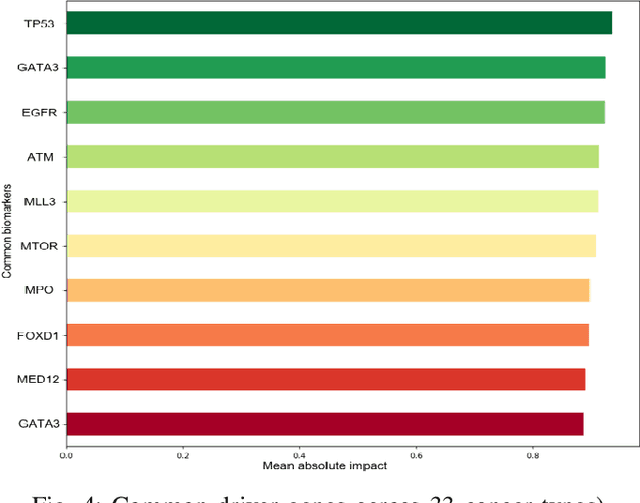
Abstract:The discovery of important biomarkers is a significant step towards understanding the molecular mechanisms of carcinogenesis; enabling accurate diagnosis for, and prognosis of, a certain cancer type. Before recommending any diagnosis, genomics data such as gene expressions(GE) and clinical outcomes need to be analyzed. However, complex nature, high dimensionality, and heterogeneity in genomics data make the overall analysis challenging. Convolutional neural networks(CNN) have shown tremendous success in solving such problems. However, neural network models are perceived mostly as `black box' methods because of their not well-understood internal functioning. However, interpretability is important to provide insights on why a given cancer case has a certain type. Besides, finding the most important biomarkers can help in recommending more accurate treatments and drug repositioning. In this paper, we propose a new approach called OncoNetExplainer to make explainable predictions of cancer types based on GE data. We used genomics data about 9,074 cancer patients covering 33 different cancer types from the Pan-Cancer Atlas on which we trained CNN and VGG16 networks using guided-gradient class activation maps++(GradCAM++). Further, we generate class-specific heat maps to identify significant biomarkers and computed feature importance in terms of mean absolute impact to rank top genes across all the cancer types. Quantitative and qualitative analyses show that both models exhibit high confidence at predicting the cancer types correctly giving an average precision of 96.25%. To provide comparisons with the baselines, we identified top genes, and cancer-specific driver genes using gradient boosted trees and SHapley Additive exPlanations(SHAP). Finally, our findings were validated with the annotations provided by the TumorPortal.
* In proc. of 19th IEEE International Conference on Bioinformatics and Bioengineering(IEEE BIBE 2019)
Structured Query Construction via Knowledge Graph Embedding
Sep 06, 2019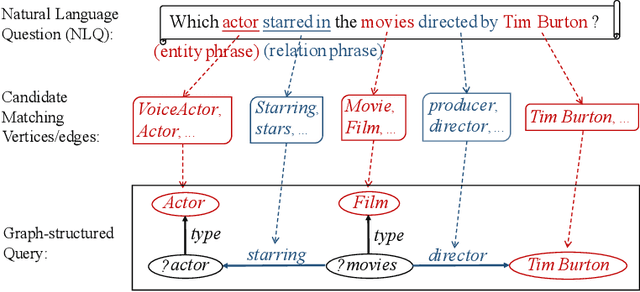
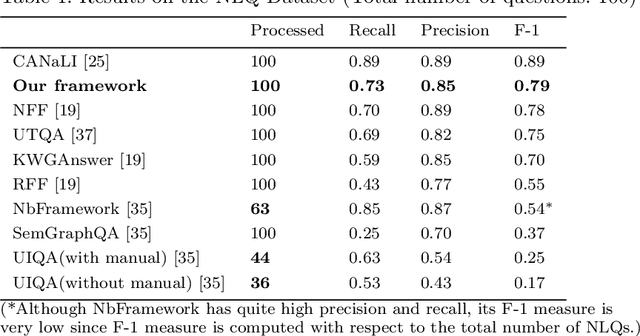
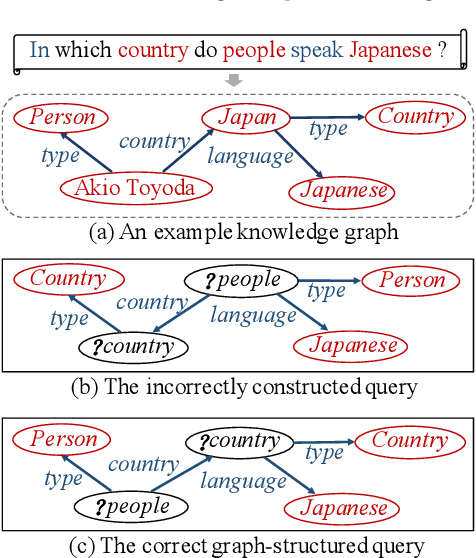
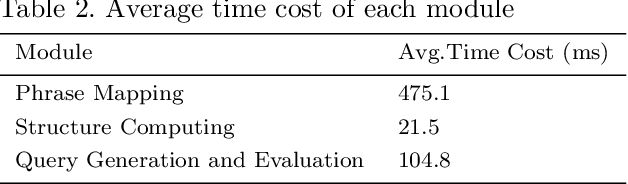
Abstract:In order to facilitate the accesses of general users to knowledge graphs, an increasing effort is being exerted to construct graph-structured queries of given natural language questions. At the core of the construction is to deduce the structure of the target query and determine the vertices/edges which constitute the query. Existing query construction methods rely on question understanding and conventional graph-based algorithms which lead to inefficient and degraded performances facing complex natural language questions over knowledge graphs with large scales. In this paper, we focus on this problem and propose a novel framework standing on recent knowledge graph embedding techniques. Our framework first encodes the underlying knowledge graph into a low-dimensional embedding space by leveraging generalized local knowledge graphs. Given a natural language question, the learned embedding representations of the knowledge graph are utilized to compute the query structure and assemble vertices/edges into the target query. Extensive experiments were conducted on the benchmark dataset, and the results demonstrate that our framework outperforms state-of-the-art baseline models regarding effectiveness and efficiency.
Message Passing for Complex Question Answering over Knowledge Graphs
Aug 19, 2019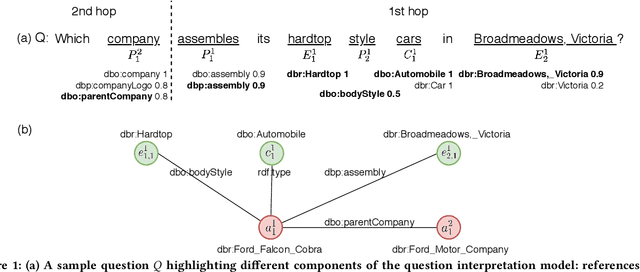
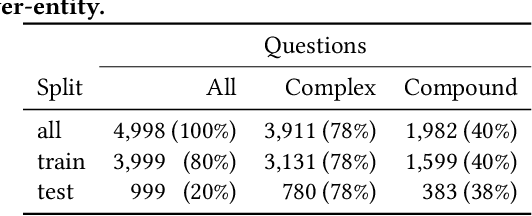
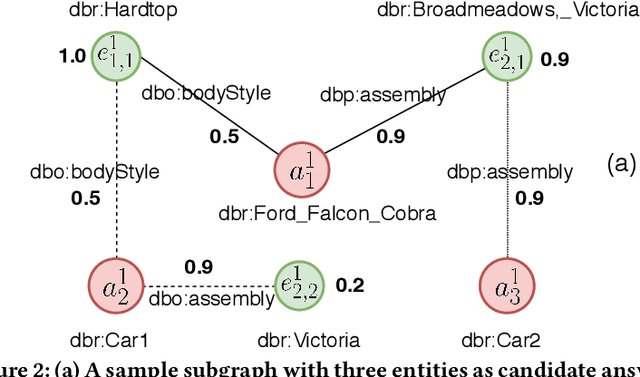
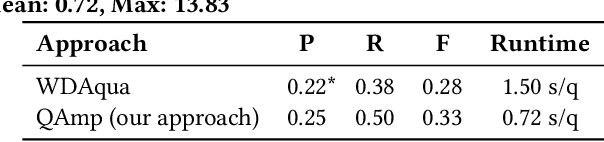
Abstract:Question answering over knowledge graphs (KGQA) has evolved from simple single-fact questions to complex questions that require graph traversal and aggregation. We propose a novel approach for complex KGQA that uses unsupervised message passing, which propagates confidence scores obtained by parsing an input question and matching terms in the knowledge graph to a set of possible answers. First, we identify entity, relationship, and class names mentioned in a natural language question, and map these to their counterparts in the graph. Then, the confidence scores of these mappings propagate through the graph structure to locate the answer entities. Finally, these are aggregated depending on the identified question type. This approach can be efficiently implemented as a series of sparse matrix multiplications mimicking joins over small local subgraphs. Our evaluation results show that the proposed approach outperforms the state-of-the-art on the LC-QuAD benchmark. Moreover, we show that the performance of the approach depends only on the quality of the question interpretation results, i.e., given a correct relevance score distribution, our approach always produces a correct answer ranking. Our error analysis reveals correct answers missing from the benchmark dataset and inconsistencies in the DBpedia knowledge graph. Finally, we provide a comprehensive evaluation of the proposed approach accompanied with an ablation study and an error analysis, which showcase the pitfalls for each of the question answering components in more detail.
Transferring knowledge from monitored to unmonitored areas for forecasting parking spaces
Aug 07, 2019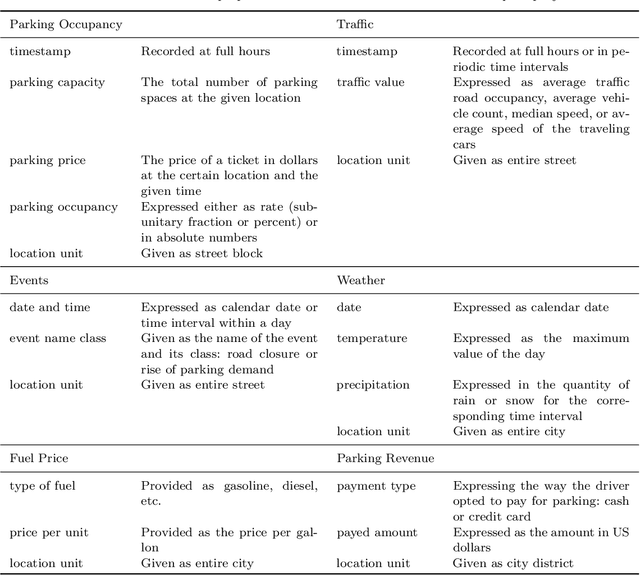
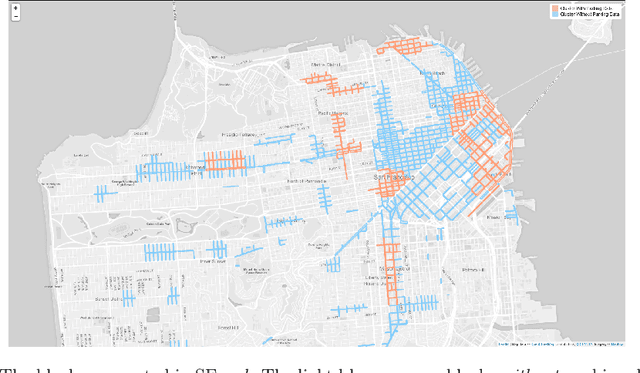
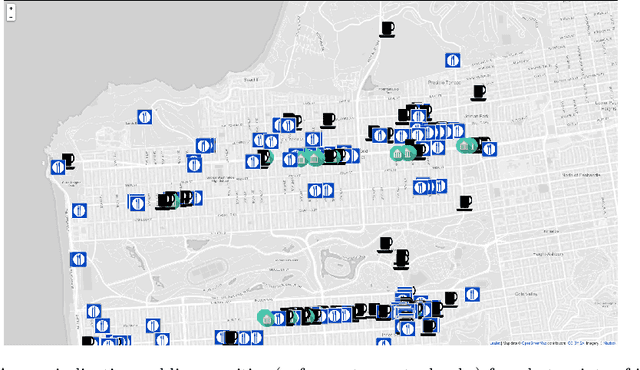
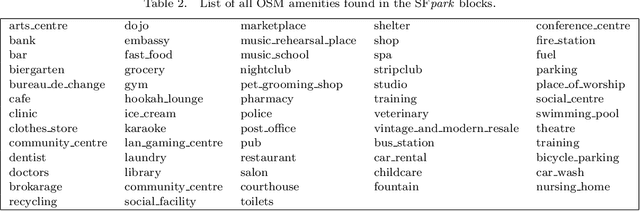
Abstract:Smart cities around the world have begun monitoring parking areas in order to estimate available parking spots and help drivers looking for parking. The current results are promising, indeed. However, existing approaches are limited by the high cost of sensors that need to be installed throughout the city in order to achieve an accurate estimation. This work investigates the extension of estimating parking information from areas equipped with sensors to areas where they are missing. To this end, the similarity between city neighborhoods is determined based on background data, i.e., from geographic information systems. Using the derived similarity values, we analyze the adaptation of occupancy rates from monitored- to unmonitored parking areas.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge