Hongfang Liu
N3C Natural Language Processing
A Cross-institutional Evaluation on Breast Cancer Phenotyping NLP Algorithms on Electronic Health Records
Mar 15, 2023Abstract:Objective: The generalizability of clinical large language models is usually ignored during the model development process. This study evaluated the generalizability of BERT-based clinical NLP models across different clinical settings through a breast cancer phenotype extraction task. Materials and Methods: Two clinical corpora of breast cancer patients were collected from the electronic health records from the University of Minnesota and the Mayo Clinic, and annotated following the same guideline. We developed three types of NLP models (i.e., conditional random field, bi-directional long short-term memory and CancerBERT) to extract cancer phenotypes from clinical texts. The models were evaluated for their generalizability on different test sets with different learning strategies (model transfer vs. locally trained). The entity coverage score was assessed with their association with the model performances. Results: We manually annotated 200 and 161 clinical documents at UMN and MC, respectively. The corpora of the two institutes were found to have higher similarity between the target entities than the overall corpora. The CancerBERT models obtained the best performances among the independent test sets from two clinical institutes and the permutation test set. The CancerBERT model developed in one institute and further fine-tuned in another institute achieved reasonable performance compared to the model developed on local data (micro-F1: 0.925 vs 0.932). Conclusions: The results indicate the CancerBERT model has the best learning ability and generalizability among the three types of clinical NLP models. The generalizability of the models was found to be correlated with the similarity of the target entities between the corpora.
Detecting Reddit Users with Depression Using a Hybrid Neural Network
Feb 03, 2023



Abstract:Depression is a widespread mental health issue, affecting an estimated 3.8% of the global population. It is also one of the main contributors to disability worldwide. Recently it is becoming popular for individuals to use social media platforms (e.g., Reddit) to express their difficulties and health issues (e.g., depression) and seek support from other users in online communities. It opens great opportunities to automatically identify social media users with depression by parsing millions of posts for potential interventions. Deep learning methods have begun to dominate in the field of machine learning and natural language processing (NLP) because of their ease of use, efficient processing, and state-of-the-art results on many NLP tasks. In this work, we propose a hybrid deep learning model which combines a pretrained sentence BERT (SBERT) and convolutional neural network (CNN) to detect individuals with depression with their Reddit posts. The sentence BERT is used to learn the meaningful representation of semantic information in each post. CNN enables the further transformation of those embeddings and the temporal identification of behavioral patterns of users. We trained and evaluated the model performance to identify Reddit users with depression by utilizing the Self-reported Mental Health Diagnoses (SMHD) data. The hybrid deep learning model achieved an accuracy of 0.86 and an F1 score of 0.86 and outperformed the state-of-the-art documented result (F1 score of 0.79) by other machine learning models in the literature. The results show the feasibility of the hybrid model to identify individuals with depression. Although the hybrid model is validated to detect depression with Reddit posts, it can be easily tuned and applied to other text classification tasks and different clinical applications.
Discrimination, calibration, and point estimate accuracy of GRU-D-Weibull architecture for real-time individualized endpoint prediction
Dec 19, 2022



Abstract:Real-time individual endpoint prediction has always been a challenging task but of great clinic utility for both patients and healthcare providers. With 6,879 chronic kidney disease stage 4 (CKD4) patients as a use case, we explored the feasibility and performance of gated recurrent units with decay that models Weibull probability density function (GRU-D-Weibull) as a semi-parametric longitudinal model for real-time individual endpoint prediction. GRU-D-Weibull has a maximum C-index of 0.77 at 4.3 years of follow-up, compared to 0.68 achieved by competing models. The L1-loss of GRU-D-Weibull is ~66% of XGB(AFT), ~60% of MTLR, and ~30% of AFT model at CKD4 index date. The average absolute L1-loss of GRU-D-Weibull is around one year, with a minimum of 40% Parkes serious error after index date. GRU-D-Weibull is not calibrated and significantly underestimates true survival probability. Feature importance tests indicate blood pressure becomes increasingly important during follow-up, while eGFR and blood albumin are less important. Most continuous features have non-linear/parabola impact on predicted survival time, and the results are generally consistent with existing knowledge. GRU-D-Weibull as a semi-parametric temporal model shows advantages in built-in parameterization of missing, native support for asynchronously arrived measurement, capability of output both probability and point estimates at arbitrary time point for arbitrary prediction horizon, improved discrimination and point estimate accuracy after incorporating newly arrived data. Further research on its performance with more comprehensive input features, in-process or post-process calibration are warranted to benefit CKD4 or alike terminally-ill patients.
The NLP Sandbox: an efficient model-to-data system to enable federated and unbiased evaluation of clinical NLP models
Jun 28, 2022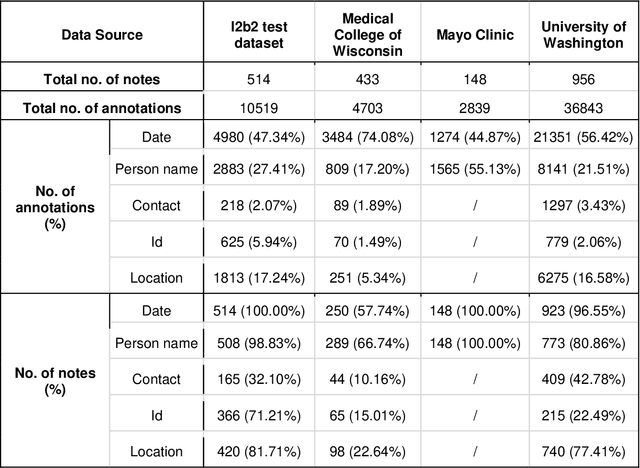
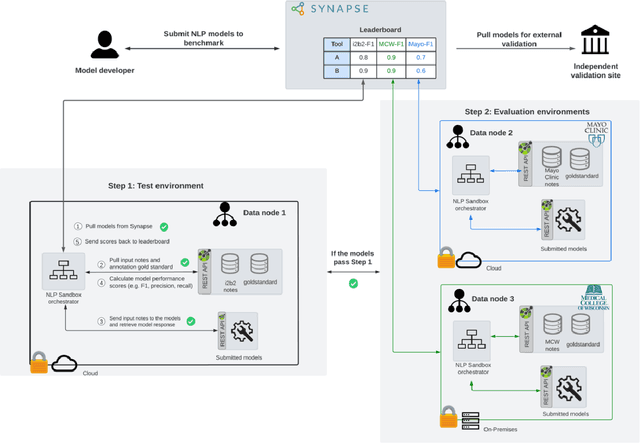
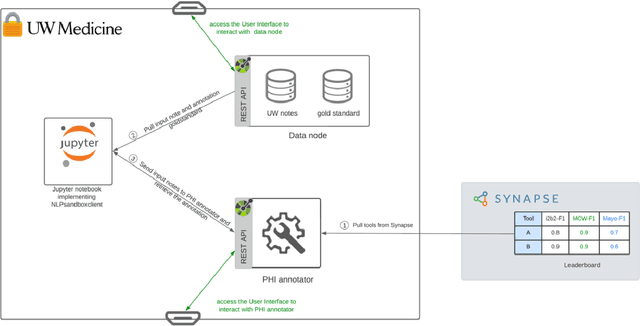
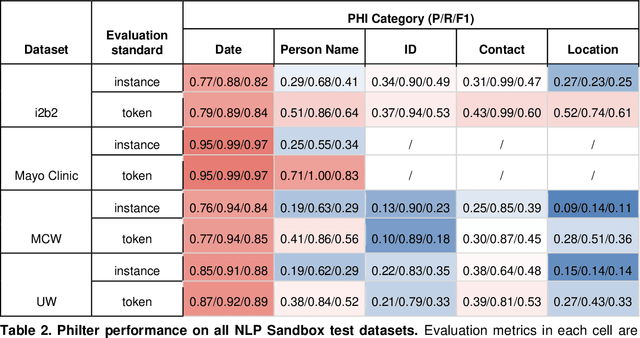
Abstract:Objective The evaluation of natural language processing (NLP) models for clinical text de-identification relies on the availability of clinical notes, which is often restricted due to privacy concerns. The NLP Sandbox is an approach for alleviating the lack of data and evaluation frameworks for NLP models by adopting a federated, model-to-data approach. This enables unbiased federated model evaluation without the need for sharing sensitive data from multiple institutions. Materials and Methods We leveraged the Synapse collaborative framework, containerization software, and OpenAPI generator to build the NLP Sandbox (nlpsandbox.io). We evaluated two state-of-the-art NLP de-identification focused annotation models, Philter and NeuroNER, using data from three institutions. We further validated model performance using data from an external validation site. Results We demonstrated the usefulness of the NLP Sandbox through de-identification clinical model evaluation. The external developer was able to incorporate their model into the NLP Sandbox template and provide user experience feedback. Discussion We demonstrated the feasibility of using the NLP Sandbox to conduct a multi-site evaluation of clinical text de-identification models without the sharing of data. Standardized model and data schemas enable smooth model transfer and implementation. To generalize the NLP Sandbox, work is required on the part of data owners and model developers to develop suitable and standardized schemas and to adapt their data or model to fit the schemas. Conclusions The NLP Sandbox lowers the barrier to utilizing clinical data for NLP model evaluation and facilitates federated, multi-site, unbiased evaluation of NLP models.
RxWhyQA: a clinical question-answering dataset with the challenge of multi-answer questions
Jan 07, 2022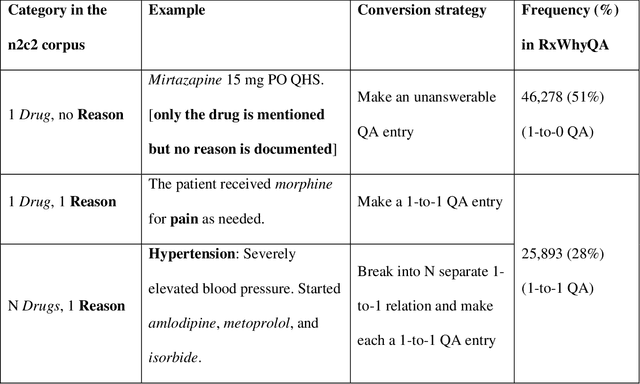
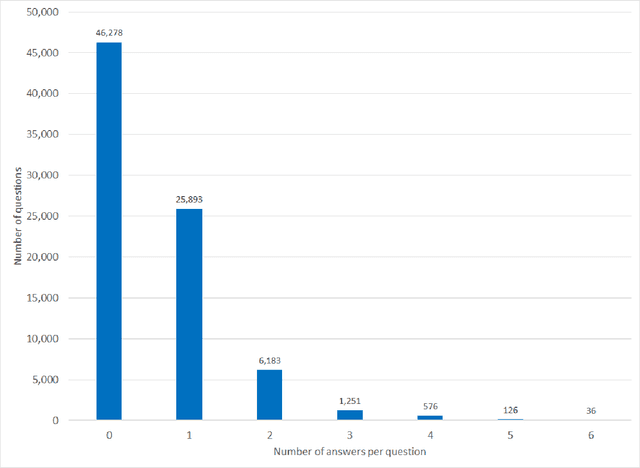


Abstract:Objectives Create a dataset for the development and evaluation of clinical question-answering (QA) systems that can handle multi-answer questions. Materials and Methods We leveraged the annotated relations from the 2018 National NLP Clinical Challenges (n2c2) corpus to generate a QA dataset. The 1-to-0 and 1-to-N drug-reason relations formed the unanswerable and multi-answer entries, which represent challenging scenarios lacking in the existing clinical QA datasets. Results The result RxWhyQA dataset contains 91,440 QA entries, of which half are unanswerable, and 21% (n=19,269) of the answerable ones require multiple answers. The dataset conforms to the community-vetted Stanford Question Answering Dataset (SQuAD) format. Discussion The RxWhyQA is useful for comparing different systems that need to handle the zero- and multi-answer challenges, demanding dual mitigation of both false positive and false negative answers. Conclusion We created and shared a clinical QA dataset with a focus on multi-answer questions to represent real-world scenarios.
An Open Natural Language Processing Development Framework for EHR-based Clinical Research: A case demonstration using the National COVID Cohort Collaborative (N3C)
Oct 20, 2021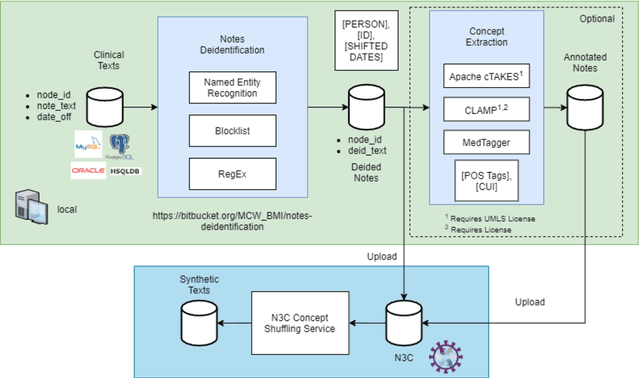
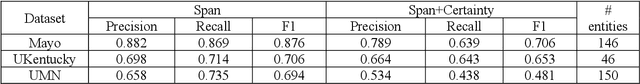
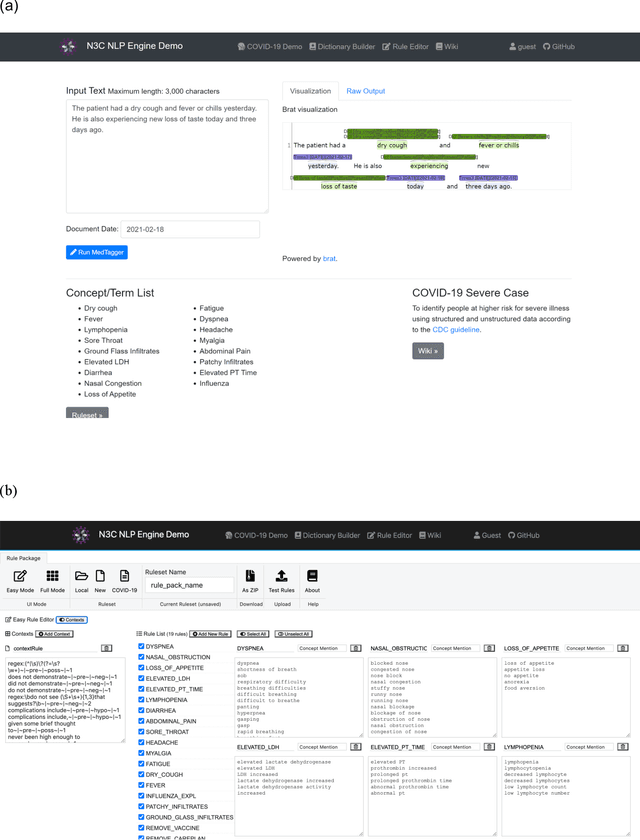
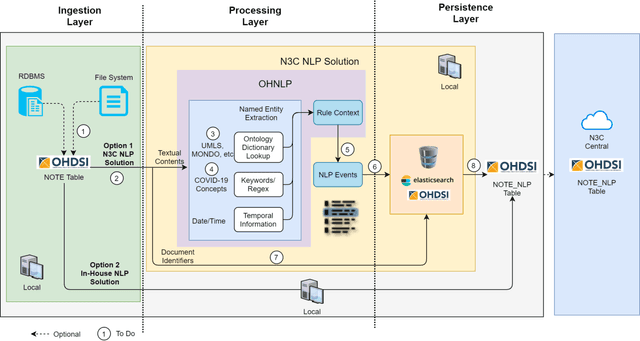
Abstract:While we pay attention to the latest advances in clinical natural language processing (NLP), we can notice some resistance in the clinical and translational research community to adopt NLP models due to limited transparency, Interpretability and usability. Built upon our previous work, in this study, we proposed an open natural language processing development framework and evaluated it through the implementation of NLP algorithms for the National COVID Cohort Collaborative (N3C). Based on the interests in information extraction from COVID-19 related clinical notes, our work includes 1) an open data annotation process using COVID-19 signs and symptoms as the use case, 2) a community-driven ruleset composing platform, and 3) a synthetic text data generation workflow to generate texts for information extraction tasks without involving human subjects. The generated corpora derived out of the texts from multiple intuitions and gold standard annotation are tested on a single institution's rule set has the performances in F1 score of 0.876, 0.706 and 0.694, respectively. The study as a consortium effort of the N3C NLP subgroup demonstrates the feasibility of creating a federated NLP algorithm development and benchmarking platform to enhance multi-institution clinical NLP study.
CancerBERT: a BERT model for Extracting Breast Cancer Phenotypes from Electronic Health Records
Aug 25, 2021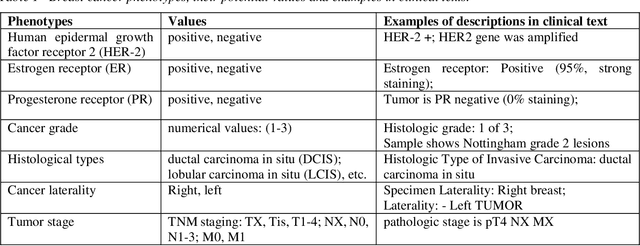
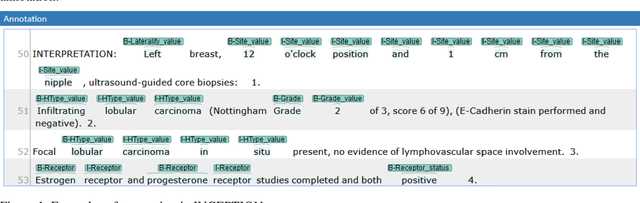

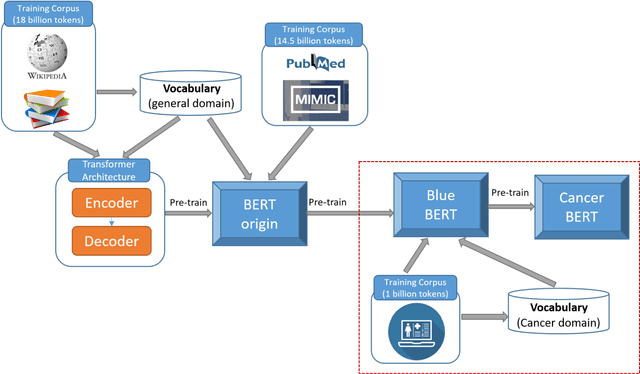
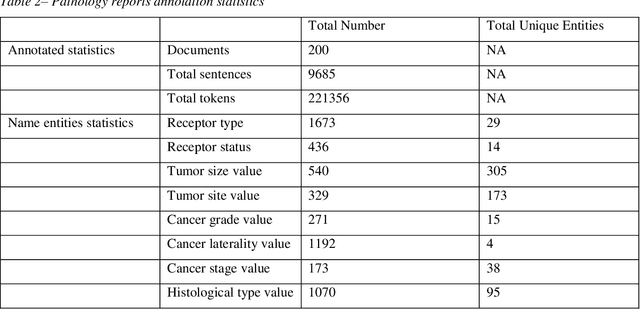
Abstract:Accurate extraction of breast cancer patients' phenotypes is important for clinical decision support and clinical research. Current models do not take full advantage of cancer domain-specific corpus, whether pre-training Bidirectional Encoder Representations from Transformer model on cancer-specific corpus could improve the performances of extracting breast cancer phenotypes from texts data remains to be explored. The objective of this study is to develop and evaluate the CancerBERT model for extracting breast cancer phenotypes from clinical texts in electronic health records. This data used in the study included 21,291 breast cancer patients diagnosed from 2010 to 2020, patients' clinical notes and pathology reports were collected from the University of Minnesota Clinical Data Repository (UMN). Results: About 3 million clinical notes and pathology reports in electronic health records for 21,291 breast cancer patients were collected to train the CancerBERT model. 200 pathology reports and 50 clinical notes of breast cancer patients that contain 9,685 sentences and 221,356 tokens were manually annotated by two annotators. 20% of the annotated data was used as a test set. Our CancerBERT model achieved the best performance with macro F1 scores equal to 0.876 (95% CI, 0.896-0.902) for exact match and 0.904 (95% CI, 0.896-0.902) for the lenient match. The NER models we developed would facilitate the automated information extraction from clinical texts to further help clinical decision support. Conclusions and Relevance: In this study, we focused on the breast cancer-related concepts extraction from EHR data and obtained a comprehensive annotated dataset that contains 7 types of breast cancer-related concepts. The CancerBERT model with customized vocabulary could significantly improve the performance for extracting breast cancer phenotypes from clinical texts.
An Empirical Study of UMLS Concept Extraction from Clinical Notes using Boolean Combination Ensembles
Aug 04, 2021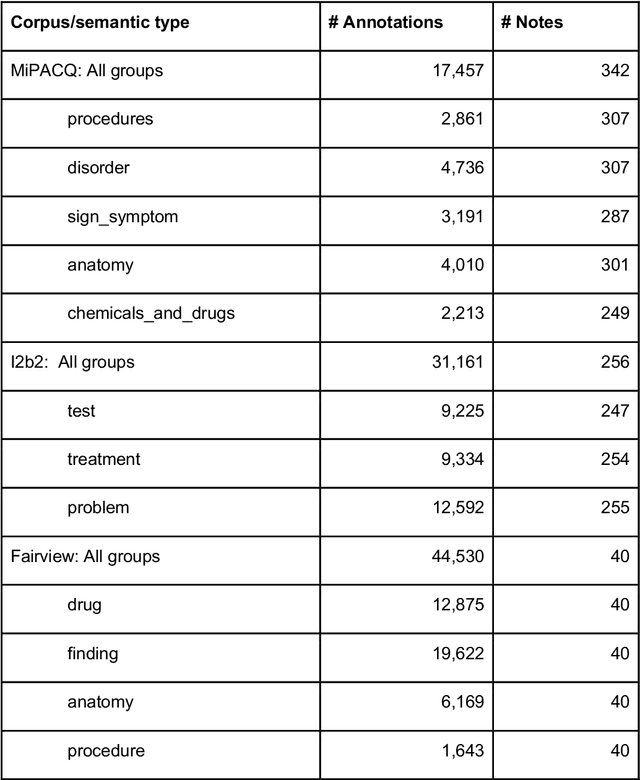
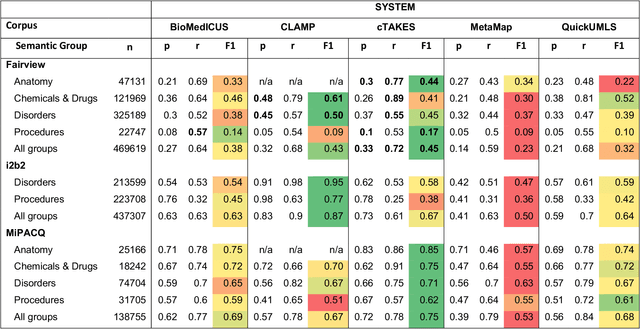
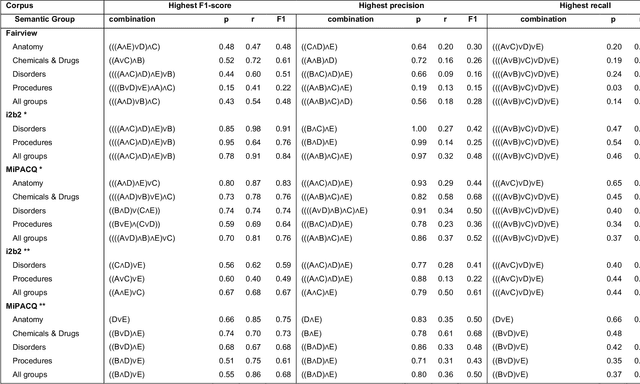
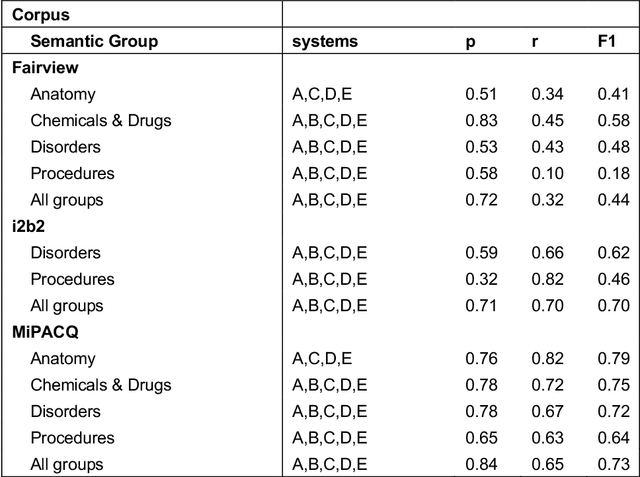
Abstract:Our objective in this study is to investigate the behavior of Boolean operators on combining annotation output from multiple Natural Language Processing (NLP) systems across multiple corpora and to assess how filtering by aggregation of Unified Medical Language System (UMLS) Metathesaurus concepts affects system performance for Named Entity Recognition (NER) of UMLS concepts. We used three corpora annotated for UMLS concepts: 2010 i2b2 VA challenge set (31,161 annotations), Multi-source Integrated Platform for Answering Clinical Questions (MiPACQ) corpus (17,457 annotations including UMLS concept unique identifiers), and Fairview Health Services corpus (44,530 annotations). Our results showed that for UMLS concept matching, Boolean ensembling of the MiPACQ corpus trended towards higher performance over individual systems. Use of an approximate grid-search can help optimize the precision-recall tradeoff and can provide a set of heuristics for choosing an optimal set of ensembles.
Leveraging a Joint of Phenotypic and Genetic Features on Cancer Patient Subgrouping
Mar 30, 2021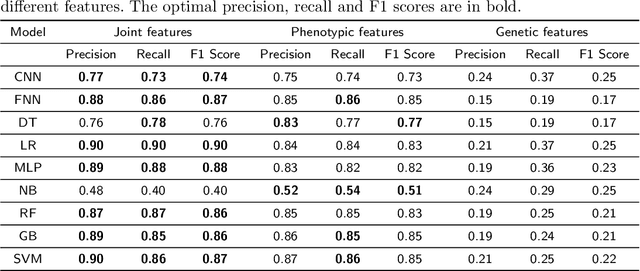
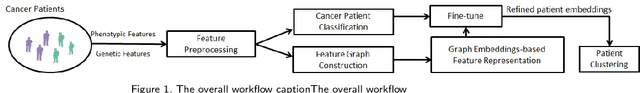
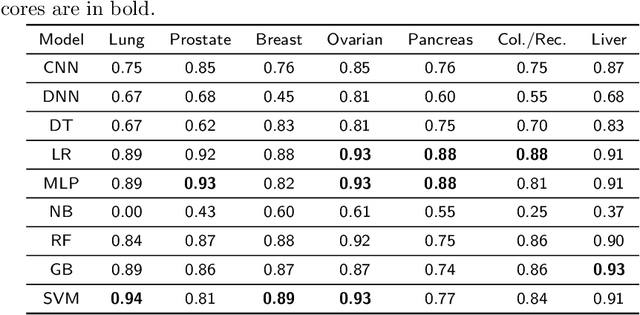
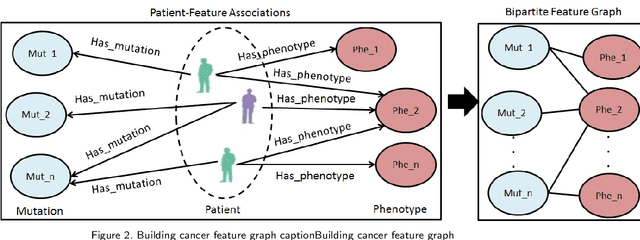
Abstract:Cancer is responsible for millions of deaths worldwide every year. Although significant progress has been achieved in cancer medicine, many issues remain to be addressed for improving cancer therapy. Appropriate cancer patient stratification is the prerequisite for selecting appropriate treatment plan, as cancer patients are of known heterogeneous genetic make-ups and phenotypic differences. In this study, built upon deep phenotypic characterizations extractable from Mayo Clinic electronic health records (EHRs) and genetic test reports for a collection of cancer patients, we developed a system leveraging a joint of phenotypic and genetic features for cancer patient subgrouping. The workflow is roughly divided into three parts: feature preprocessing, cancer patient classification, and cancer patient clustering based. In feature preprocessing step, we performed filtering, retaining the most relevant features. In cancer patient classification, we utilized joint categorical features to build a patient-feature matrix and applied nine different machine learning models, Random Forests (RF), Decision Tree (DT), Support Vector Machine (SVM), Naive Bayes (NB), Logistic Regression (LR), Multilayer Perceptron (MLP), Gradient Boosting (GB), Convolutional Neural Network (CNN), and Feedforward Neural Network (FNN), for classification purposes. Finally, in the cancer patient clustering step, we leveraged joint embeddings features and patient-feature associations to build an undirected feature graph and then trained the cancer feature node embeddings.
Comparisons of Graph Neural Networks on Cancer Classification Leveraging a Joint of Phenotypic and Genetic Features
Jan 14, 2021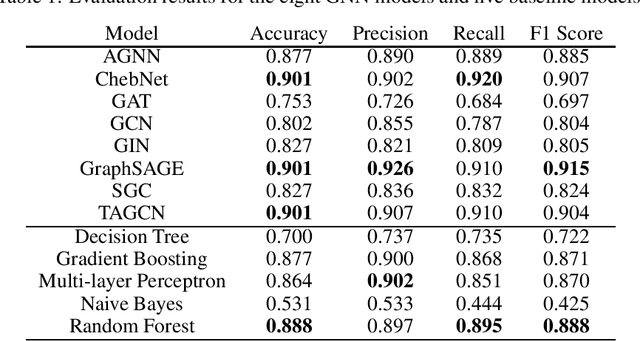
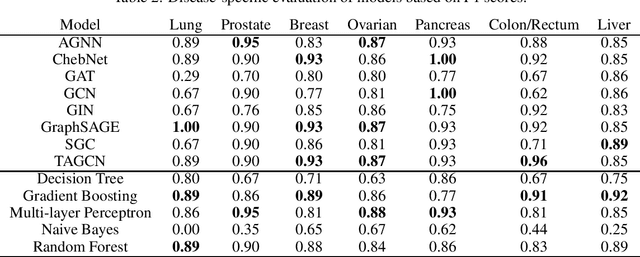
Abstract:Cancer is responsible for millions of deaths worldwide every year. Although significant progress hasbeen achieved in cancer medicine, many issues remain to be addressed for improving cancer therapy.Appropriate cancer patient stratification is the prerequisite for selecting appropriate treatment plan, ascancer patients are of known heterogeneous genetic make-ups and phenotypic differences. In thisstudy, built upon deep phenotypic characterizations extractable from Mayo Clinic electronic healthrecords (EHRs) and genetic test reports for a collection of cancer patients, we evaluated variousgraph neural networks (GNNs) leveraging a joint of phenotypic and genetic features for cancer typeclassification. Models were applied and fine-tuned on the Mayo Clinic cancer disease dataset. Theassessment was done through the reported accuracy, precision, recall, and F1 values as well as throughF1 scores based on the disease class. Per our evaluation results, GNNs on average outperformed thebaseline models with mean statistics always being higher that those of the baseline models (0.849 vs0.772 for accuracy, 0.858 vs 0.794 for precision, 0.843 vs 0.759 for recall, and 0.843 vs 0.855 for F1score). Among GNNs, ChebNet, GraphSAGE, and TAGCN showed the best performance, while GATshowed the worst. We applied and compared eight GNN models including AGNN, ChebNet, GAT,GCN, GIN, GraphSAGE, SGC, and TAGCN on the Mayo Clinic cancer disease dataset and assessedtheir performance as well as compared them with each other and with more conventional machinelearning models such as decision tree, gradient boosting, multi-layer perceptron, naive bayes, andrandom forest which we used as the baselines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge