Junhan Chang
Safe-SDL:Establishing Safety Boundaries and Control Mechanisms for AI-Driven Self-Driving Laboratories
Feb 13, 2026Abstract:The emergence of Self-Driving Laboratories (SDLs) transforms scientific discovery methodology by integrating AI with robotic automation to create closed-loop experimental systems capable of autonomous hypothesis generation, experimentation, and analysis. While promising to compress research timelines from years to weeks, their deployment introduces unprecedented safety challenges differing from traditional laboratories or purely digital AI. This paper presents Safe-SDL, a comprehensive framework for establishing robust safety boundaries and control mechanisms in AI-driven autonomous laboratories. We identify and analyze the critical ``Syntax-to-Safety Gap'' -- the disconnect between AI-generated syntactically correct commands and their physical safety implications -- as the central challenge in SDL deployment. Our framework addresses this gap through three synergistic components: (1) formally defined Operational Design Domains (ODDs) that constrain system behavior within mathematically verified boundaries, (2) Control Barrier Functions (CBFs) that provide real-time safety guarantees through continuous state-space monitoring, and (3) a novel Transactional Safety Protocol (CRUTD) that ensures atomic consistency between digital planning and physical execution. We ground our theoretical contributions through analysis of existing implementations including UniLabOS and the Osprey architecture, demonstrating how these systems instantiate key safety principles. Evaluation against the LabSafety Bench reveals that current foundation models exhibit significant safety failures, demonstrating that architectural safety mechanisms are essential rather than optional. Our framework provides both theoretical foundations and practical implementation guidance for safe deployment of autonomous scientific systems, establishing the groundwork for responsible acceleration of AI-driven discovery.
Bohrium + SciMaster: Building the Infrastructure and Ecosystem for Agentic Science at Scale
Dec 23, 2025



Abstract:AI agents are emerging as a practical way to run multi-step scientific workflows that interleave reasoning with tool use and verification, pointing to a shift from isolated AI-assisted steps toward \emph{agentic science at scale}. This shift is increasingly feasible, as scientific tools and models can be invoked through stable interfaces and verified with recorded execution traces, and increasingly necessary, as AI accelerates scientific output and stresses the peer-review and publication pipeline, raising the bar for traceability and credible evaluation. However, scaling agentic science remains difficult: workflows are hard to observe and reproduce; many tools and laboratory systems are not agent-ready; execution is hard to trace and govern; and prototype AI Scientist systems are often bespoke, limiting reuse and systematic improvement from real workflow signals. We argue that scaling agentic science requires an infrastructure-and-ecosystem approach, instantiated in Bohrium+SciMaster. Bohrium acts as a managed, traceable hub for AI4S assets -- akin to a HuggingFace of AI for Science -- that turns diverse scientific data, software, compute, and laboratory systems into agent-ready capabilities. SciMaster orchestrates these capabilities into long-horizon scientific workflows, on which scientific agents can be composed and executed. Between infrastructure and orchestration, a \emph{scientific intelligence substrate} organizes reusable models, knowledge, and components into executable building blocks for workflow reasoning and action, enabling composition, auditability, and improvement through use. We demonstrate this stack with eleven representative master agents in real workflows, achieving orders-of-magnitude reductions in end-to-end scientific cycle time and generating execution-grounded signals from real workloads at multi-million scale.
Uni-SMART: Universal Science Multimodal Analysis and Research Transformer
Mar 15, 2024



Abstract:In scientific research and its application, scientific literature analysis is crucial as it allows researchers to build on the work of others. However, the fast growth of scientific knowledge has led to a massive increase in scholarly articles, making in-depth literature analysis increasingly challenging and time-consuming. The emergence of Large Language Models (LLMs) has offered a new way to address this challenge. Known for their strong abilities in summarizing texts, LLMs are seen as a potential tool to improve the analysis of scientific literature. However, existing LLMs have their own limits. Scientific literature often includes a wide range of multimodal elements, such as molecular structure, tables, and charts, which are hard for text-focused LLMs to understand and analyze. This issue points to the urgent need for new solutions that can fully understand and analyze multimodal content in scientific literature. To answer this demand, we present Uni-SMART (Universal Science Multimodal Analysis and Research Transformer), an innovative model designed for in-depth understanding of multimodal scientific literature. Through rigorous quantitative evaluation across several domains, Uni-SMART demonstrates superior performance over leading text-focused LLMs. Furthermore, our exploration extends to practical applications, including patent infringement detection and nuanced analysis of charts. These applications not only highlight Uni-SMART's adaptability but also its potential to revolutionize how we interact with scientific literature.
SciAssess: Benchmarking LLM Proficiency in Scientific Literature Analysis
Mar 15, 2024



Abstract:Recent breakthroughs in Large Language Models (LLMs) have revolutionized natural language understanding and generation, igniting a surge of interest in leveraging these technologies in the field of scientific literature analysis. Existing benchmarks, however, inadequately evaluate the proficiency of LLMs in scientific literature analysis, especially in scenarios involving complex comprehension and multimodal data. In response, we introduced SciAssess, a benchmark tailored for the in-depth analysis of scientific literature, crafted to provide a thorough assessment of LLMs' efficacy. SciAssess focuses on evaluating LLMs' abilities in memorization, comprehension, and analysis within the context of scientific literature analysis. It includes representative tasks from diverse scientific fields, such as general chemistry, organic materials, and alloy materials. And rigorous quality control measures ensure its reliability in terms of correctness, anonymization, and copyright compliance. SciAssess evaluates leading LLMs, including GPT-4, GPT-3.5, and Gemini, identifying their strengths and aspects for improvement and supporting the ongoing development of LLM applications in scientific literature analysis. SciAssess and its resources are made available at https://sci-assess.github.io, offering a valuable tool for advancing LLM capabilities in scientific literature analysis.
A Perspective on Deep Learning for Molecular Modeling and Simulations
Apr 25, 2020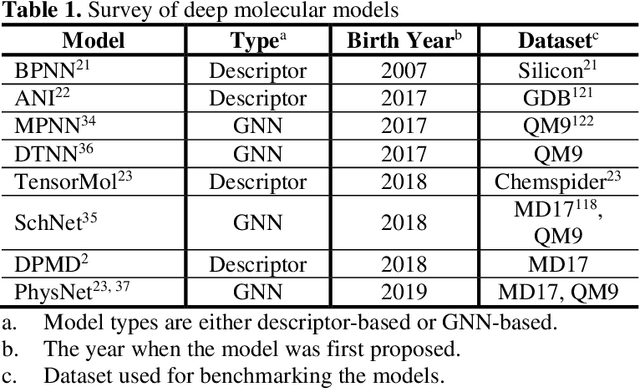
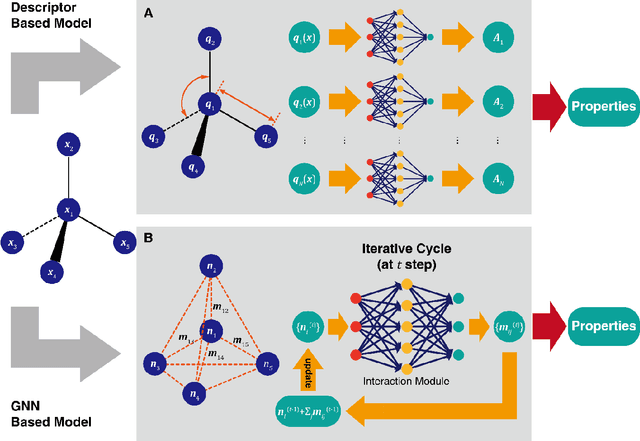
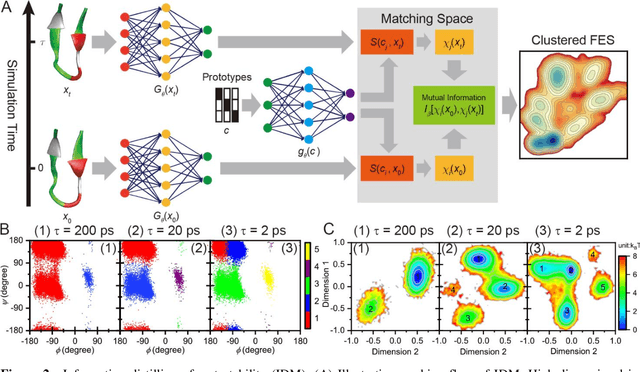
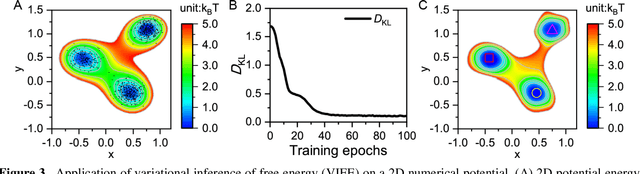
Abstract:Deep learning is transforming many areas in science, and it has great potential in modeling molecular systems. However, unlike the mature deployment of deep learning in computer vision and natural language processing, its development in molecular modeling and simulations is still at an early stage, largely because the inductive biases of molecules are completely different from those of images or texts. Footed on these differences, we first reviewed the limitations of traditional deep learning models from the perspective of molecular physics, and wrapped up some relevant technical advancement at the interface between molecular modeling and deep learning. We do not focus merely on the ever more complex neural network models, instead, we emphasize the theories and ideas behind modern deep learning. We hope that transacting these ideas into molecular modeling will create new opportunities. For this purpose, we summarized several representative applications, ranging from supervised to unsupervised and reinforcement learning, and discussed their connections with the emerging trends in deep learning. Finally, we outlook promising directions which may help address the existing issues in the current framework of deep molecular modeling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge