Harald Binder
Assessing Multimodal Chronic Wound Embeddings with Expert Triplet Agreement
Apr 02, 2026Abstract:Recessive dystrophic epidermolysis bullosa (RDEB) is a rare genetic skin disorder for which clinicians greatly benefit from finding similar cases using images and clinical text. However, off-the-shelf foundation models do not reliably capture clinically meaningful features for this heterogeneous, long-tail disease, and structured measurement of agreement with experts is challenging. To address these gaps, we propose evaluating embedding spaces with expert ordinal comparisons (triplet judgments), which are fast to collect and encode implicit clinical similarity knowledge. We further introduce TriDerm, a multimodal framework that learns interpretable wound representations from small cohorts by integrating wound imagery, boundary masks, and expert reports. On the vision side, TriDerm adapts visual foundation models to RDEB using wound-level attention pooling and non-contrastive representation learning. For text, we prompt large language models with comparison queries and recover medically meaningful representations via soft ordinal embeddings (SOE). We show that visual and textual modalities capture complementary aspects of wound phenotype, and that fusing both modalities yields 73.5% agreement with experts, outperforming the best off-the-shelf single-modality foundation model by over 5.6 percentage points. We make the expert annotation tool, model code and representative dataset samples publicly available.
Embedding interpretable $\ell_1$-regression into neural networks for uncovering temporal structure in cell imaging
Mar 03, 2026Abstract:While artificial neural networks excel in unsupervised learning of non-sparse structure, classical statistical regression techniques offer better interpretability, in particular when sparseness is enforced by $\ell_1$ regularization, enabling identification of which factors drive observed dynamics. We investigate how these two types of approaches can be optimally combined, exemplarily considering two-photon calcium imaging data where sparse autoregressive dynamics are to be extracted. We propose embedding a vector autoregressive (VAR) model as an interpretable regression technique into a convolutional autoencoder, which provides dimension reduction for tractable temporal modeling. A skip connection separately addresses non-sparse static spatial information, selectively channeling sparse structure into the $\ell_1$-regularized VAR. $\ell_1$-estimation of regression parameters is enabled by differentiating through the piecewise linear solution path. This is contrasted with approaches where the autoencoder does not adapt to the VAR model. Having an embedded statistical model also enables a testing approach for comparing temporal sequences from the same observational unit. Additionally, contribution maps visualize which spatial regions drive the learned dynamics.
A statistical perspective on transformers for small longitudinal cohort data
Feb 18, 2026Abstract:Modeling of longitudinal cohort data typically involves complex temporal dependencies between multiple variables. There, the transformer architecture, which has been highly successful in language and vision applications, allows us to account for the fact that the most recently observed time points in an individual's history may not always be the most important for the immediate future. This is achieved by assigning attention weights to observations of an individual based on a transformation of their values. One reason why these ideas have not yet been fully leveraged for longitudinal cohort data is that typically, large datasets are required. Therefore, we present a simplified transformer architecture that retains the core attention mechanism while reducing the number of parameters to be estimated, to be more suitable for small datasets with few time points. Guided by a statistical perspective on transformers, we use an autoregressive model as a starting point and incorporate attention as a kernel-based operation with temporal decay, where aggregation of multiple transformer heads, i.e. different candidate weighting schemes, is expressed as accumulating evidence on different types of underlying characteristics of individuals. This also enables a permutation-based statistical testing procedure for identifying contextual patterns. In a simulation study, the approach is shown to recover contextual dependencies even with a small number of individuals and time points. In an application to data from a resilience study, we identify temporal patterns in the dynamics of stress and mental health. This indicates that properly adapted transformers can not only achieve competitive predictive performance, but also uncover complex context dependencies in small data settings.
Contrasting Global and Patient-Specific Regression Models via a Neural Network Representation
Jan 26, 2026Abstract:When developing clinical prediction models, it can be challenging to balance between global models that are valid for all patients and personalized models tailored to individuals or potentially unknown subgroups. To aid such decisions, we propose a diagnostic tool for contrasting global regression models and patient-specific (local) regression models. The core utility of this tool is to identify where and for whom a global model may be inadequate. We focus on regression models and specifically suggest a localized regression approach that identifies regions in the predictor space where patients are not well represented by the global model. As localization becomes challenging when dealing with many predictors, we propose modeling in a dimension-reduced latent representation obtained from an autoencoder. Using such a neural network architecture for dimension reduction enables learning a latent representation simultaneously optimized for both good data reconstruction and for revealing local outcome-related associations suitable for robust localized regression. We illustrate the proposed approach with a clinical study involving patients with chronic obstructive pulmonary disease. Our findings indicate that the global model is adequate for most patients but that indeed specific subgroups benefit from personalized models. We also demonstrate how to map these subgroup models back to the original predictors, providing insight into why the global model falls short for these groups. Thus, the principal application and diagnostic yield of our tool is the identification and characterization of patients or subgroups whose outcome associations deviate from the global model.
Small Data Explainer -- The impact of small data methods in everyday life
Jul 15, 2025Abstract:The emergence of breakthrough artificial intelligence (AI) techniques has led to a renewed focus on how small data settings, i.e., settings with limited information, can benefit from such developments. This includes societal issues such as how best to include under-represented groups in data-driven policy and decision making, or the health benefits of assistive technologies such as wearables. We provide a conceptual overview, in particular contrasting small data with big data, and identify common themes from exemplary case studies and application areas. Potential solutions are described in a more detailed technical overview of current data analysis and modelling techniques, highlighting contributions from different disciplines, such as knowledge-driven modelling from statistics and data-driven modelling from computer science. By linking application settings, conceptual contributions and specific techniques, we highlight what is already feasible and suggest what an agenda for fully leveraging small data might look like.
A systematic review of challenges and proposed solutions in modeling multimodal data
May 15, 2025Abstract:Multimodal data modeling has emerged as a powerful approach in clinical research, enabling the integration of diverse data types such as imaging, genomics, wearable sensors, and electronic health records. Despite its potential to improve diagnostic accuracy and support personalized care, modeling such heterogeneous data presents significant technical challenges. This systematic review synthesizes findings from 69 studies to identify common obstacles, including missing modalities, limited sample sizes, dimensionality imbalance, interpretability issues, and finding the optimal fusion techniques. We highlight recent methodological advances, such as transfer learning, generative models, attention mechanisms, and neural architecture search that offer promising solutions. By mapping current trends and innovations, this review provides a comprehensive overview of the field and offers practical insights to guide future research and development in multimodal modeling for medical applications.
Combining propensity score methods with variational autoencoders for generating synthetic data in presence of latent sub-groups
Dec 12, 2023Abstract:In settings requiring synthetic data generation based on a clinical cohort, e.g., due to data protection regulations, heterogeneity across individuals might be a nuisance that we need to control or faithfully preserve. The sources of such heterogeneity might be known, e.g., as indicated by sub-groups labels, or might be unknown and thus reflected only in properties of distributions, such as bimodality or skewness. We investigate how such heterogeneity can be preserved and controlled when obtaining synthetic data from variational autoencoders (VAEs), i.e., a generative deep learning technique that utilizes a low-dimensional latent representation. To faithfully reproduce unknown heterogeneity reflected in marginal distributions, we propose to combine VAEs with pre-transformations. For dealing with known heterogeneity due to sub-groups, we complement VAEs with models for group membership, specifically from propensity score regression. The evaluation is performed with a realistic simulation design that features sub-groups and challenging marginal distributions. The proposed approach faithfully recovers the latter, compared to synthetic data approaches that focus purely on marginal distributions. Propensity scores add complementary information, e.g., when visualized in the latent space, and enable sampling of synthetic data with or without sub-group specific characteristics. We also illustrate the proposed approach with real data from an international stroke trial that exhibits considerable distribution differences between study sites, in addition to bimodality. These results indicate that describing heterogeneity by statistical approaches, such as propensity score regression, might be more generally useful for complementing generative deep learning for obtaining synthetic data that faithfully reflects structure from clinical cohorts.
Investigating a domain adaptation approach for integrating different measurement instruments in a longitudinal clinical registry
Dec 01, 2023Abstract:In a longitudinal clinical registry, different measurement instruments might have been used for assessing individuals at different time points. To combine them, we investigate deep learning techniques for obtaining a joint latent representation, to which the items of different measurement instruments are mapped. This corresponds to domain adaptation, an established concept in computer science for image data. Using the proposed approach as an example, we evaluate the potential of domain adaptation in a longitudinal cohort setting with a rather small number of time points, motivated by an application with different motor function measurement instruments in a registry of spinal muscular atrophy (SMA) patients. There, we model trajectories in the latent representation by ordinary differential equations (ODEs), where person-specific ODE parameters are inferred from baseline characteristics. The goodness of fit and complexity of the ODE solutions then allows to judge the measurement instrument mappings. We subsequently explore how alignment can be improved by incorporating corresponding penalty terms into model fitting. To systematically investigate the effect of differences between measurement instruments, we consider several scenarios based on modified SMA data, including scenarios where a mapping should be feasible in principle and scenarios where no perfect mapping is available. While misalignment increases in more complex scenarios, some structure is still recovered, even if the availability of measurement instruments depends on patient state. A reasonable mapping is feasible also in the more complex real SMA dataset. These results indicate that domain adaptation might be more generally useful in statistical modeling for longitudinal registry data.
A statistical approach to latent dynamic modeling with differential equations
Nov 27, 2023



Abstract:Ordinary differential equations (ODEs) can provide mechanistic models of temporally local changes of processes, where parameters are often informed by external knowledge. While ODEs are popular in systems modeling, they are less established for statistical modeling of longitudinal cohort data, e.g., in a clinical setting. Yet, modeling of local changes could also be attractive for assessing the trajectory of an individual in a cohort in the immediate future given its current status, where ODE parameters could be informed by further characteristics of the individual. However, several hurdles so far limit such use of ODEs, as compared to regression-based function fitting approaches. The potentially higher level of noise in cohort data might be detrimental to ODEs, as the shape of the ODE solution heavily depends on the initial value. In addition, larger numbers of variables multiply such problems and might be difficult to handle for ODEs. To address this, we propose to use each observation in the course of time as the initial value to obtain multiple local ODE solutions and build a combined estimator of the underlying dynamics. Neural networks are used for obtaining a low-dimensional latent space for dynamic modeling from a potentially large number of variables, and for obtaining patient-specific ODE parameters from baseline variables. Simultaneous identification of dynamic models and of a latent space is enabled by recently developed differentiable programming techniques. We illustrate the proposed approach in an application with spinal muscular atrophy patients and a corresponding simulation study. In particular, modeling of local changes in health status at any point in time is contrasted to the interpretation of functions obtained from a global regression. This more generally highlights how different application settings might demand different modeling strategies.
Deep learning and differential equations for modeling changes in individual-level latent dynamics between observation periods
Feb 15, 2022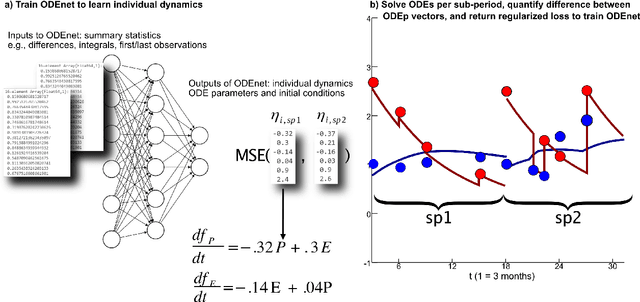
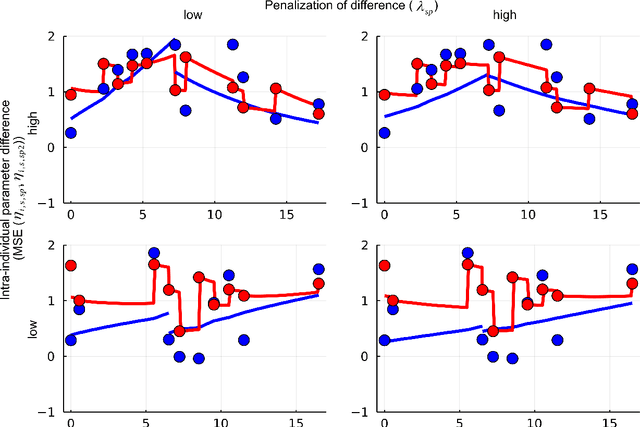
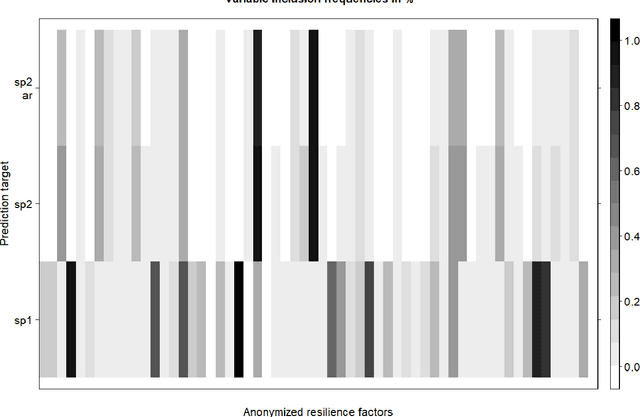
Abstract:When modeling longitudinal biomedical data, often dimensionality reduction as well as dynamic modeling in the resulting latent representation is needed. This can be achieved by artificial neural networks for dimension reduction, and differential equations for dynamic modeling of individual-level trajectories. However, such approaches so far assume that parameters of individual-level dynamics are constant throughout the observation period. Motivated by an application from psychological resilience research, we propose an extension where different sets of differential equation parameters are allowed for observation sub-periods. Still, estimation for intra-individual sub-periods is coupled for being able to fit the model also with a relatively small dataset. We subsequently derive prediction targets from individual dynamic models of resilience in the application. These serve as interpretable resilience-related outcomes, to be predicted from characteristics of individuals, measured at baseline and a follow-up time point, and selecting a small set of important predictors. Our approach is seen to successfully identify individual-level parameters of dynamic models that allows us to stably select predictors, i.e., resilience factors. Furthermore, we can identify those characteristics of individuals that are the most promising for updates at follow-up, which might inform future study design. This underlines the usefulness of our proposed deep dynamic modeling approach with changes in parameters between observation sub-periods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge