Ge Wang
Xi'an Jiaotong University
LIT-Former: Linking In-plane and Through-plane Transformers for Simultaneous CT Image Denoising and Deblurring
Feb 21, 2023Abstract:This paper studies 3D low-dose computed tomography (CT) imaging. Although various deep learning methods were developed in this context, typically they perform denoising due to low-dose and deblurring for super-resolution separately. Up to date, little work was done for simultaneous in-plane denoising and through-plane deblurring, which is important to improve clinical CT images. For this task, a straightforward method is to directly train an end-to-end 3D network. However, it demands much more training data and expensive computational costs. Here, we propose to link in-plane and through-plane transformers for simultaneous in-plane denoising and through-plane deblurring, termed as LIT-Former, which can efficiently synergize in-plane and through-plane sub-tasks for 3D CT imaging and enjoy the advantages of both convolution and transformer networks. LIT-Former has two novel designs: efficient multi-head self-attention modules (eMSM) and efficient convolutional feed-forward networks (eCFN). First, eMSM integrates in-plane 2D self-attention and through-plane 1D self-attention to efficiently capture global interactions of 3D self-attention, the core unit of transformer networks. Second, eCFN integrates 2D convolution and 1D convolution to extract local information of 3D convolution in the same fashion. As a result, the proposed LIT-Former synergizes these two sub-tasks, significantly reducing the computational complexity as compared to 3D counterparts and enabling rapid convergence. Extensive experimental results on simulated and clinical datasets demonstrate superior performance over state-of-the-art models.
QS-ADN: Quasi-Supervised Artifact Disentanglement Network for Low-Dose CT Image Denoising by Local Similarity Among Unpaired Data
Feb 08, 2023Abstract:Deep learning has been successfully applied to low-dose CT (LDCT) image denoising for reducing potential radiation risk. However, the widely reported supervised LDCT denoising networks require a training set of paired images, which is expensive to obtain and cannot be perfectly simulated. Unsupervised learning utilizes unpaired data and is highly desirable for LDCT denoising. As an example, an artifact disentanglement network (ADN) relies on unparied images and obviates the need for supervision but the results of artifact reduction are not as good as those through supervised learning.An important observation is that there is often hidden similarity among unpaired data that can be utilized. This paper introduces a new learning mode, called quasi-supervised learning, to empower the ADN for LDCT image denoising.For every LDCT image, the best matched image is first found from an unpaired normal-dose CT (NDCT) dataset. Then, the matched pairs and the corresponding matching degree as prior information are used to construct and train our ADN-type network for LDCT denoising.The proposed method is different from (but compatible with) supervised and semi-supervised learning modes and can be easily implemented by modifying existing networks. The experimental results show that the method is competitive with state-of-the-art methods in terms of noise suppression and contextual fidelity. The code and working dataset are publicly available at https://github.com/ruanyuhui/ADN-QSDL.git.
Improving CT Image Segmentation Accuracy Using StyleGAN Driven Data Augmentation
Feb 07, 2023Abstract:Medical Image Segmentation is a useful application for medical image analysis including detecting diseases and abnormalities in imaging modalities such as MRI, CT etc. Deep learning has proven to be promising for this task but usually has a low accuracy because of the lack of appropriate publicly available annotated or segmented medical datasets. In addition, the datasets that are available may have a different texture because of different dosage values or scanner properties than the images that need to be segmented. This paper presents a StyleGAN-driven approach for segmenting publicly available large medical datasets by using readily available extremely small annotated datasets in similar modalities. The approach involves augmenting the small segmented dataset and eliminating texture differences between the two datasets. The dataset is augmented by being passed through six different StyleGANs that are trained on six different style images taken from the large non-annotated dataset we want to segment. Specifically, style transfer is used to augment the training dataset. The annotations of the training dataset are hence combined with the textures of the non-annotated dataset to generate new anatomically sound images. The augmented dataset is then used to train a U-Net segmentation network which displays a significant improvement in the segmentation accuracy in segmenting the large non-annotated dataset.
Data-Efficient Protein 3D Geometric Pretraining via Refinement of Diffused Protein Structure Decoy
Feb 05, 2023Abstract:Learning meaningful protein representation is important for a variety of biological downstream tasks such as structure-based drug design. Having witnessed the success of protein sequence pretraining, pretraining for structural data which is more informative has become a promising research topic. However, there are three major challenges facing protein structure pretraining: insufficient sample diversity, physically unrealistic modeling, and the lack of protein-specific pretext tasks. To try to address these challenges, we present the 3D Geometric Pretraining. In this paper, we propose a unified framework for protein pretraining and a 3D geometric-based, data-efficient, and protein-specific pretext task: RefineDiff (Refine the Diffused Protein Structure Decoy). After pretraining our geometric-aware model with this task on limited data(less than 1% of SOTA models), we obtained informative protein representations that can achieve comparable performance for various downstream tasks.
DiffusionCT: Latent Diffusion Model for CT Image Standardization
Jan 20, 2023



Abstract:Computed tomography (CT) imaging is a widely used modality for early lung cancer diagnosis, treatment, and prognosis. Features extracted from CT images are now accepted to quantify spatial and temporal variations in tumor architecture and function. However, CT images are often acquired using scanners from different vendors with customized acquisition standards, resulting in significantly different texture features even for the same patient, posing a fundamental challenge to downstream studies. Existing CT image harmonization models rely on supervised or semi-supervised techniques, with limited performance. In this paper, we have proposed a diffusion-based CT image standardization model called DiffusionCT which works on latent space by mapping latent distribution into a standard distribution. DiffusionCT incorporates an Unet-based encoder-decoder and a diffusion model embedded in its bottleneck part. The Unet first trained without the diffusion model to learn the latent representation of the input data. The diffusion model is trained in the next training phase. All the trained models work together on image standardization. The encoded representation outputted from the Unet encoder passes through the diffusion model, and the diffusion model maps the distribution in to target standard image domain. Finally, the decode takes that transformed latent representation to synthesize a standardized image. The experimental results show that DiffusionCT significantly improves the performance of the standardization task.
Patch-Based Denoising Diffusion Probabilistic Model for Sparse-View CT Reconstruction
Nov 18, 2022



Abstract:Sparse-view computed tomography (CT) can be used to reduce radiation dose greatly but is suffers from severe image artifacts. Recently, the deep learning based method for sparse-view CT reconstruction has attracted a major attention. However, neural networks often have a limited ability to remove the artifacts when they only work in the image domain. Deep learning-based sinogram processing can achieve a better anti-artifact performance, but it inevitably requires feature maps of the whole image in a video memory, which makes handling large-scale or three-dimensional (3D) images rather challenging. In this paper, we propose a patch-based denoising diffusion probabilistic model (DDPM) for sparse-view CT reconstruction. A DDPM network based on patches extracted from fully sampled projection data is trained and then used to inpaint down-sampled projection data. The network does not require paired full-sampled and down-sampled data, enabling unsupervised learning. Since the data processing is patch-based, the deep learning workflow can be distributed in parallel, overcoming the memory problem of large-scale data. Our experiments show that the proposed method can effectively suppress few-view artifacts while faithfully preserving textural details.
Using Context-to-Vector with Graph Retrofitting to Improve Word Embeddings
Oct 30, 2022



Abstract:Although contextualized embeddings generated from large-scale pre-trained models perform well in many tasks, traditional static embeddings (e.g., Skip-gram, Word2Vec) still play an important role in low-resource and lightweight settings due to their low computational cost, ease of deployment, and stability. In this paper, we aim to improve word embeddings by 1) incorporating more contextual information from existing pre-trained models into the Skip-gram framework, which we call Context-to-Vec; 2) proposing a post-processing retrofitting method for static embeddings independent of training by employing priori synonym knowledge and weighted vector distribution. Through extrinsic and intrinsic tasks, our methods are well proven to outperform the baselines by a large margin.
Low-Dose CT Using Denoising Diffusion Probabilistic Model for 20$\times$ Speedup
Sep 29, 2022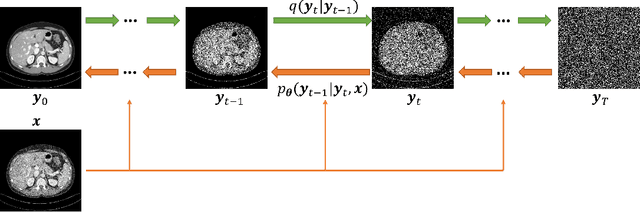
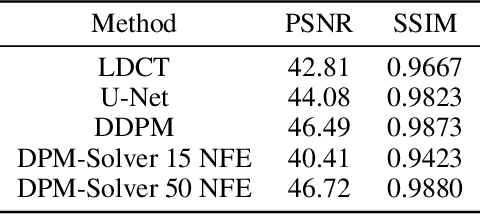
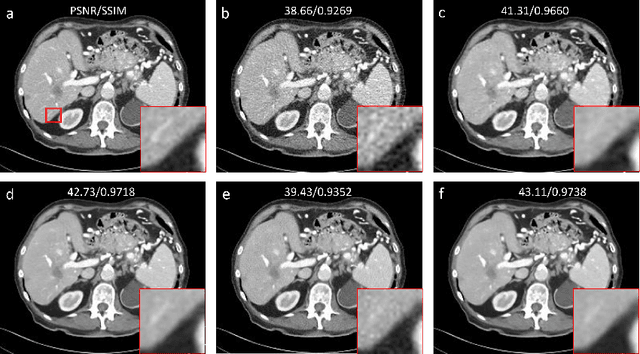

Abstract:Low-dose computed tomography (LDCT) is an important topic in the field of radiology over the past decades. LDCT reduces ionizing radiation-induced patient health risks but it also results in a low signal-to-noise ratio (SNR) and a potential compromise in the diagnostic performance. In this paper, to improve the LDCT denoising performance, we introduce the conditional denoising diffusion probabilistic model (DDPM) and show encouraging results with a high computational efficiency. Specifically, given the high sampling cost of the original DDPM model, we adapt the fast ordinary differential equation (ODE) solver for a much-improved sampling efficiency. The experiments show that the accelerated DDPM can achieve 20x speedup without compromising image quality.
Conversion Between CT and MRI Images Using Diffusion and Score-Matching Models
Sep 29, 2022
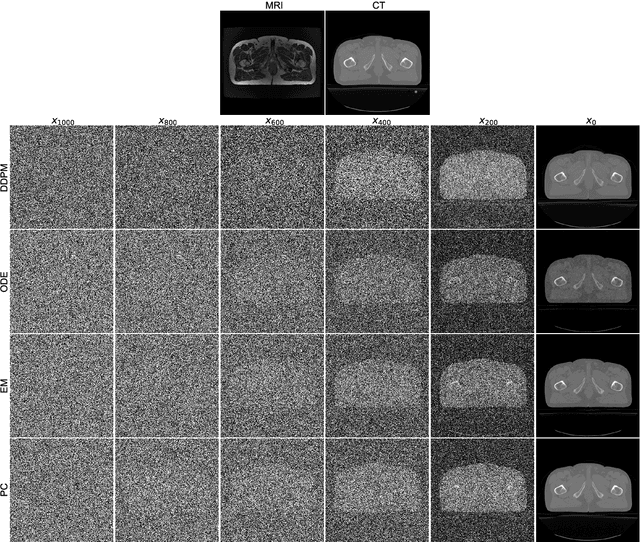
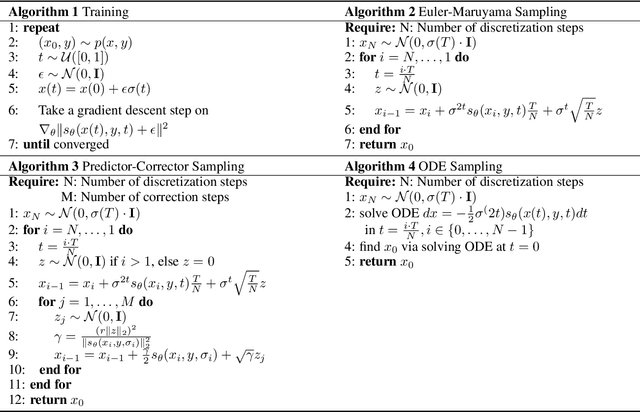


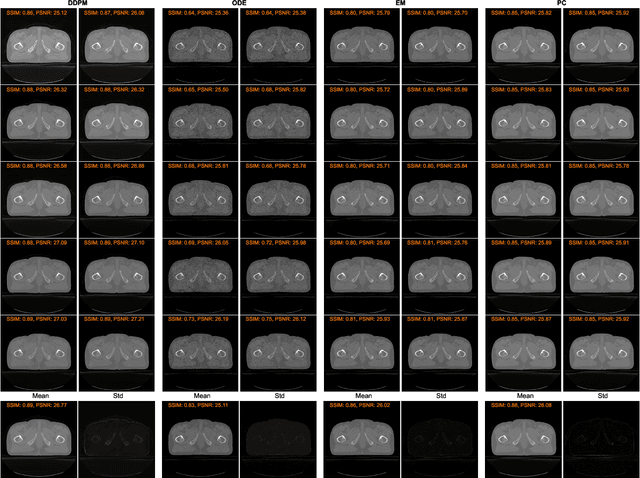
Abstract:MRI and CT are most widely used medical imaging modalities. It is often necessary to acquire multi-modality images for diagnosis and treatment such as radiotherapy planning. However, multi-modality imaging is not only costly but also introduces misalignment between MRI and CT images. To address this challenge, computational conversion is a viable approach between MRI and CT images, especially from MRI to CT images. In this paper, we propose to use an emerging deep learning framework called diffusion and score-matching models in this context. Specifically, we adapt denoising diffusion probabilistic and score-matching models, use four different sampling strategies, and compare their performance metrics with that using a convolutional neural network and a generative adversarial network model. Our results show that the diffusion and score-matching models generate better synthetic CT images than the CNN and GAN models. Furthermore, we investigate the uncertainties associated with the diffusion and score-matching networks using the Monte-Carlo method, and improve the results by averaging their Monte-Carlo outputs. Our study suggests that diffusion and score-matching models are powerful to generate high quality images conditioned on an image obtained using a complementary imaging modality, analytically rigorous with clear explainability, and highly competitive with CNNs and GANs for image synthesis.
Quasi-supervised Learning for Super-resolution PET
Sep 03, 2022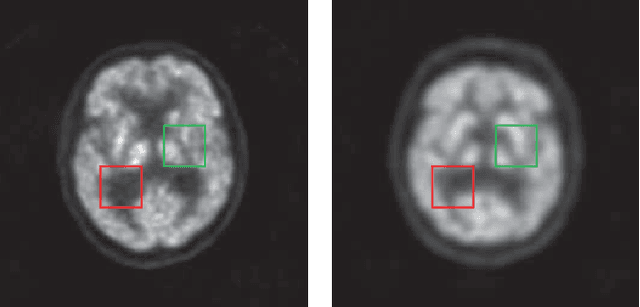
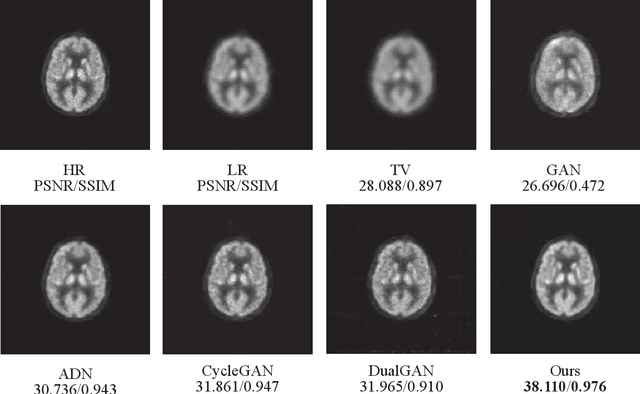
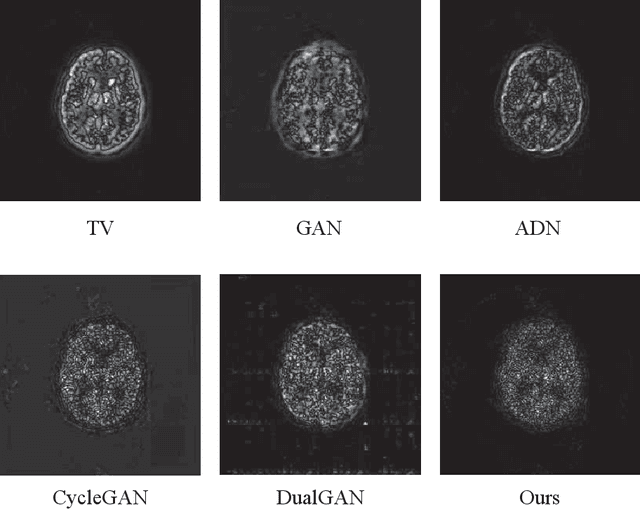
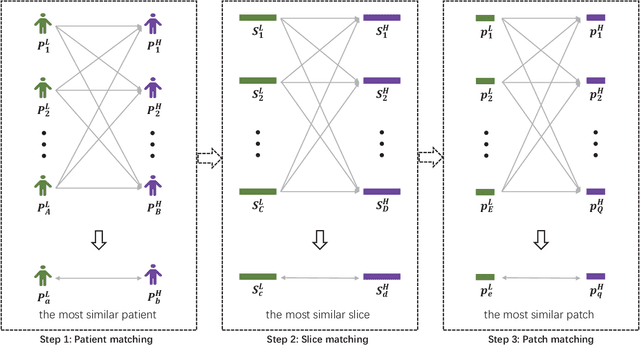
Abstract:Low resolution of positron emission tomography (PET) limits its diagnostic performance. Deep learning has been successfully applied to achieve super-resolution PET. However, commonly used supervised learning methods in this context require many pairs of low- and high-resolution (LR and HR) PET images. Although unsupervised learning utilizes unpaired images, the results are not as good as that obtained with supervised deep learning. In this paper, we propose a quasi-supervised learning method, which is a new type of weakly-supervised learning methods, to recover HR PET images from LR counterparts by leveraging similarity between unpaired LR and HR image patches. Specifically, LR image patches are taken from a patient as inputs, while the most similar HR patches from other patients are found as labels. The similarity between the matched HR and LR patches serves as a prior for network construction. Our proposed method can be implemented by designing a new network or modifying an existing network. As an example in this study, we have modified the cycle-consistent generative adversarial network (CycleGAN) for super-resolution PET. Our numerical and experimental results qualitatively and quantitatively show the merits of our method relative to the state-ofthe-art methods. The code is publicly available at https://github.com/PigYang-ops/CycleGAN-QSDL.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge