Jayashree Kalpathy-Cramer
The Massachusetts General Hospital, USA and University of Colorado, USA
QU-BraTS: MICCAI BraTS 2020 Challenge on Quantifying Uncertainty in Brain Tumor Segmentation -- Analysis of Ranking Metrics and Benchmarking Results
Dec 19, 2021
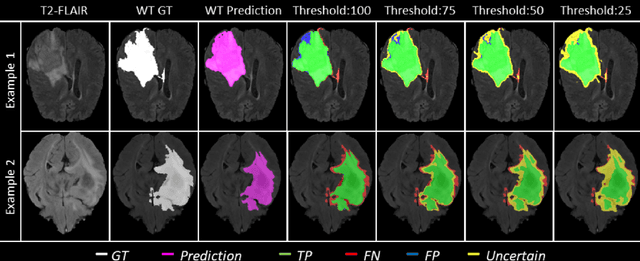
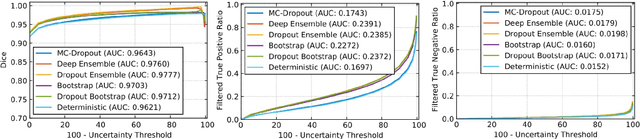
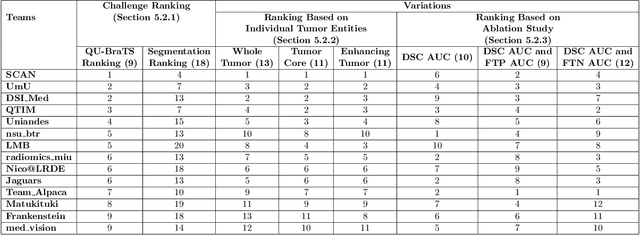
Abstract:Deep learning (DL) models have provided the state-of-the-art performance in a wide variety of medical imaging benchmarking challenges, including the Brain Tumor Segmentation (BraTS) challenges. However, the task of focal pathology multi-compartment segmentation (e.g., tumor and lesion sub-regions) is particularly challenging, and potential errors hinder the translation of DL models into clinical workflows. Quantifying the reliability of DL model predictions in the form of uncertainties, could enable clinical review of the most uncertain regions, thereby building trust and paving the way towards clinical translation. Recently, a number of uncertainty estimation methods have been introduced for DL medical image segmentation tasks. Developing metrics to evaluate and compare the performance of uncertainty measures will assist the end-user in making more informed decisions. In this study, we explore and evaluate a metric developed during the BraTS 2019-2020 task on uncertainty quantification (QU-BraTS), and designed to assess and rank uncertainty estimates for brain tumor multi-compartment segmentation. This metric (1) rewards uncertainty estimates that produce high confidence in correct assertions, and those that assign low confidence levels at incorrect assertions, and (2) penalizes uncertainty measures that lead to a higher percentages of under-confident correct assertions. We further benchmark the segmentation uncertainties generated by 14 independent participating teams of QU-BraTS 2020, all of which also participated in the main BraTS segmentation task. Overall, our findings confirm the importance and complementary value that uncertainty estimates provide to segmentation algorithms, and hence highlight the need for uncertainty quantification in medical image analyses. Our evaluation code is made publicly available at https://github.com/RagMeh11/QU-BraTS.
Monte Carlo dropout increases model repeatability
Nov 12, 2021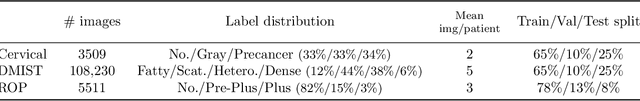
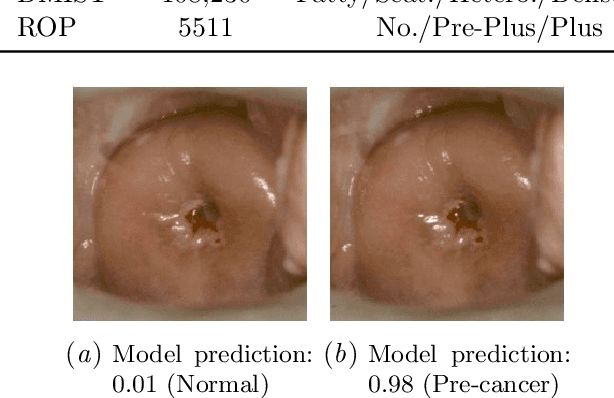
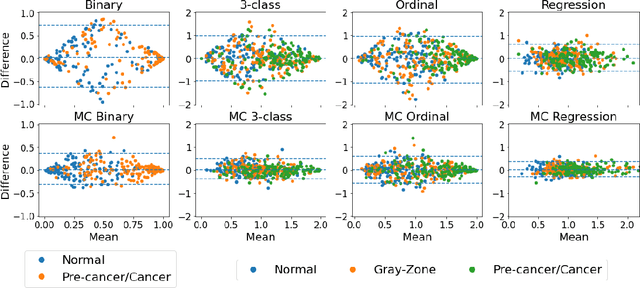
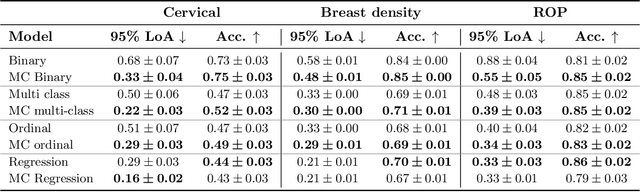
Abstract:The integration of artificial intelligence into clinical workflows requires reliable and robust models. Among the main features of robustness is repeatability. Much attention is given to classification performance without assessing the model repeatability, leading to the development of models that turn out to be unusable in practice. In this work, we evaluate the repeatability of four model types on images from the same patient that were acquired during the same visit. We study the performance of binary, multi-class, ordinal, and regression models on three medical image analysis tasks: cervical cancer screening, breast density estimation, and retinopathy of prematurity classification. Moreover, we assess the impact of sampling Monte Carlo dropout predictions at test time on classification performance and repeatability. Leveraging Monte Carlo predictions significantly increased repeatability for all tasks on the binary, multi-class, and ordinal models leading to an average reduction of the 95% limits of agreement by 17% points.
Not Color Blind: AI Predicts Racial Identity from Black and White Retinal Vessel Segmentations
Sep 28, 2021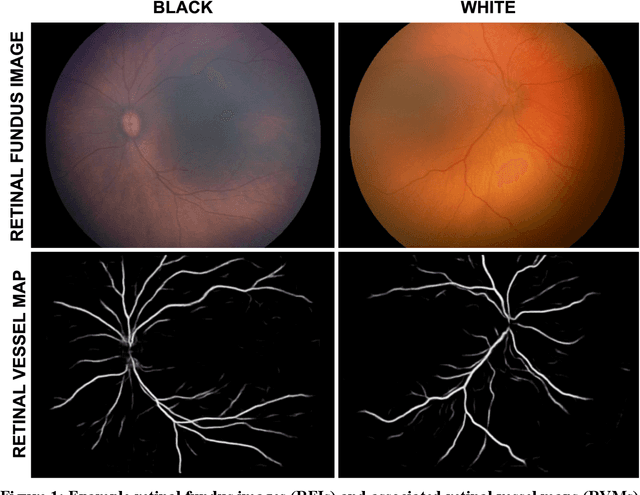
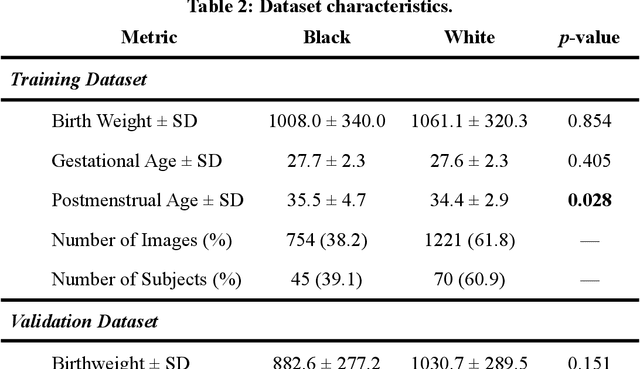
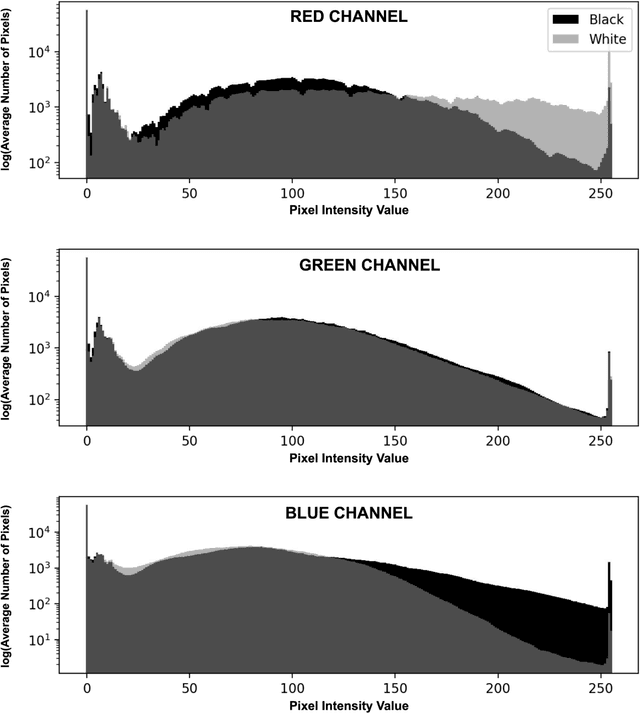
Abstract:Background: Artificial intelligence (AI) may demonstrate racial bias when skin or choroidal pigmentation is present in medical images. Recent studies have shown that convolutional neural networks (CNNs) can predict race from images that were not previously thought to contain race-specific features. We evaluate whether grayscale retinal vessel maps (RVMs) of patients screened for retinopathy of prematurity (ROP) contain race-specific features. Methods: 4095 retinal fundus images (RFIs) were collected from 245 Black and White infants. A U-Net generated RVMs from RFIs, which were subsequently thresholded, binarized, or skeletonized. To determine whether RVM differences between Black and White eyes were physiological, CNNs were trained to predict race from color RFIs, raw RVMs, and thresholded, binarized, or skeletonized RVMs. Area under the precision-recall curve (AUC-PR) was evaluated. Findings: CNNs predicted race from RFIs near perfectly (image-level AUC-PR: 0.999, subject-level AUC-PR: 1.000). Raw RVMs were almost as informative as color RFIs (image-level AUC-PR: 0.938, subject-level AUC-PR: 0.995). Ultimately, CNNs were able to detect whether RFIs or RVMs were from Black or White babies, regardless of whether images contained color, vessel segmentation brightness differences were nullified, or vessel segmentation widths were normalized. Interpretation: AI can detect race from grayscale RVMs that were not thought to contain racial information. Two potential explanations for these findings are that: retinal vessels physiologically differ between Black and White babies or the U-Net segments the retinal vasculature differently for various fundus pigmentations. Either way, the implications remain the same: AI algorithms have potential to demonstrate racial bias in practice, even when preliminary attempts to remove such information from the underlying images appear to be successful.
Deploying clinical machine learning? Consider the following...
Sep 14, 2021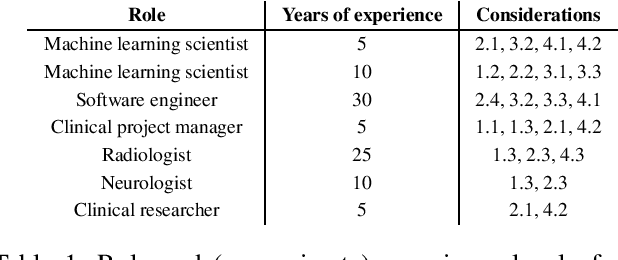
Abstract:Despite the intense attention and investment into clinical machine learning (CML) research, relatively few applications convert to clinical practice. While research is important in advancing the state-of-the-art, translation is equally important in bringing these technologies into a position to ultimately impact patient care and live up to extensive expectations surrounding AI in healthcare. To better characterize a holistic perspective among researchers and practitioners, we survey several participants with experience in developing CML for clinical deployment about their learned experiences. We collate these insights and identify several main categories of barriers and pitfalls in order to better design and develop clinical machine learning applications.
Fair Conformal Predictors for Applications in Medical Imaging
Sep 09, 2021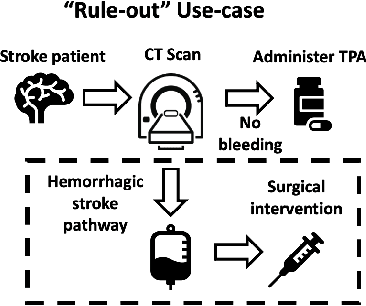
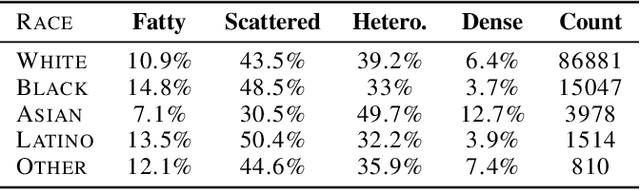
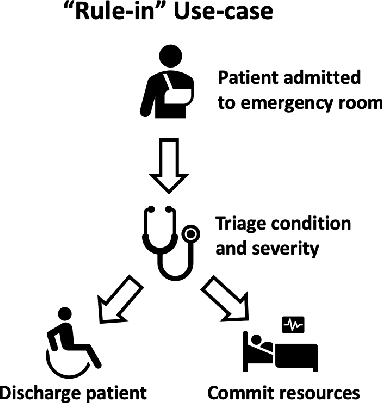
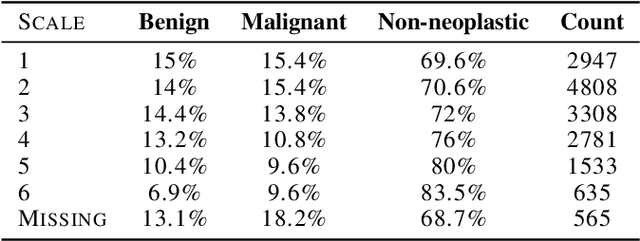
Abstract:Deep learning has the potential to augment many components of the clinical workflow, such as medical image interpretation. However, the translation of these black box algorithms into clinical practice has been marred by the relative lack of transparency compared to conventional machine learning methods, hindering in clinician trust in the systems for critical medical decision-making. Specifically, common deep learning approaches do not have intuitive ways of expressing uncertainty with respect to cases that might require further human review. Furthermore, the possibility of algorithmic bias has caused hesitancy regarding the use of developed algorithms in clinical settings. To these ends, we explore how conformal methods can complement deep learning models by providing both clinically intuitive way (by means of confidence prediction sets) of expressing model uncertainty as well as facilitating model transparency in clinical workflows. In this paper, we conduct a field survey with clinicians to assess clinical use-cases of conformal predictions. Next, we conduct experiments with a mammographic breast density and dermatology photography datasets to demonstrate the utility of conformal predictions in "rule-in" and "rule-out" disease scenarios. Further, we show that conformal predictors can be used to equalize coverage with respect to patient demographics such as race and skin tone. We find that a conformal predictions to be a promising framework with potential to increase clinical usability and transparency for better collaboration between deep learning algorithms and clinicians.
Evaluating subgroup disparity using epistemic uncertainty in mammography
Jul 15, 2021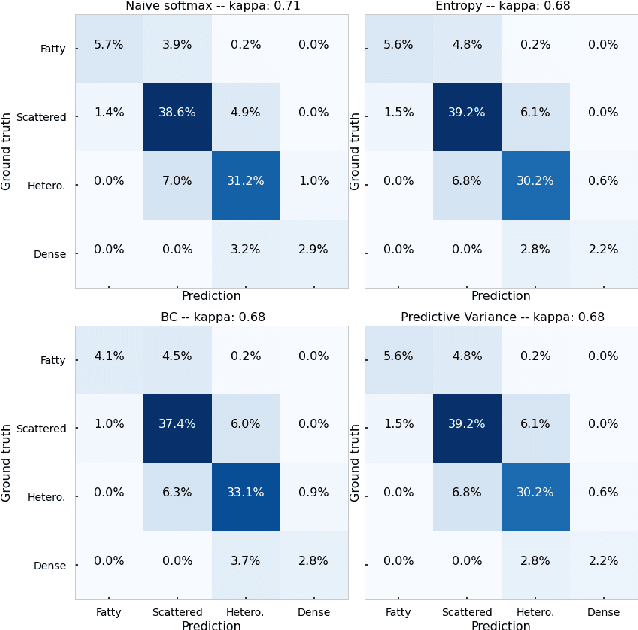
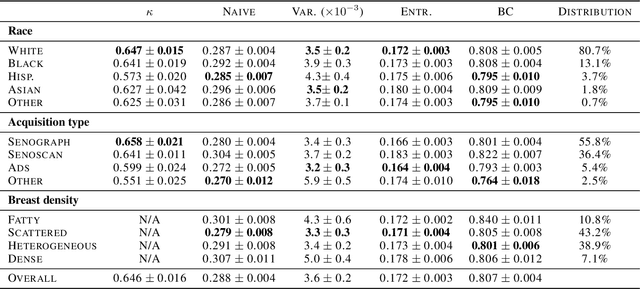
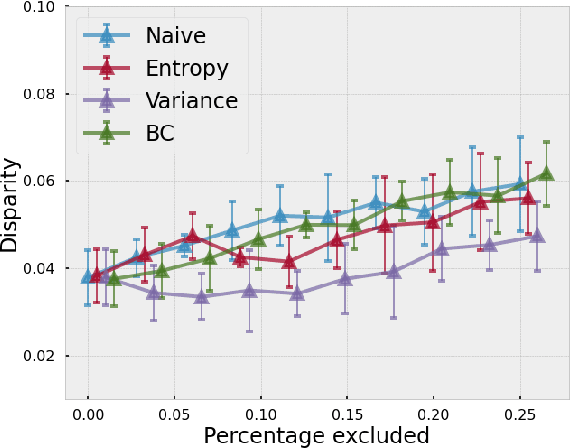
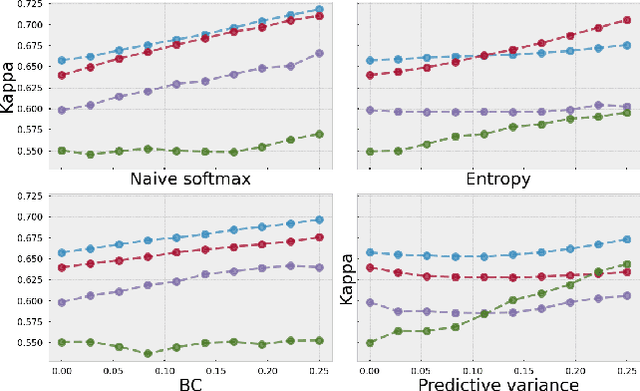
Abstract:As machine learning (ML) continue to be integrated into healthcare systems that affect clinical decision making, new strategies will need to be incorporated in order to effectively detect and evaluate subgroup disparities to ensure accountability and generalizability in clinical workflows. In this paper, we explore how epistemic uncertainty can be used to evaluate disparity in patient demographics (race) and data acquisition (scanner) subgroups for breast density assessment on a dataset of 108,190 mammograms collected from 33 clinical sites. Our results show that even if aggregate performance is comparable, the choice of uncertainty quantification metric can significantly the subgroup level. We hope this analysis can promote further work on how uncertainty can be leveraged to increase transparency of machine learning applications for clinical deployment.
SplitAVG: A heterogeneity-aware federated deep learning method for medical imaging
Jul 06, 2021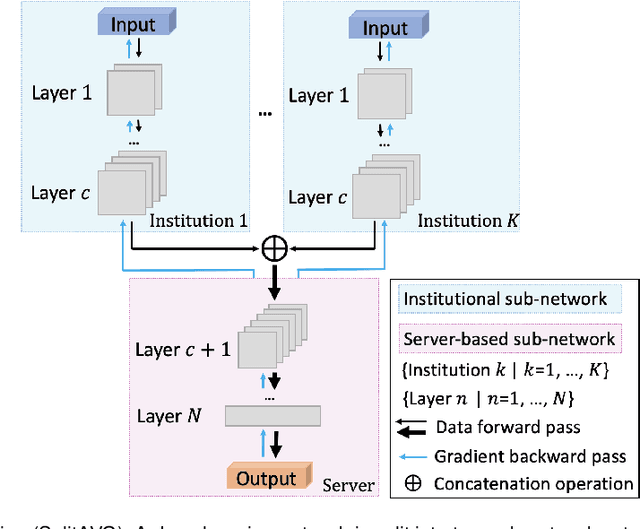
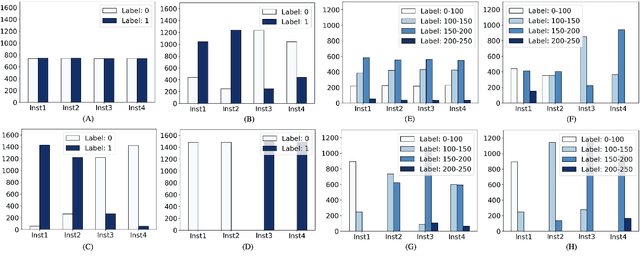
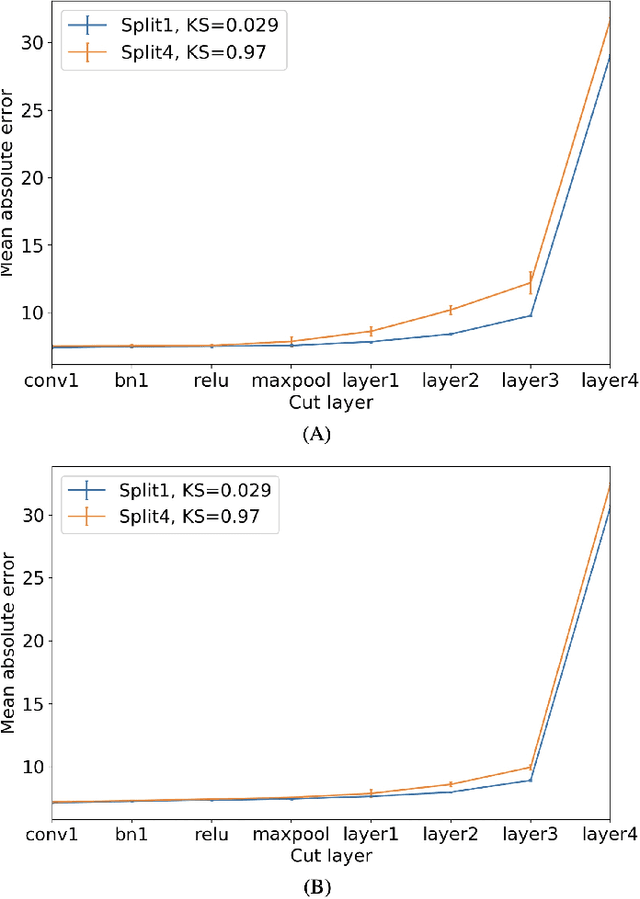
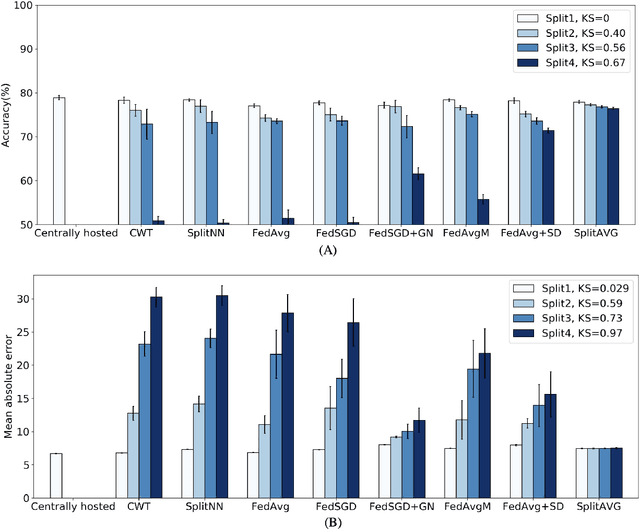
Abstract:Federated learning is an emerging research paradigm for enabling collaboratively training deep learning models without sharing patient data. However, the data from different institutions are usually heterogeneous across institutions, which may reduce the performance of models trained using federated learning. In this study, we propose a novel heterogeneity-aware federated learning method, SplitAVG, to overcome the performance drops from data heterogeneity in federated learning. Unlike previous federated methods that require complex heuristic training or hyper parameter tuning, our SplitAVG leverages the simple network split and feature map concatenation strategies to encourage the federated model training an unbiased estimator of the target data distribution. We compare SplitAVG with seven state-of-the-art federated learning methods, using centrally hosted training data as the baseline on a suite of both synthetic and real-world federated datasets. We find that the performance of models trained using all the comparison federated learning methods degraded significantly with the increasing degrees of data heterogeneity. In contrast, SplitAVG method achieves comparable results to the baseline method under all heterogeneous settings, that it achieves 96.2% of the accuracy and 110.4% of the mean absolute error obtained by the baseline in a diabetic retinopathy binary classification dataset and a bone age prediction dataset, respectively, on highly heterogeneous data partitions. We conclude that SplitAVG method can effectively overcome the performance drops from variability in data distributions across institutions. Experimental results also show that SplitAVG can be adapted to different base networks and generalized to various types of medical imaging tasks.
The RSNA-ASNR-MICCAI BraTS 2021 Benchmark on Brain Tumor Segmentation and Radiogenomic Classification
Jul 05, 2021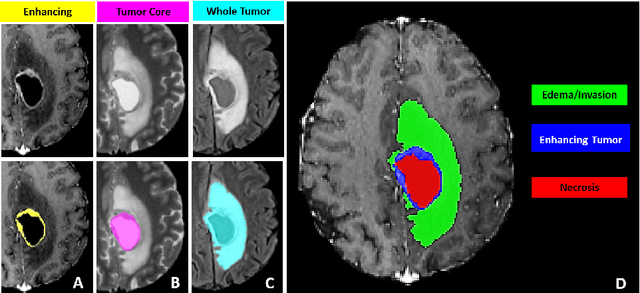
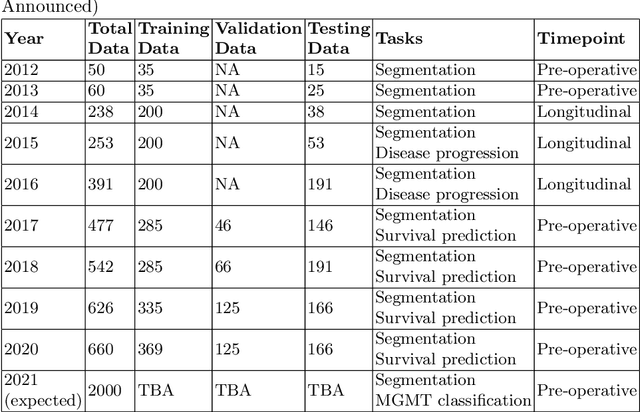
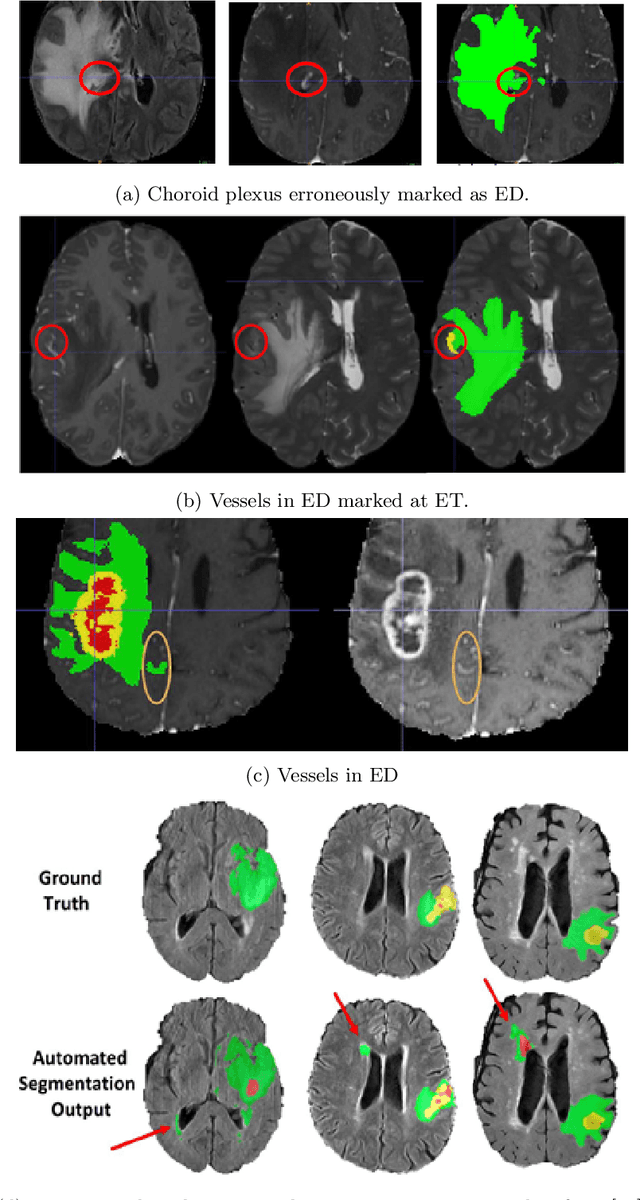
Abstract:The BraTS 2021 challenge celebrates its 10th anniversary and is jointly organized by the Radiological Society of North America (RSNA), the American Society of Neuroradiology (ASNR), and the Medical Image Computing and Computer Assisted Interventions (MICCAI) society. Since its inception, BraTS has been focusing on being a common benchmarking venue for brain glioma segmentation algorithms, with well-curated multi-institutional multi-parametric magnetic resonance imaging (mpMRI) data. Gliomas are the most common primary malignancies of the central nervous system, with varying degrees of aggressiveness and prognosis. The RSNA-ASNR-MICCAI BraTS 2021 challenge targets the evaluation of computational algorithms assessing the same tumor compartmentalization, as well as the underlying tumor's molecular characterization, in pre-operative baseline mpMRI data from 2,000 patients. Specifically, the two tasks that BraTS 2021 focuses on are: a) the segmentation of the histologically distinct brain tumor sub-regions, and b) the classification of the tumor's O[6]-methylguanine-DNA methyltransferase (MGMT) promoter methylation status. The performance evaluation of all participating algorithms in BraTS 2021 will be conducted through the Sage Bionetworks Synapse platform (Task 1) and Kaggle (Task 2), concluding in distributing to the top ranked participants monetary awards of $60,000 collectively.
Highdicom: A Python library for standardized encoding of image annotations and machine learning model outputs in pathology and radiology
Jun 14, 2021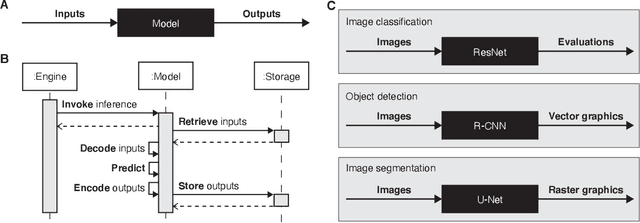
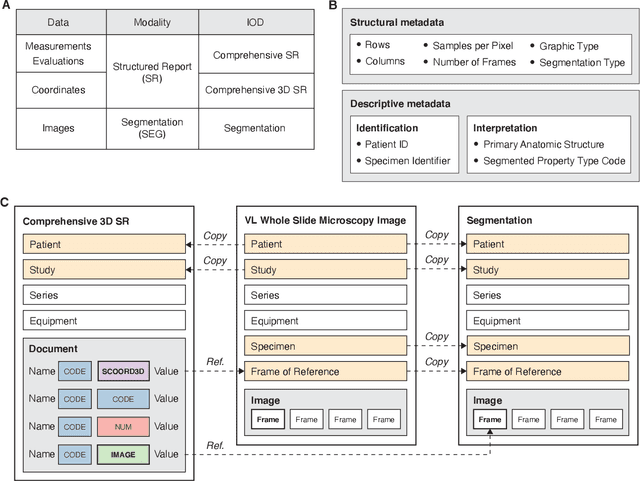
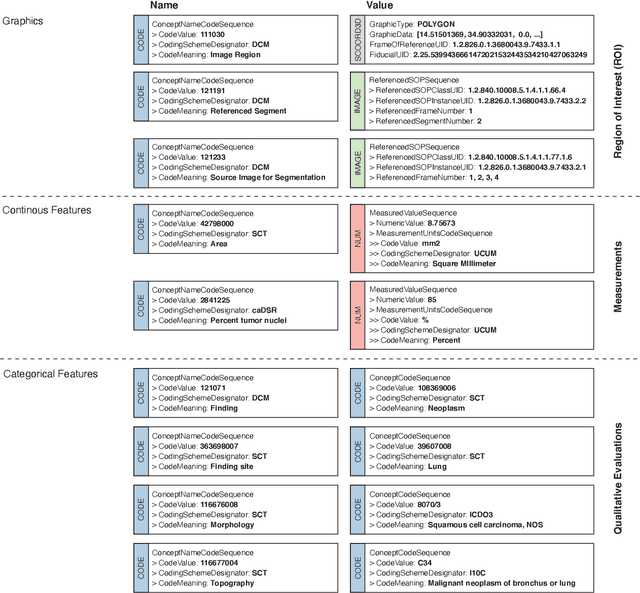
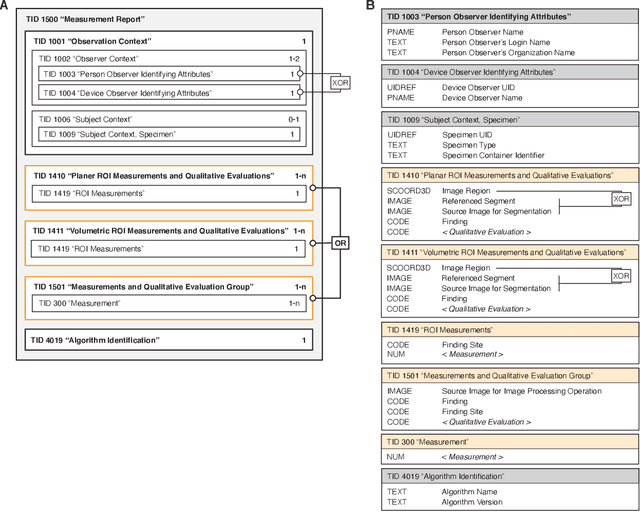
Abstract:Machine learning is revolutionizing image-based diagnostics in pathology and radiology. ML models have shown promising results in research settings, but their lack of interoperability has been a major barrier for clinical integration and evaluation. The DICOM a standard specifies Information Object Definitions and Services for the representation and communication of digital images and related information, including image-derived annotations and analysis results. However, the complexity of the standard represents an obstacle for its adoption in the ML community and creates a need for software libraries and tools that simplify working with data sets in DICOM format. Here we present the highdicom library, which provides a high-level application programming interface for the Python programming language that abstracts low-level details of the standard and enables encoding and decoding of image-derived information in DICOM format in a few lines of Python code. The highdicom library ties into the extensive Python ecosystem for image processing and machine learning. Simultaneously, by simplifying creation and parsing of DICOM-compliant files, highdicom achieves interoperability with the medical imaging systems that hold the data used to train and run ML models, and ultimately communicate and store model outputs for clinical use. We demonstrate through experiments with slide microscopy and computed tomography imaging, that, by bridging these two ecosystems, highdicom enables developers to train and evaluate state-of-the-art ML models in pathology and radiology while remaining compliant with the DICOM standard and interoperable with clinical systems at all stages. To promote standardization of ML research and streamline the ML model development and deployment process, we made the library available free and open-source.
Addressing catastrophic forgetting for medical domain expansion
Mar 24, 2021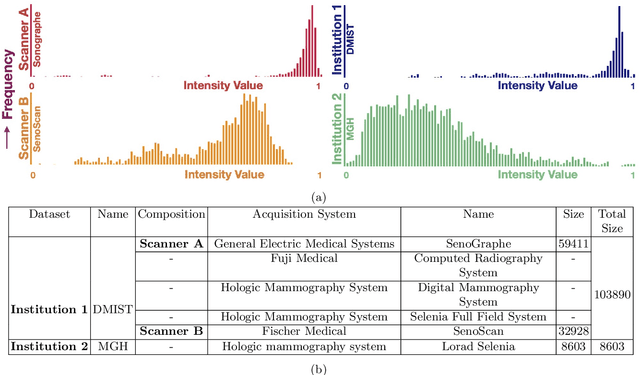
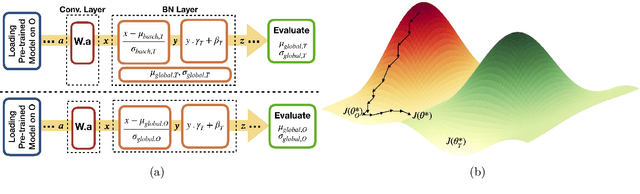
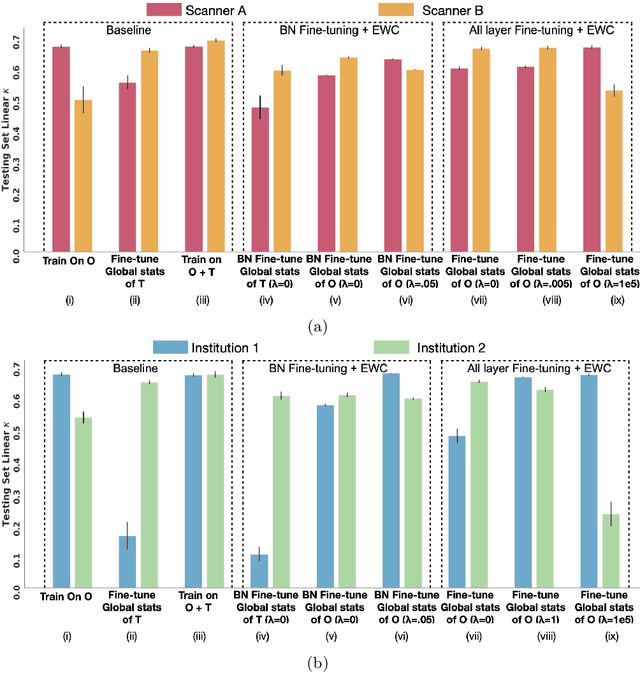
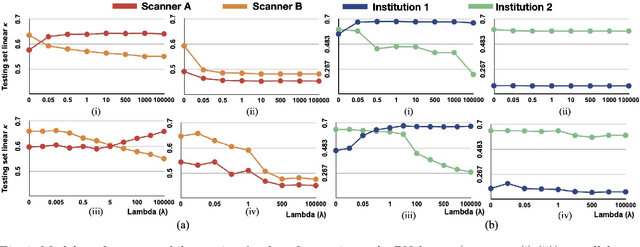
Abstract:Model brittleness is a key concern when deploying deep learning models in real-world medical settings. A model that has high performance at one institution may suffer a significant decline in performance when tested at other institutions. While pooling datasets from multiple institutions and retraining may provide a straightforward solution, it is often infeasible and may compromise patient privacy. An alternative approach is to fine-tune the model on subsequent institutions after training on the original institution. Notably, this approach degrades model performance at the original institution, a phenomenon known as catastrophic forgetting. In this paper, we develop an approach to address catastrophic forget-ting based on elastic weight consolidation combined with modulation of batch normalization statistics under two scenarios: first, for expanding the domain from one imaging system's data to another imaging system's, and second, for expanding the domain from a large multi-institutional dataset to another single institution dataset. We show that our approach outperforms several other state-of-the-art approaches and provide theoretical justification for the efficacy of batch normalization modulation. The results of this study are generally applicable to the deployment of any clinical deep learning model which requires domain expansion.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge