Praveer Singh
A generalized framework to predict continuous scores from medical ordinal labels
May 30, 2023



Abstract:Many variables of interest in clinical medicine, like disease severity, are recorded using discrete ordinal categories such as normal/mild/moderate/severe. These labels are used to train and evaluate disease severity prediction models. However, ordinal categories represent a simplification of an underlying continuous severity spectrum. Using continuous scores instead of ordinal categories is more sensitive to detecting small changes in disease severity over time. Here, we present a generalized framework that accurately predicts continuously valued variables using only discrete ordinal labels during model development. We found that for three clinical prediction tasks, models that take the ordinal relationship of the training labels into account outperformed conventional multi-class classification models. Particularly the continuous scores generated by ordinal classification and regression models showed a significantly higher correlation with expert rankings of disease severity and lower mean squared errors compared to the multi-class classification models. Furthermore, the use of MC dropout significantly improved the ability of all evaluated deep learning approaches to predict continuously valued scores that truthfully reflect the underlying continuous target variable. We showed that accurate continuously valued predictions can be generated even if the model development only involves discrete ordinal labels. The novel framework has been validated on three different clinical prediction tasks and has proven to bridge the gap between discrete ordinal labels and the underlying continuously valued variables.
Towards More Efficient Data Valuation in Healthcare Federated Learning using Ensembling
Sep 12, 2022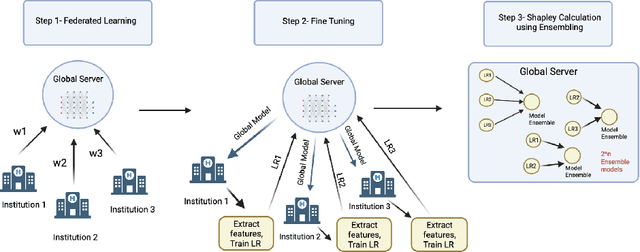
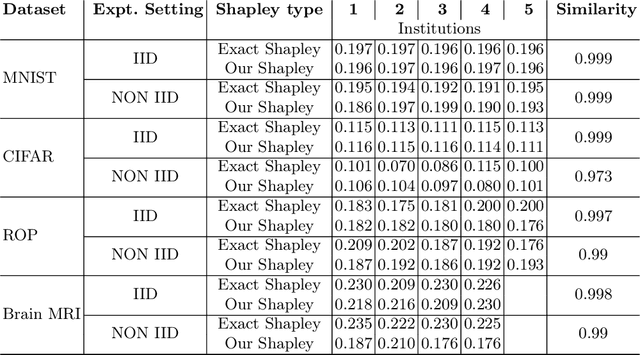
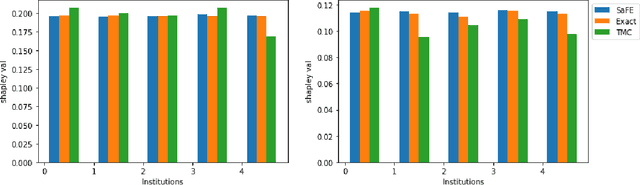
Abstract:Federated Learning (FL) wherein multiple institutions collaboratively train a machine learning model without sharing data is becoming popular. Participating institutions might not contribute equally, some contribute more data, some better quality data or some more diverse data. To fairly rank the contribution of different institutions, Shapley value (SV) has emerged as the method of choice. Exact SV computation is impossibly expensive, especially when there are hundreds of contributors. Existing SV computation techniques use approximations. However, in healthcare where the number of contributing institutions are likely not of a colossal scale, computing exact SVs is still exorbitantly expensive, but not impossible. For such settings, we propose an efficient SV computation technique called SaFE (Shapley Value for Federated Learning using Ensembling). We empirically show that SaFE computes values that are close to exact SVs, and that it performs better than current SV approximations. This is particularly relevant in medical imaging setting where widespread heterogeneity across institutions is rampant and fast accurate data valuation is required to determine the contribution of each participant in multi-institutional collaborative learning.
Estimating Test Performance for AI Medical Devices under Distribution Shift with Conformal Prediction
Jul 12, 2022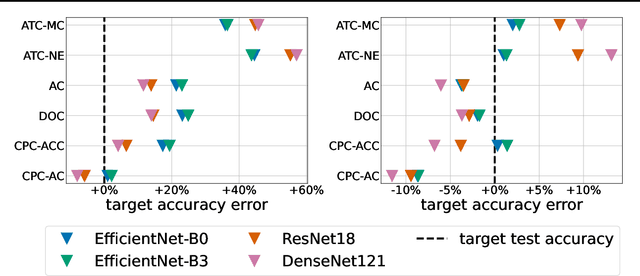
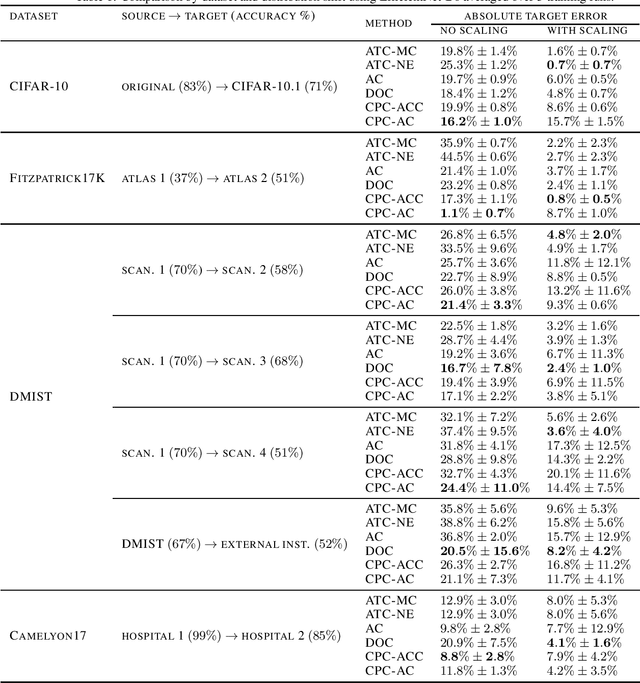
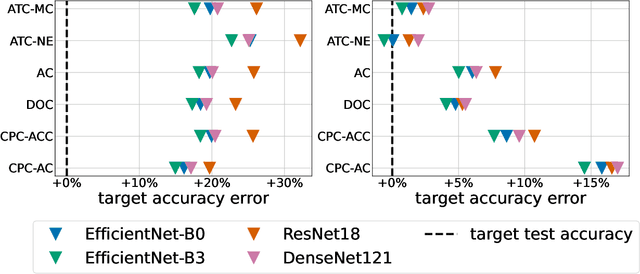
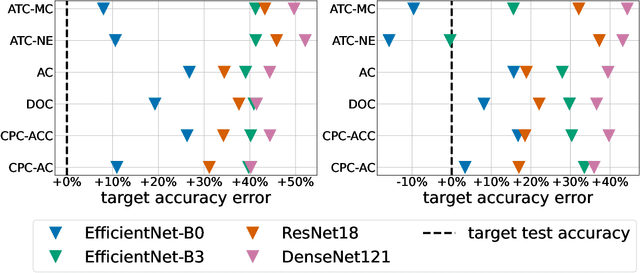
Abstract:Estimating the test performance of software AI-based medical devices under distribution shifts is crucial for evaluating the safety, efficiency, and usability prior to clinical deployment. Due to the nature of regulated medical device software and the difficulty in acquiring large amounts of labeled medical datasets, we consider the task of predicting the test accuracy of an arbitrary black-box model on an unlabeled target domain without modification to the original training process or any distributional assumptions of the original source data (i.e. we treat the model as a "black-box" and only use the predicted output responses). We propose a "black-box" test estimation technique based on conformal prediction and evaluate it against other methods on three medical imaging datasets (mammography, dermatology, and histopathology) under several clinically relevant types of distribution shift (institution, hardware scanner, atlas, hospital). We hope that by promoting practical and effective estimation techniques for black-box models, manufacturers of medical devices will develop more standardized and realistic evaluation procedures to improve the robustness and trustworthiness of clinical AI tools.
Three Applications of Conformal Prediction for Rating Breast Density in Mammography
Jun 23, 2022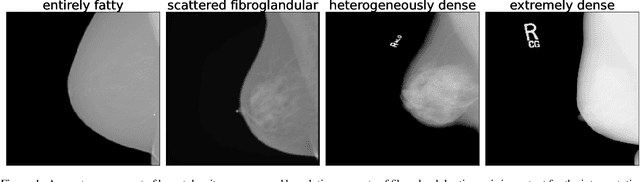

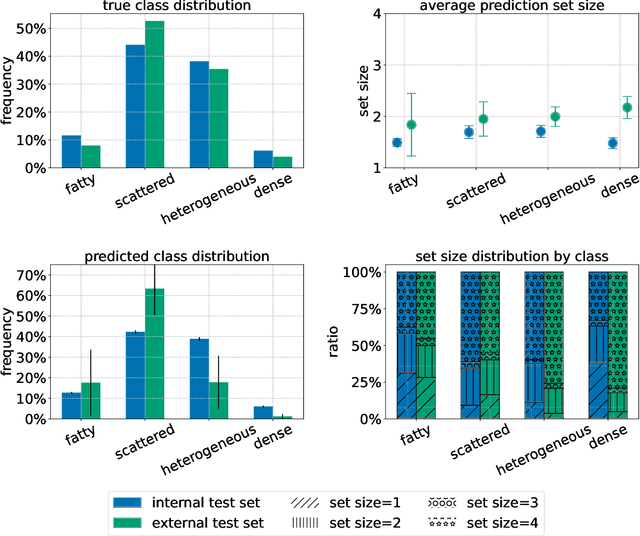
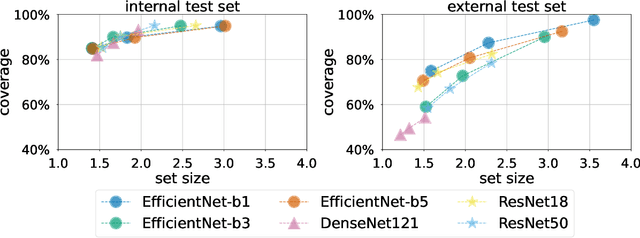
Abstract:Breast cancer is the most common cancers and early detection from mammography screening is crucial in improving patient outcomes. Assessing mammographic breast density is clinically important as the denser breasts have higher risk and are more likely to occlude tumors. Manual assessment by experts is both time-consuming and subject to inter-rater variability. As such, there has been increased interest in the development of deep learning methods for mammographic breast density assessment. Despite deep learning having demonstrated impressive performance in several prediction tasks for applications in mammography, clinical deployment of deep learning systems in still relatively rare; historically, mammography Computer-Aided Diagnoses (CAD) have over-promised and failed to deliver. This is in part due to the inability to intuitively quantify uncertainty of the algorithm for the clinician, which would greatly enhance usability. Conformal prediction is well suited to increase reliably and trust in deep learning tools but they lack realistic evaluations on medical datasets. In this paper, we present a detailed analysis of three possible applications of conformal prediction applied to medical imaging tasks: distribution shift characterization, prediction quality improvement, and subgroup fairness analysis. Our results show the potential of distribution-free uncertainty quantification techniques to enhance trust on AI algorithms and expedite their translation to usage.
QU-BraTS: MICCAI BraTS 2020 Challenge on Quantifying Uncertainty in Brain Tumor Segmentation -- Analysis of Ranking Metrics and Benchmarking Results
Dec 19, 2021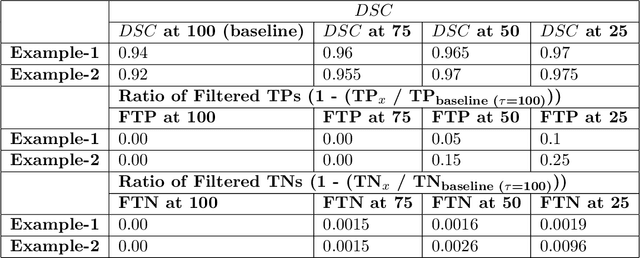
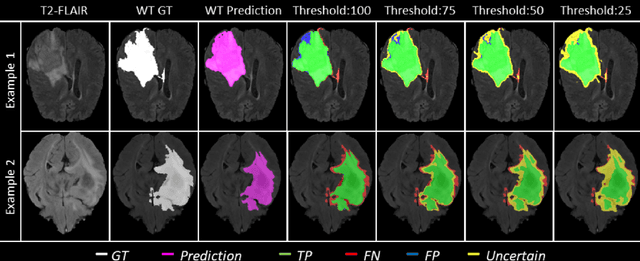
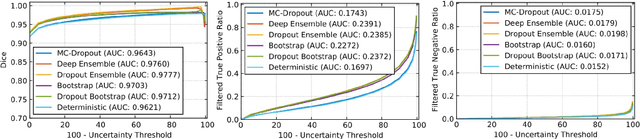
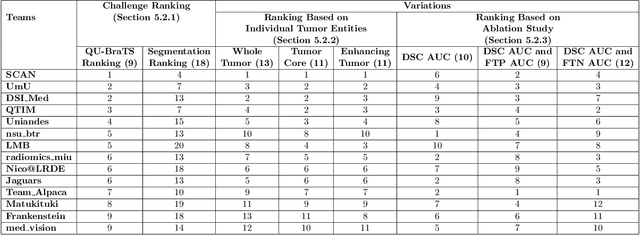
Abstract:Deep learning (DL) models have provided the state-of-the-art performance in a wide variety of medical imaging benchmarking challenges, including the Brain Tumor Segmentation (BraTS) challenges. However, the task of focal pathology multi-compartment segmentation (e.g., tumor and lesion sub-regions) is particularly challenging, and potential errors hinder the translation of DL models into clinical workflows. Quantifying the reliability of DL model predictions in the form of uncertainties, could enable clinical review of the most uncertain regions, thereby building trust and paving the way towards clinical translation. Recently, a number of uncertainty estimation methods have been introduced for DL medical image segmentation tasks. Developing metrics to evaluate and compare the performance of uncertainty measures will assist the end-user in making more informed decisions. In this study, we explore and evaluate a metric developed during the BraTS 2019-2020 task on uncertainty quantification (QU-BraTS), and designed to assess and rank uncertainty estimates for brain tumor multi-compartment segmentation. This metric (1) rewards uncertainty estimates that produce high confidence in correct assertions, and those that assign low confidence levels at incorrect assertions, and (2) penalizes uncertainty measures that lead to a higher percentages of under-confident correct assertions. We further benchmark the segmentation uncertainties generated by 14 independent participating teams of QU-BraTS 2020, all of which also participated in the main BraTS segmentation task. Overall, our findings confirm the importance and complementary value that uncertainty estimates provide to segmentation algorithms, and hence highlight the need for uncertainty quantification in medical image analyses. Our evaluation code is made publicly available at https://github.com/RagMeh11/QU-BraTS.
Not Color Blind: AI Predicts Racial Identity from Black and White Retinal Vessel Segmentations
Sep 28, 2021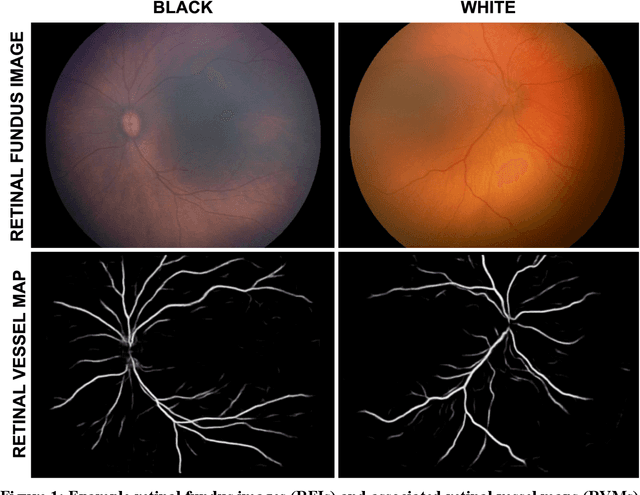
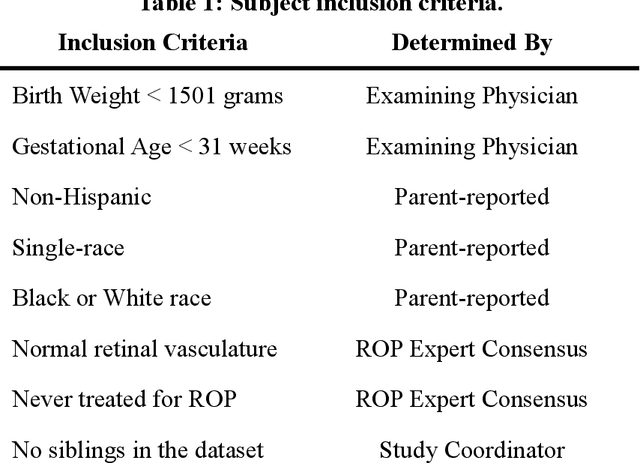
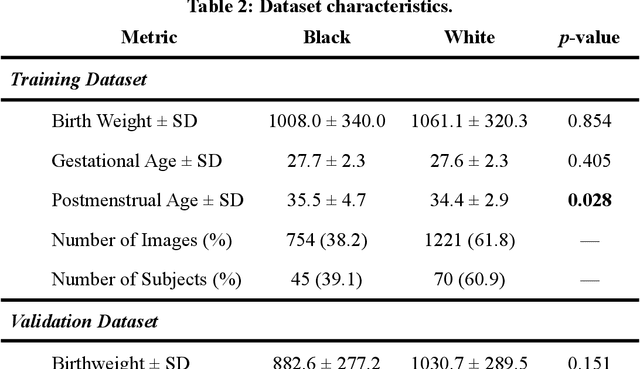
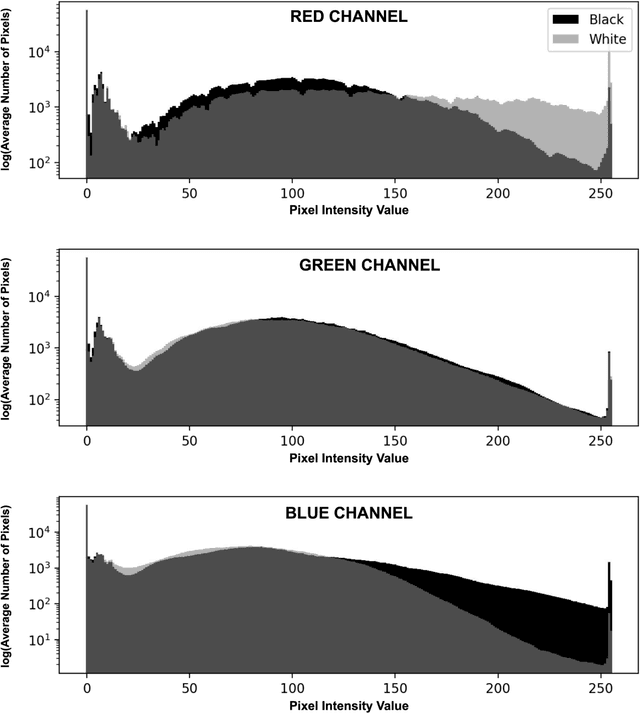
Abstract:Background: Artificial intelligence (AI) may demonstrate racial bias when skin or choroidal pigmentation is present in medical images. Recent studies have shown that convolutional neural networks (CNNs) can predict race from images that were not previously thought to contain race-specific features. We evaluate whether grayscale retinal vessel maps (RVMs) of patients screened for retinopathy of prematurity (ROP) contain race-specific features. Methods: 4095 retinal fundus images (RFIs) were collected from 245 Black and White infants. A U-Net generated RVMs from RFIs, which were subsequently thresholded, binarized, or skeletonized. To determine whether RVM differences between Black and White eyes were physiological, CNNs were trained to predict race from color RFIs, raw RVMs, and thresholded, binarized, or skeletonized RVMs. Area under the precision-recall curve (AUC-PR) was evaluated. Findings: CNNs predicted race from RFIs near perfectly (image-level AUC-PR: 0.999, subject-level AUC-PR: 1.000). Raw RVMs were almost as informative as color RFIs (image-level AUC-PR: 0.938, subject-level AUC-PR: 0.995). Ultimately, CNNs were able to detect whether RFIs or RVMs were from Black or White babies, regardless of whether images contained color, vessel segmentation brightness differences were nullified, or vessel segmentation widths were normalized. Interpretation: AI can detect race from grayscale RVMs that were not thought to contain racial information. Two potential explanations for these findings are that: retinal vessels physiologically differ between Black and White babies or the U-Net segments the retinal vasculature differently for various fundus pigmentations. Either way, the implications remain the same: AI algorithms have potential to demonstrate racial bias in practice, even when preliminary attempts to remove such information from the underlying images appear to be successful.
Deploying clinical machine learning? Consider the following...
Sep 14, 2021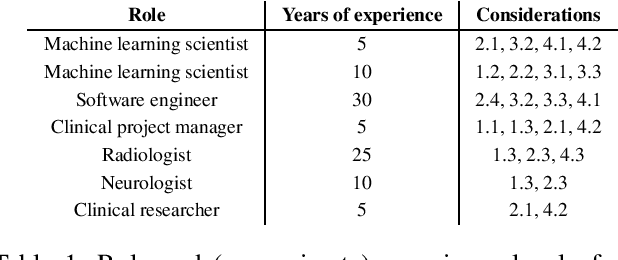
Abstract:Despite the intense attention and investment into clinical machine learning (CML) research, relatively few applications convert to clinical practice. While research is important in advancing the state-of-the-art, translation is equally important in bringing these technologies into a position to ultimately impact patient care and live up to extensive expectations surrounding AI in healthcare. To better characterize a holistic perspective among researchers and practitioners, we survey several participants with experience in developing CML for clinical deployment about their learned experiences. We collate these insights and identify several main categories of barriers and pitfalls in order to better design and develop clinical machine learning applications.
SplitAVG: A heterogeneity-aware federated deep learning method for medical imaging
Jul 06, 2021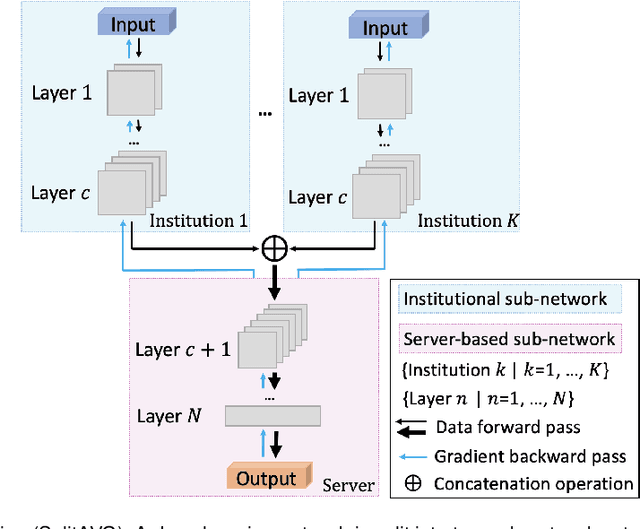
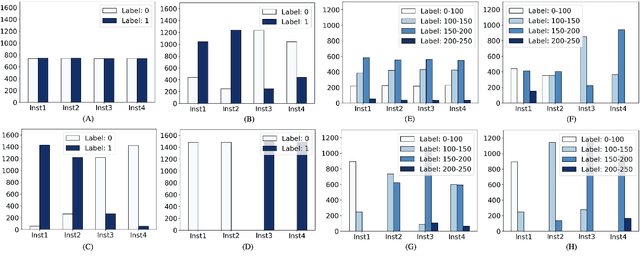
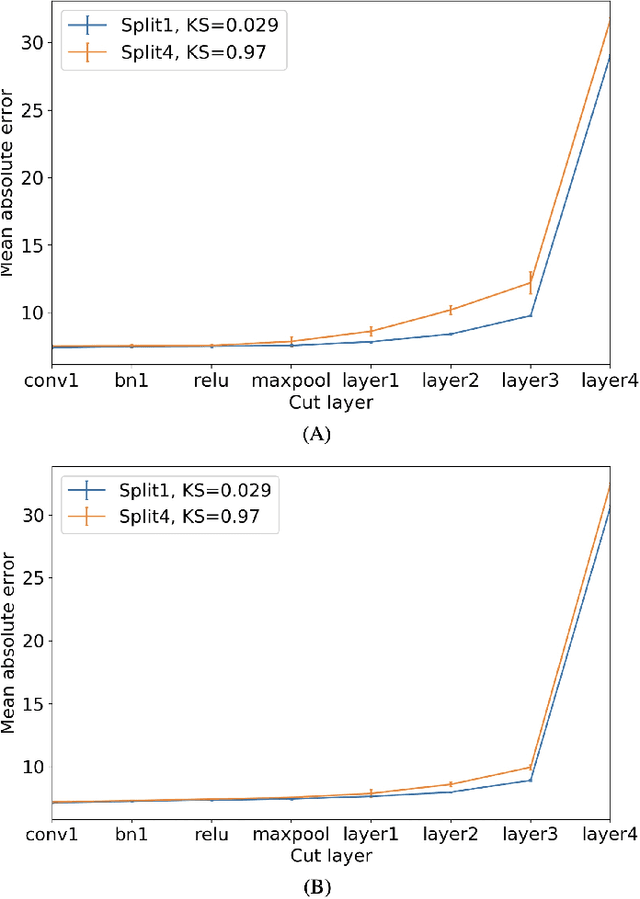
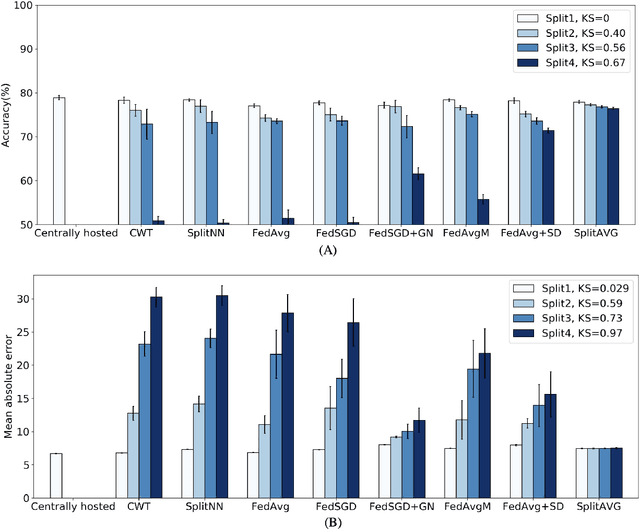
Abstract:Federated learning is an emerging research paradigm for enabling collaboratively training deep learning models without sharing patient data. However, the data from different institutions are usually heterogeneous across institutions, which may reduce the performance of models trained using federated learning. In this study, we propose a novel heterogeneity-aware federated learning method, SplitAVG, to overcome the performance drops from data heterogeneity in federated learning. Unlike previous federated methods that require complex heuristic training or hyper parameter tuning, our SplitAVG leverages the simple network split and feature map concatenation strategies to encourage the federated model training an unbiased estimator of the target data distribution. We compare SplitAVG with seven state-of-the-art federated learning methods, using centrally hosted training data as the baseline on a suite of both synthetic and real-world federated datasets. We find that the performance of models trained using all the comparison federated learning methods degraded significantly with the increasing degrees of data heterogeneity. In contrast, SplitAVG method achieves comparable results to the baseline method under all heterogeneous settings, that it achieves 96.2% of the accuracy and 110.4% of the mean absolute error obtained by the baseline in a diabetic retinopathy binary classification dataset and a bone age prediction dataset, respectively, on highly heterogeneous data partitions. We conclude that SplitAVG method can effectively overcome the performance drops from variability in data distributions across institutions. Experimental results also show that SplitAVG can be adapted to different base networks and generalized to various types of medical imaging tasks.
Addressing catastrophic forgetting for medical domain expansion
Mar 24, 2021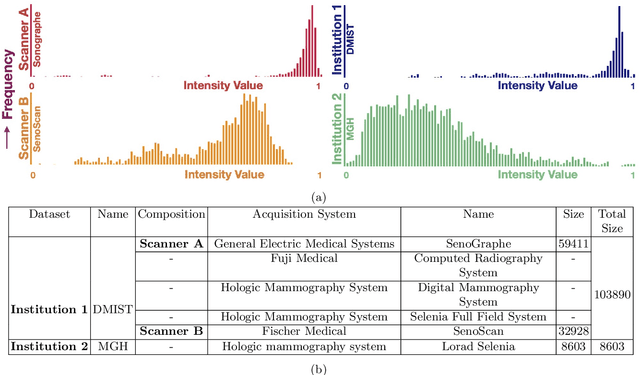
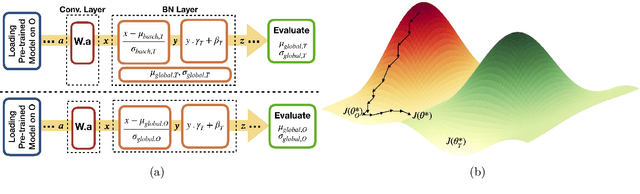
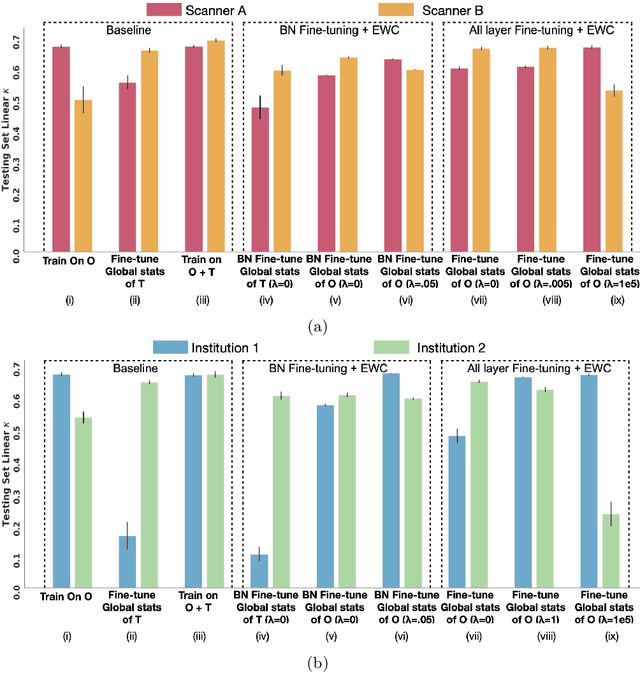
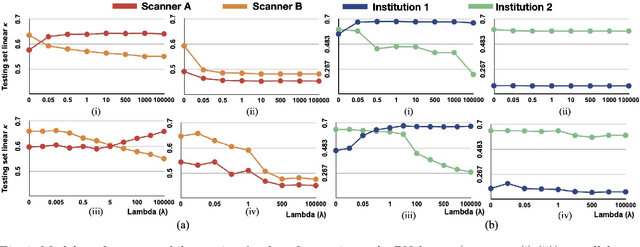
Abstract:Model brittleness is a key concern when deploying deep learning models in real-world medical settings. A model that has high performance at one institution may suffer a significant decline in performance when tested at other institutions. While pooling datasets from multiple institutions and retraining may provide a straightforward solution, it is often infeasible and may compromise patient privacy. An alternative approach is to fine-tune the model on subsequent institutions after training on the original institution. Notably, this approach degrades model performance at the original institution, a phenomenon known as catastrophic forgetting. In this paper, we develop an approach to address catastrophic forget-ting based on elastic weight consolidation combined with modulation of batch normalization statistics under two scenarios: first, for expanding the domain from one imaging system's data to another imaging system's, and second, for expanding the domain from a large multi-institutional dataset to another single institution dataset. We show that our approach outperforms several other state-of-the-art approaches and provide theoretical justification for the efficacy of batch normalization modulation. The results of this study are generally applicable to the deployment of any clinical deep learning model which requires domain expansion.
The unreasonable effectiveness of Batch-Norm statistics in addressing catastrophic forgetting across medical institutions
Nov 16, 2020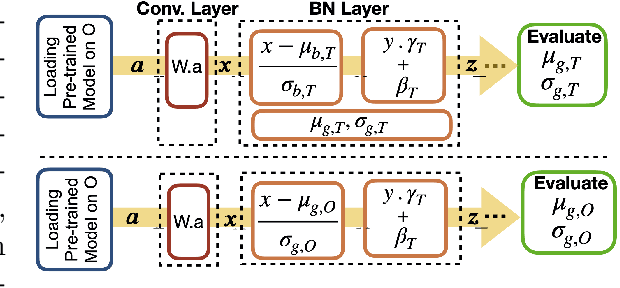
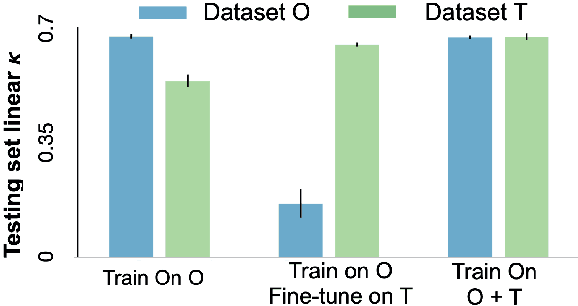
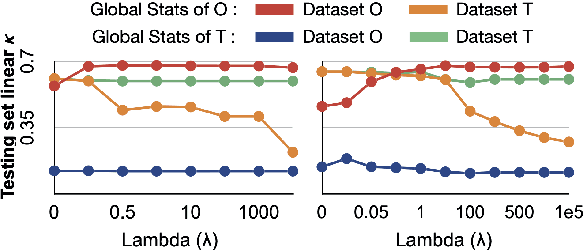
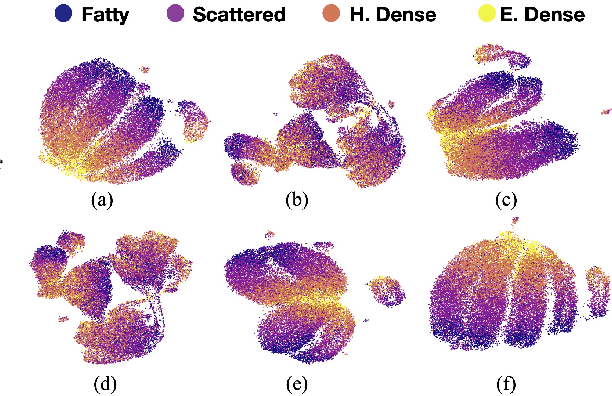
Abstract:Model brittleness is a primary concern when deploying deep learning models in medical settings owing to inter-institution variations, like patient demographics and intra-institution variation, such as multiple scanner types. While simply training on the combined datasets is fraught with data privacy limitations, fine-tuning the model on subsequent institutions after training it on the original institution results in a decrease in performance on the original dataset, a phenomenon called catastrophic forgetting. In this paper, we investigate trade-off between model refinement and retention of previously learned knowledge and subsequently address catastrophic forgetting for the assessment of mammographic breast density. More specifically, we propose a simple yet effective approach, adapting Elastic weight consolidation (EWC) using the global batch normalization (BN) statistics of the original dataset. The results of this study provide guidance for the deployment of clinical deep learning models where continuous learning is needed for domain expansion.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge