Eric Horvitz
Capabilities of GPT-4 on Medical Challenge Problems
Mar 20, 2023



Abstract:Large language models (LLMs) have demonstrated remarkable capabilities in natural language understanding and generation across various domains, including medicine. We present a comprehensive evaluation of GPT-4, a state-of-the-art LLM, on medical competency examinations and benchmark datasets. GPT-4 is a general-purpose model that is not specialized for medical problems through training or engineered to solve clinical tasks. Our analysis covers two sets of official practice materials for the USMLE, a three-step examination program used to assess clinical competency and grant licensure in the United States. We also evaluate performance on the MultiMedQA suite of benchmark datasets. Beyond measuring model performance, experiments were conducted to investigate the influence of test questions containing both text and images on model performance, probe for memorization of content during training, and study probability calibration, which is of critical importance in high-stakes applications like medicine. Our results show that GPT-4, without any specialized prompt crafting, exceeds the passing score on USMLE by over 20 points and outperforms earlier general-purpose models (GPT-3.5) as well as models specifically fine-tuned on medical knowledge (Med-PaLM, a prompt-tuned version of Flan-PaLM 540B). In addition, GPT-4 is significantly better calibrated than GPT-3.5, demonstrating a much-improved ability to predict the likelihood that its answers are correct. We also explore the behavior of the model qualitatively through a case study that shows the ability of GPT-4 to explain medical reasoning, personalize explanations to students, and interactively craft new counterfactual scenarios around a medical case. Implications of the findings are discussed for potential uses of GPT-4 in medical education, assessment, and clinical practice, with appropriate attention to challenges of accuracy and safety.
Benchmarking Spatial Relationships in Text-to-Image Generation
Dec 20, 2022



Abstract:Spatial understanding is a fundamental aspect of computer vision and integral for human-level reasoning about images, making it an important component for grounded language understanding. While recent large-scale text-to-image synthesis (T2I) models have shown unprecedented improvements in photorealism, it is unclear whether they have reliable spatial understanding capabilities. We investigate the ability of T2I models to generate correct spatial relationships among objects and present VISOR, an evaluation metric that captures how accurately the spatial relationship described in text is generated in the image. To benchmark existing models, we introduce a large-scale challenge dataset SR2D that contains sentences describing two objects and the spatial relationship between them. We construct and harness an automated evaluation pipeline that employs computer vision to recognize objects and their spatial relationships, and we employ it in a large-scale evaluation of T2I models. Our experiments reveal a surprising finding that, although recent state-of-the-art T2I models exhibit high image quality, they are severely limited in their ability to generate multiple objects or the specified spatial relations such as left/right/above/below. Our analyses demonstrate several biases and artifacts of T2I models such as the difficulty with generating multiple objects, a bias towards generating the first object mentioned, spatially inconsistent outputs for equivalent relationships, and a correlation between object co-occurrence and spatial understanding capabilities. We conduct a human study that shows the alignment between VISOR and human judgment about spatial understanding. We offer the SR2D dataset and the VISOR metric to the community in support of T2I spatial reasoning research.
Reading Between the Lines: Modeling User Behavior and Costs in AI-Assisted Programming
Oct 25, 2022



Abstract:AI code-recommendation systems (CodeRec), such as Copilot, can assist programmers inside an IDE by suggesting and autocompleting arbitrary code; potentially improving their productivity. To understand how these AI improve programmers in a coding session, we need to understand how they affect programmers' behavior. To make progress, we studied GitHub Copilot, and developed CUPS -- a taxonomy of 12 programmer activities common to AI code completion systems. We then conducted a study with 21 programmers who completed coding tasks and used our labeling tool to retrospectively label their sessions with CUPS. We analyze over 3000 label instances, and visualize the results with timelines and state machines to profile programmer-CodeRec interaction. This reveals novel insights into the distribution and patterns of programmer behavior, as well as inefficiencies and time costs. Finally, we use these insights to inform future interventions to improve AI-assisted programming and human-AI interaction.
On the Horizon: Interactive and Compositional Deepfakes
Sep 21, 2022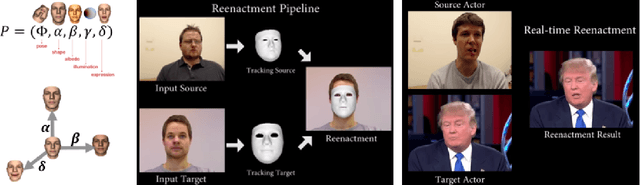
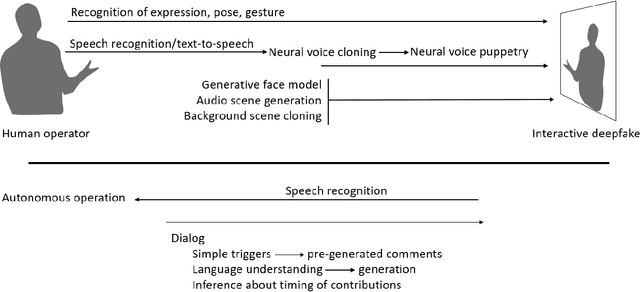
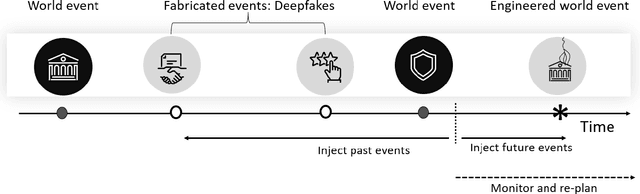
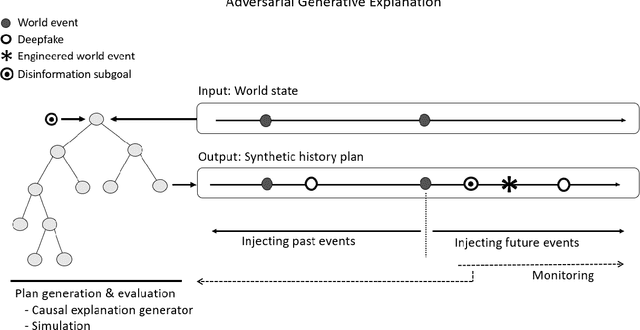
Abstract:Over a five-year period, computing methods for generating high-fidelity, fictional depictions of people and events moved from exotic demonstrations by computer science research teams into ongoing use as a tool of disinformation. The methods, referred to with the portmanteau of "deepfakes," have been used to create compelling audiovisual content. Here, I share challenges ahead with malevolent uses of two classes of deepfakes that we can expect to come into practice with costly implications for society: interactive and compositional deepfakes. Interactive deepfakes have the capability to impersonate people with realistic interactive behaviors, taking advantage of advances in multimodal interaction. Compositional deepfakes leverage synthetic content in larger disinformation plans that integrate sets of deepfakes over time with observed, expected, and engineered world events to create persuasive synthetic histories. Synthetic histories can be constructed manually but may one day be guided by adversarial generative explanation (AGE) techniques. In the absence of mitigations, interactive and compositional deepfakes threaten to move us closer to a post-epistemic world, where fact cannot be distinguished from fiction. I shall describe interactive and compositional deepfakes and reflect about cautions and potential mitigations to defend against them.
Who Goes First? Influences of Human-AI Workflow on Decision Making in Clinical Imaging
May 19, 2022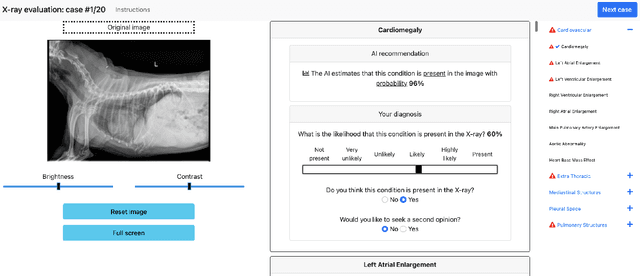
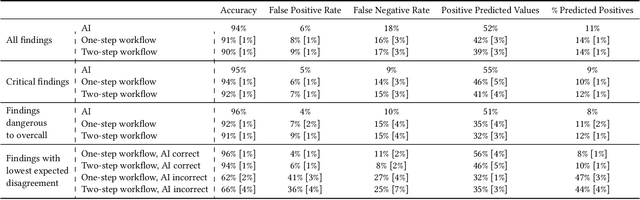
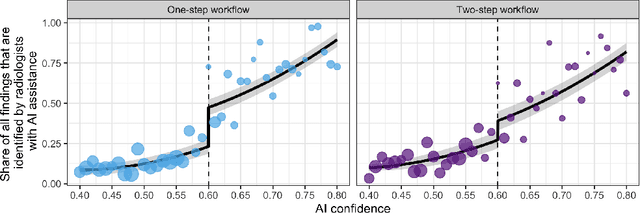

Abstract:Details of the designs and mechanisms in support of human-AI collaboration must be considered in the real-world fielding of AI technologies. A critical aspect of interaction design for AI-assisted human decision making are policies about the display and sequencing of AI inferences within larger decision-making workflows. We have a poor understanding of the influences of making AI inferences available before versus after human review of a diagnostic task at hand. We explore the effects of providing AI assistance at the start of a diagnostic session in radiology versus after the radiologist has made a provisional decision. We conducted a user study where 19 veterinary radiologists identified radiographic findings present in patients' X-ray images, with the aid of an AI tool. We employed two workflow configurations to analyze (i) anchoring effects, (ii) human-AI team diagnostic performance and agreement, (iii) time spent and confidence in decision making, and (iv) perceived usefulness of the AI. We found that participants who are asked to register provisional responses in advance of reviewing AI inferences are less likely to agree with the AI regardless of whether the advice is accurate and, in instances of disagreement with the AI, are less likely to seek the second opinion of a colleague. These participants also reported the AI advice to be less useful. Surprisingly, requiring provisional decisions on cases in advance of the display of AI inferences did not lengthen the time participants spent on the task. The study provides generalizable and actionable insights for the deployment of clinical AI tools in human-in-the-loop systems and introduces a methodology for studying alternative designs for human-AI collaboration. We make our experimental platform available as open source to facilitate future research on the influence of alternate designs on human-AI workflows.
A Computational Inflection for Scientific Discovery
May 04, 2022
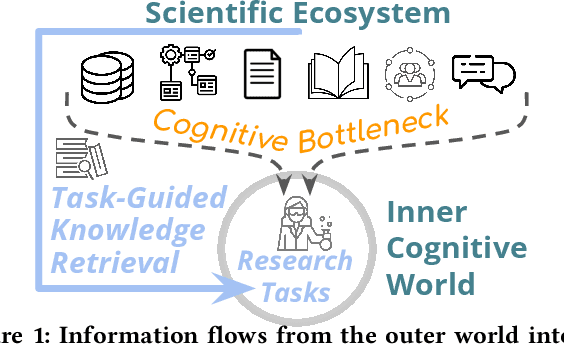
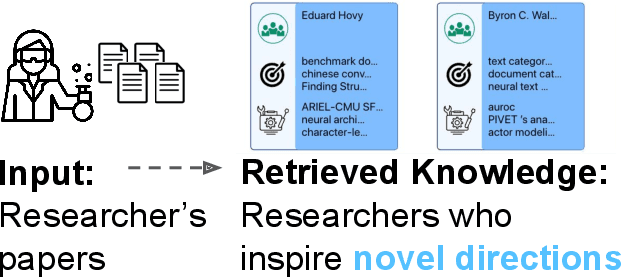
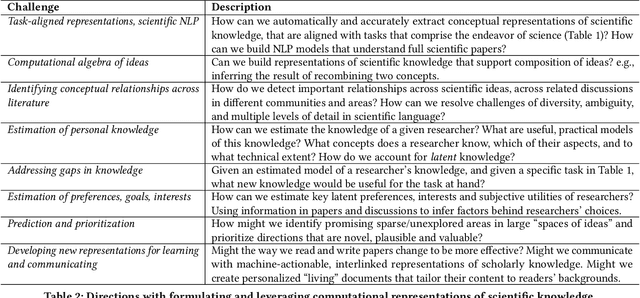
Abstract:We stand at the foot of a significant inflection in the trajectory of scientific discovery. As society continues on its fast-paced digital transformation, so does humankind's collective scientific knowledge and discourse. We now read and write papers in digitized form, and a great deal of the formal and informal processes of science are captured digitally -- including papers, preprints and books, code and datasets, conference presentations, and interactions in social networks and communication platforms. The transition has led to the growth of a tremendous amount of information, opening exciting opportunities for computational models and systems that analyze and harness it. In parallel, exponential growth in data processing power has fueled remarkable advances in AI, including self-supervised neural models capable of learning powerful representations from large-scale unstructured text without costly human supervision. The confluence of societal and computational trends suggests that computer science is poised to ignite a revolution in the scientific process itself. However, the explosion of scientific data, results and publications stands in stark contrast to the constancy of human cognitive capacity. While scientific knowledge is expanding with rapidity, our minds have remained static, with severe limitations on the capacity for finding, assimilating and manipulating information. We propose a research agenda of task-guided knowledge retrieval, in which systems counter humans' bounded capacity by ingesting corpora of scientific knowledge and retrieving inspirations, explanations, solutions and evidence synthesized to directly augment human performance on salient tasks in scientific endeavors. We present initial progress on methods and prototypes, and lay out important opportunities and challenges ahead with computational approaches that have the potential to revolutionize science.
Computational Lens on Cognition: Study Of Autobiographical Versus Imagined Stories With Large-Scale Language Models
Jan 07, 2022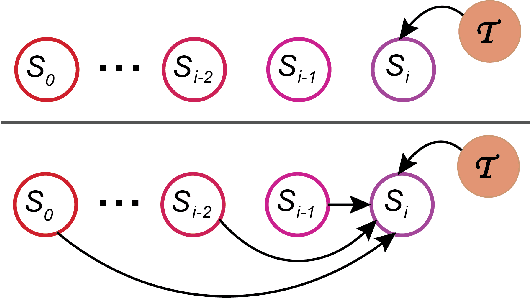
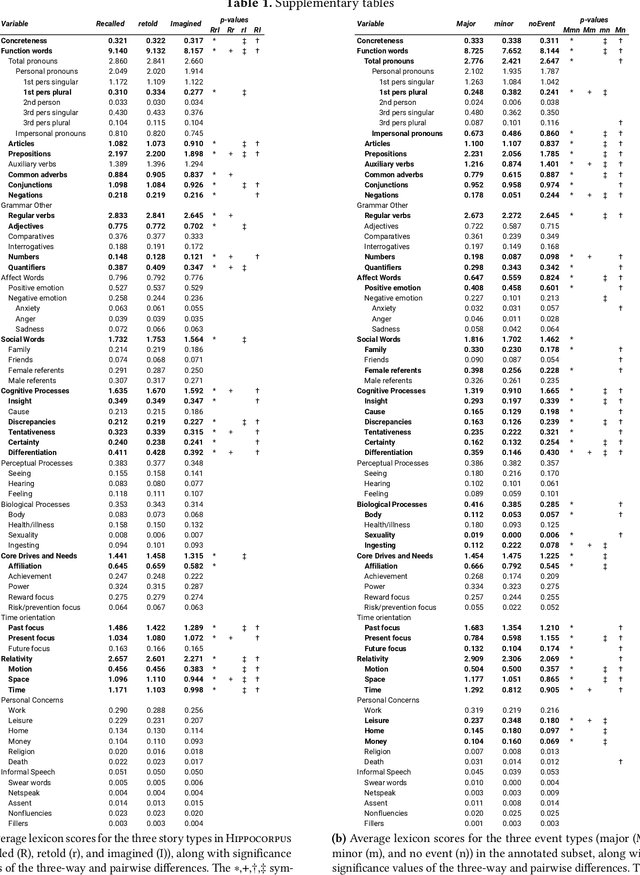
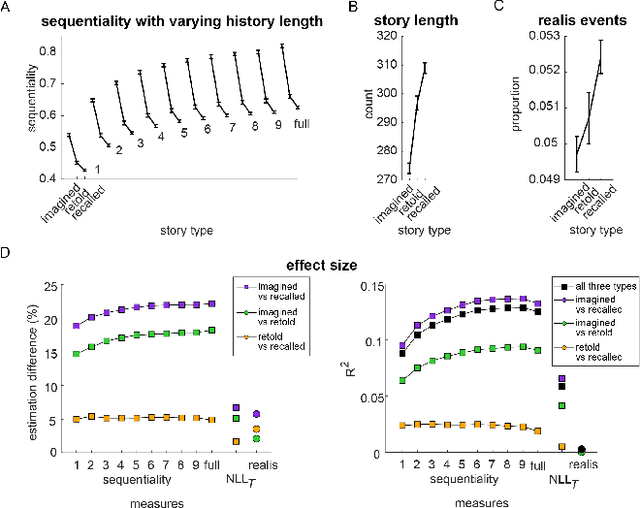
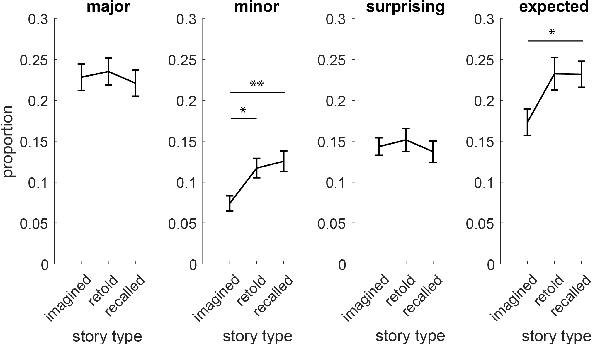
Abstract:Lifelong experiences and learned knowledge lead to shared expectations about how common situations tend to unfold. Such knowledge enables people to interpret story narratives and identify salient events effortlessly. We study differences in the narrative flow of events in autobiographical versus imagined stories using GPT-3, one of the largest neural language models created to date. The diary-like stories were written by crowdworkers about either a recently experienced event or an imagined event on the same topic. To analyze the narrative flow of events of these stories, we measured sentence *sequentiality*, which compares the probability of a sentence with and without its preceding story context. We found that imagined stories have higher sequentiality than autobiographical stories, and that the sequentiality of autobiographical stories is higher when they are retold than when freshly recalled. Through an annotation of events in story sentences, we found that the story types contain similar proportions of major salient events, but that the autobiographical stories are denser in factual minor events. Furthermore, in comparison to imagined stories, autobiographical stories contain more concrete words and words related to the first person, cognitive processes, time, space, numbers, social words, and core drives and needs. Our findings highlight the opportunity to investigate memory and cognition with large-scale statistical language models.
Ideal Partition of Resources for Metareasoning
Oct 18, 2021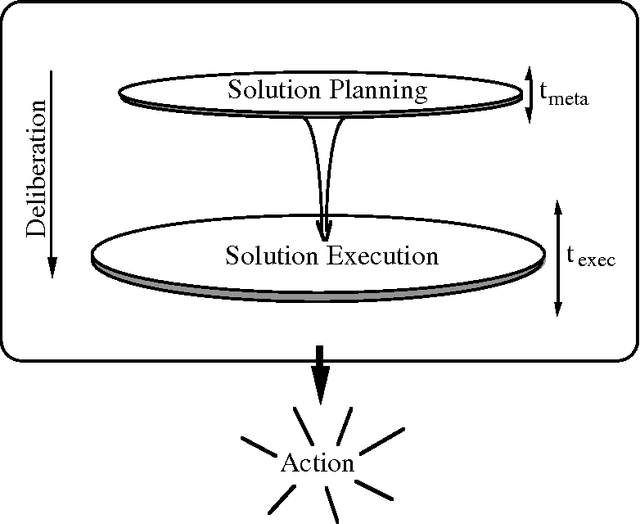
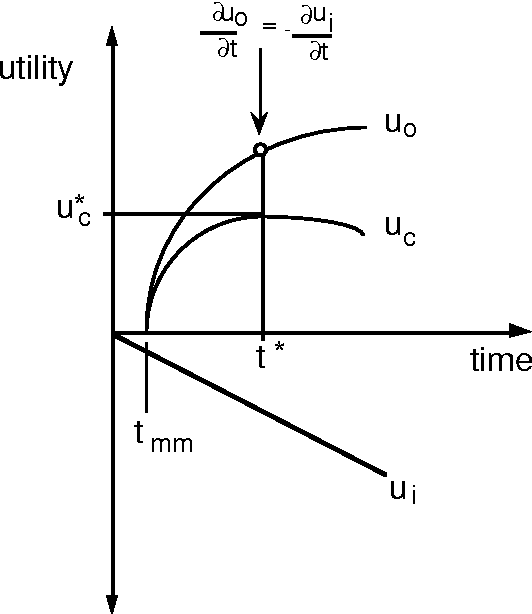
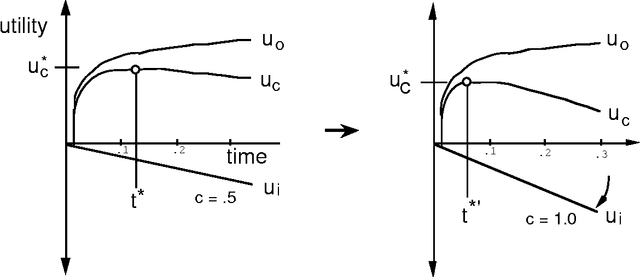
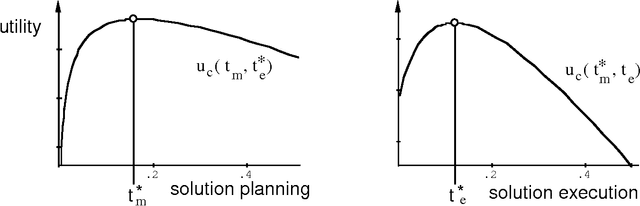
Abstract:We can achieve significant gains in the value of computation by metareasoning about the nature or extent of base-level problem solving before executing a solution. However, resources that are irrevocably committed to metareasoning are not available for executing a solution. Thus, it is important to determine the portion of resources we wish to apply to metareasoning and control versus to the execution of a solution plan. Recent research on rational agency has highlighted the importance of limiting the consumption of resources by metareasoning machinery. We shall introduce the metareasoning-partition problem--the problem of ideally apportioning costly reasoning resources to planning a solution versus applying resource to executing a solution to a problem. We exercise prototypical metareasoning-partition models to probe the relationships between time allocated to metareasoning and to execution for different problem classes. Finally, we examine the value of metareasoning in the context of our functional analyses.
Bursting Scientific Filter Bubbles: Boosting Innovation via Novel Author Discovery
Sep 10, 2021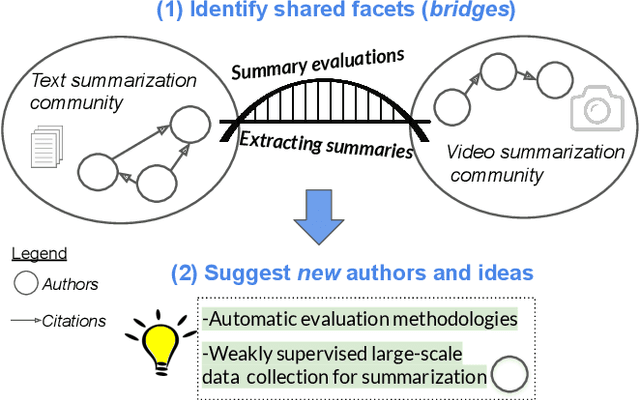
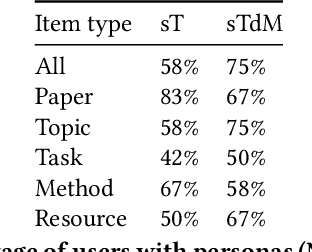
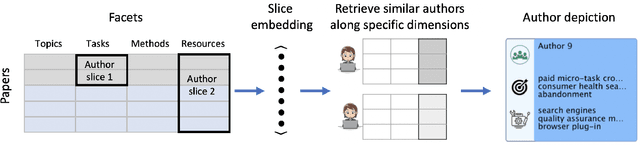
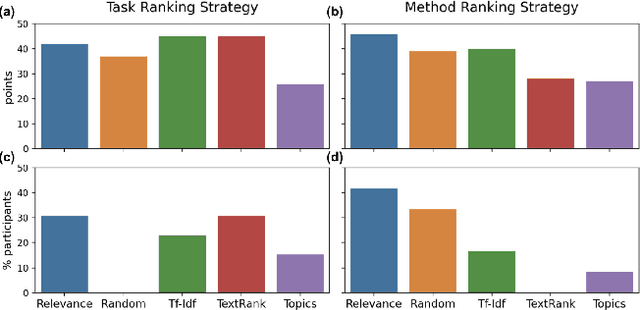
Abstract:Isolated silos of scientific research and the growing challenge of information overload limit awareness across the literature and hinder innovation. Algorithmic curation and recommendation, which often prioritize relevance, can further reinforce these informational "filter bubbles." In response, we describe Bridger, a system for facilitating discovery of scholars and their work, to explore design tradeoffs between relevant and novel recommendations. We construct a faceted representation of authors with information gleaned from their papers and inferred author personas, and use it to develop an approach that locates commonalities ("bridges") and contrasts between scientists -- retrieving partially similar authors rather than aiming for strict similarity. In studies with computer science researchers, this approach helps users discover authors considered useful for generating novel research directions, outperforming a state-of-art neural model. In addition to recommending new content, we also demonstrate an approach for displaying it in a manner that boosts researchers' ability to understand the work of authors with whom they are unfamiliar. Finally, our analysis reveals that Bridger connects authors who have different citation profiles, publish in different venues, and are more distant in social co-authorship networks, raising the prospect of bridging diverse communities and facilitating discovery.
A Search Engine for Discovery of Scientific Challenges and Directions
Sep 10, 2021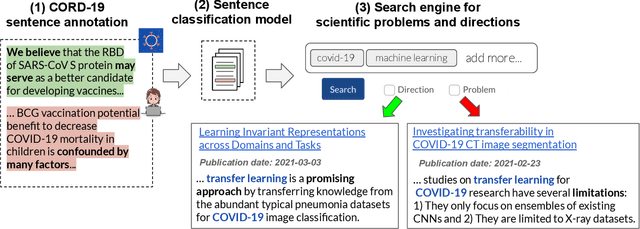


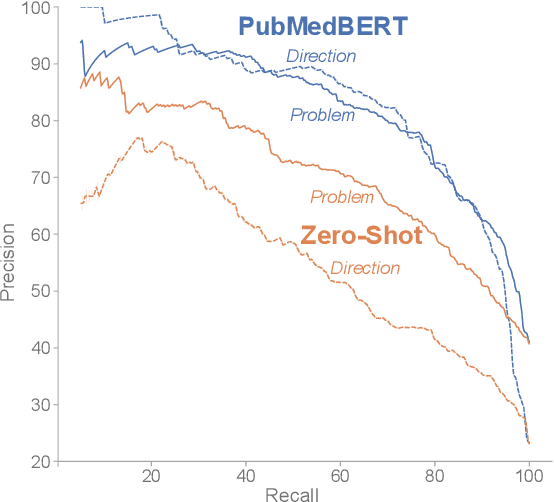
Abstract:Keeping track of scientific challenges, advances and emerging directions is a fundamental part of research. However, researchers face a flood of papers that hinders discovery of important knowledge. In biomedicine, this directly impacts human lives. To address this problem, we present a novel task of extraction and search of scientific challenges and directions, to facilitate rapid knowledge discovery. We construct and release an expert-annotated corpus of texts sampled from full-length papers, labeled with novel semantic categories that generalize across many types of challenges and directions. We focus on a large corpus of interdisciplinary work relating to the COVID-19 pandemic, ranging from biomedicine to areas such as AI and economics. We apply a model trained on our data to identify challenges and directions across the corpus and build a dedicated search engine. In experiments with 19 researchers and clinicians using our system, we outperform a popular scientific search engine in assisting knowledge discovery. Finally, we show that models trained on our resource generalize to the wider biomedical domain and to AI papers, highlighting its broad utility. We make our data, model and search engine publicly available. https://challenges.apps.allenai.org/
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge