Jevin West
Temporal Narrative Monitoring in Dynamic Information Environments
Mar 18, 2026Abstract:Comprehending the information environment (IE) during crisis events is challenging due to the rapid change and abstract nature of the domain. Many approaches focus on snapshots via classification methods or network approaches to describe the IE in crisis, ignoring the temporal nature of how information changed over time. This work presents a system-oriented framework for modeling emerging narratives as temporally evolving semantic structures without requiring prior label specification. By integrating semantic embeddings, density-based clustering, and rolling temporal linkage, the framework represents narratives as persistent yet adaptive entities within a shared semantic space. We apply the methodology to a real-world crisis event and evaluate system behavior through stratified cluster validation and temporal lifecycle analysis. Results demonstrate high cluster coherence and reveal heterogeneous narrative lifecycles characterized by both transient fragments and stable narrative anchors. We ground our approach in situational awareness theory, supporting perception and comprehension of the IE by transforming unstructured social media streams into interpretable, temporally structured representations. The resulting system provides a methodology for monitoring and decision support in dynamic information environments.
LLM Chain Ensembles for Scalable and Accurate Data Annotation
Oct 16, 2024



Abstract:The ability of large language models (LLMs) to perform zero-shot classification makes them viable solutions for data annotation in rapidly evolving domains where quality labeled data is often scarce and costly to obtain. However, the large-scale deployment of LLMs can be prohibitively expensive. This paper introduces an LLM chain ensemble methodology that aligns multiple LLMs in a sequence, routing data subsets to subsequent models based on classification uncertainty. This approach leverages the strengths of individual LLMs within a broader system, allowing each model to handle data points where it exhibits the highest confidence, while forwarding more complex cases to potentially more robust models. Our results show that the chain ensemble method often exceeds the performance of the best individual model in the chain and achieves substantial cost savings, making LLM chain ensembles a practical and efficient solution for large-scale data annotation challenges.
LLM Confidence Evaluation Measures in Zero-Shot CSS Classification
Oct 16, 2024

Abstract:Assessing classification confidence is critical for leveraging large language models (LLMs) in automated labeling tasks, especially in the sensitive domains presented by Computational Social Science (CSS) tasks. In this paper, we make three key contributions: (1) we propose an uncertainty quantification (UQ) performance measure tailored for data annotation tasks, (2) we compare, for the first time, five different UQ strategies across three distinct LLMs and CSS data annotation tasks, (3) we introduce a novel UQ aggregation strategy that effectively identifies low-confidence LLM annotations and disproportionately uncovers data incorrectly labeled by the LLMs. Our results demonstrate that our proposed UQ aggregation strategy improves upon existing methods andcan be used to significantly improve human-in-the-loop data annotation processes.
RED-CT: A Systems Design Methodology for Using LLM-labeled Data to Train and Deploy Edge Classifiers for Computational Social Science
Aug 15, 2024Abstract:Large language models (LLMs) have enhanced our ability to rapidly analyze and classify unstructured natural language data. However, concerns regarding cost, network limitations, and security constraints have posed challenges for their integration into work processes. In this study, we adopt a systems design approach to employing LLMs as imperfect data annotators for downstream supervised learning tasks, introducing novel system intervention measures aimed at improving classification performance. Our methodology outperforms LLM-generated labels in seven of eight tests, demonstrating an effective strategy for incorporating LLMs into the design and deployment of specialized, supervised learning models present in many industry use cases.
Response: Emergent analogical reasoning in large language models
Aug 30, 2023



Abstract:In their recent Nature Human Behaviour paper, "Emergent analogical reasoning in large language models," (Webb, Holyoak, and Lu, 2023) the authors argue that "large language models such as GPT-3 have acquired an emergent ability to find zero-shot solutions to a broad range of analogy problems." In this response, we provide counterexamples of the letter string analogies. In our tests, GPT-3 fails to solve even the easiest variants of the problems presented in the original paper. Zero-shot reasoning is an extraordinary claim that requires extraordinary evidence. We do not see that evidence in our experiments. To strengthen claims of humanlike reasoning such as zero-shot reasoning, it is important that the field develop approaches that rule out data memorization.
Bursting Scientific Filter Bubbles: Boosting Innovation via Novel Author Discovery
Sep 10, 2021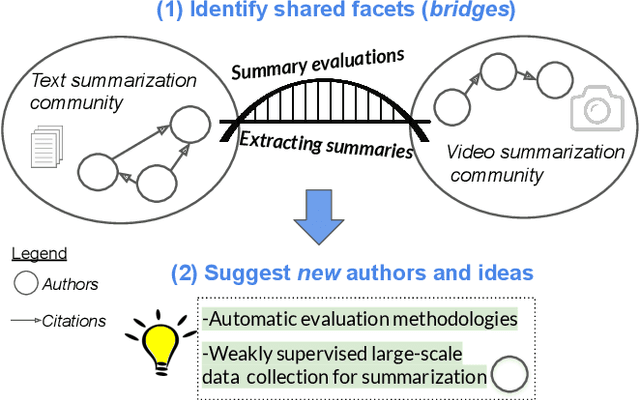
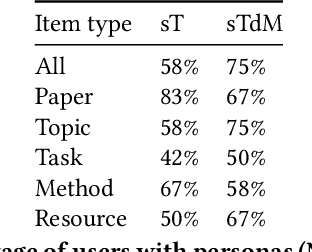
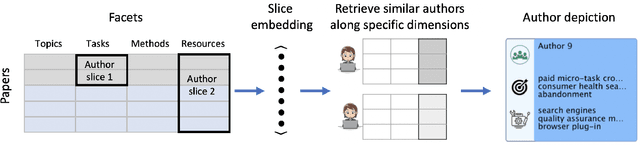
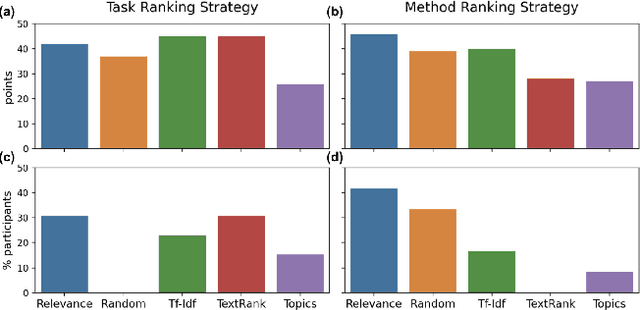
Abstract:Isolated silos of scientific research and the growing challenge of information overload limit awareness across the literature and hinder innovation. Algorithmic curation and recommendation, which often prioritize relevance, can further reinforce these informational "filter bubbles." In response, we describe Bridger, a system for facilitating discovery of scholars and their work, to explore design tradeoffs between relevant and novel recommendations. We construct a faceted representation of authors with information gleaned from their papers and inferred author personas, and use it to develop an approach that locates commonalities ("bridges") and contrasts between scientists -- retrieving partially similar authors rather than aiming for strict similarity. In studies with computer science researchers, this approach helps users discover authors considered useful for generating novel research directions, outperforming a state-of-art neural model. In addition to recommending new content, we also demonstrate an approach for displaying it in a manner that boosts researchers' ability to understand the work of authors with whom they are unfamiliar. Finally, our analysis reveals that Bridger connects authors who have different citation profiles, publish in different venues, and are more distant in social co-authorship networks, raising the prospect of bridging diverse communities and facilitating discovery.
SciSight: Combining faceted navigation and research group detection for COVID-19 exploratory scientific search
May 27, 2020
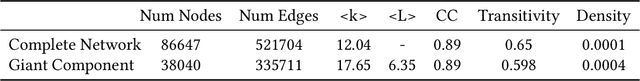
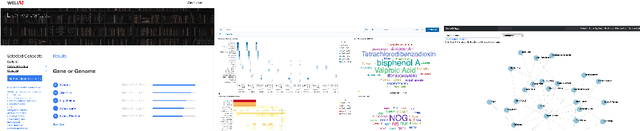
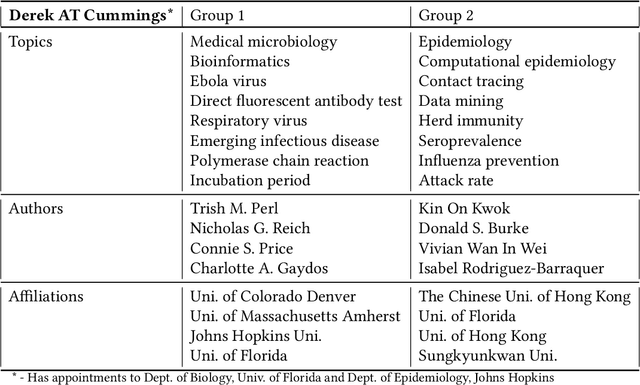
Abstract:The COVID-19 pandemic has sparked unprecedented mobilization of scientists, already generating thousands of new papers that join a litany of previous biomedical work in related areas. This deluge of information makes it hard for researchers to keep track of their own research area, let alone explore new directions. Standard search engines are designed primarily for targeted search and are not geared for discovery or making connections that are not obvious from reading individual papers. In this paper, we present our ongoing work on SciSight, a novel framework for exploratory search of COVID-19 research. Based on formative interviews with scientists and a review of existing tools, we build and integrate two key capabilities: first, exploring interactions between biomedical facets (e.g., proteins, genes, drugs, diseases, patient characteristics); and second, discovering groups of researchers and how they are connected. We extract entities using a language model pre-trained on several biomedical information extraction tasks, and enrich them with data from the Microsoft Academic Graph (MAG). To find research groups automatically, we use hierarchical clustering with overlap to allow authors, as they do, to belong to multiple groups. Finally, we introduce a novel presentation of these groups based on both topical and social affinities, allowing users to drill down from groups to papers to associations between entities, and update query suggestions on the fly with the goal of facilitating exploratory navigation. SciSight has thus far served over 10K users with over 30K page views and 13% returning users. Preliminary user interviews with biomedical researchers suggest that SciSight complements current approaches and helps find new and relevant knowledge.
Stem-ming the Tide: Predicting STEM attrition using student transcript data
Aug 28, 2017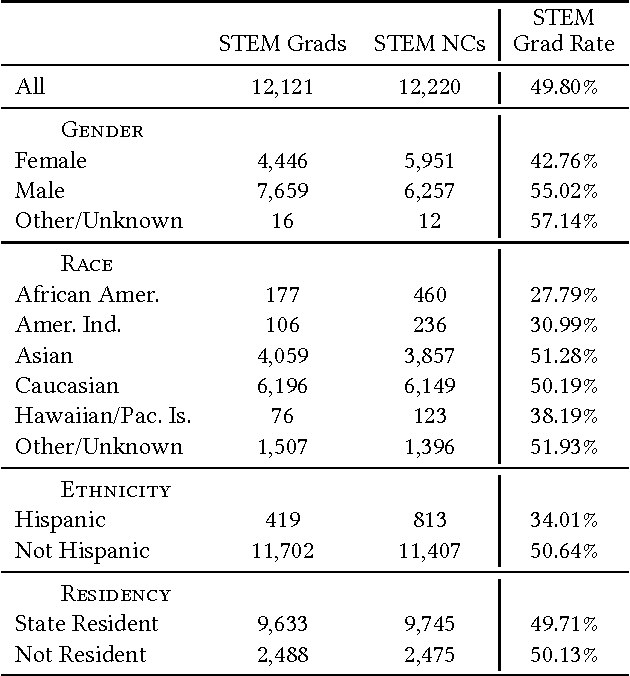
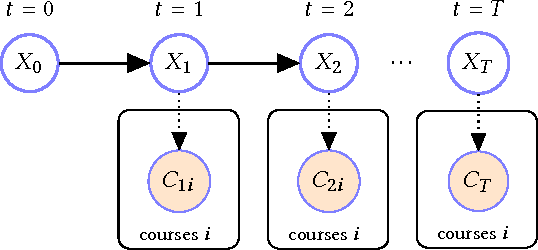
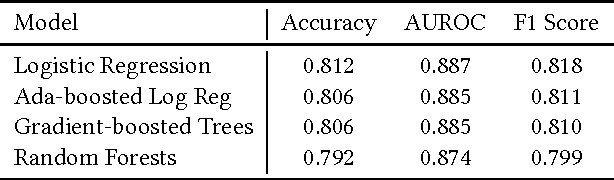
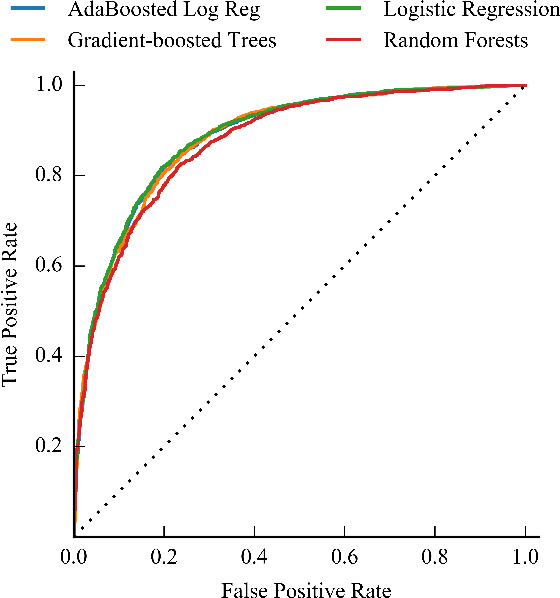
Abstract:Science, technology, engineering, and math (STEM) fields play growing roles in national and international economies by driving innovation and generating high salary jobs. Yet, the US is lagging behind other highly industrialized nations in terms of STEM education and training. Furthermore, many economic forecasts predict a rising shortage of domestic STEM-trained professions in the US for years to come. One potential solution to this deficit is to decrease the rates at which students leave STEM-related fields in higher education, as currently over half of all students intending to graduate with a STEM degree eventually attrite. However, little quantitative research at scale has looked at causes of STEM attrition, let alone the use of machine learning to examine how well this phenomenon can be predicted. In this paper, we detail our efforts to model and predict dropout from STEM fields using one of the largest known datasets used for research on students at a traditional campus setting. Our results suggest that attrition from STEM fields can be accurately predicted with data that is routinely collected at universities using only information on students' first academic year. We also propose a method to model student STEM intentions for each academic term to better understand the timing of STEM attrition events. We believe these results show great promise in using machine learning to improve STEM retention in traditional and non-traditional campus settings.
Predicting Student Dropout in Higher Education
Mar 07, 2017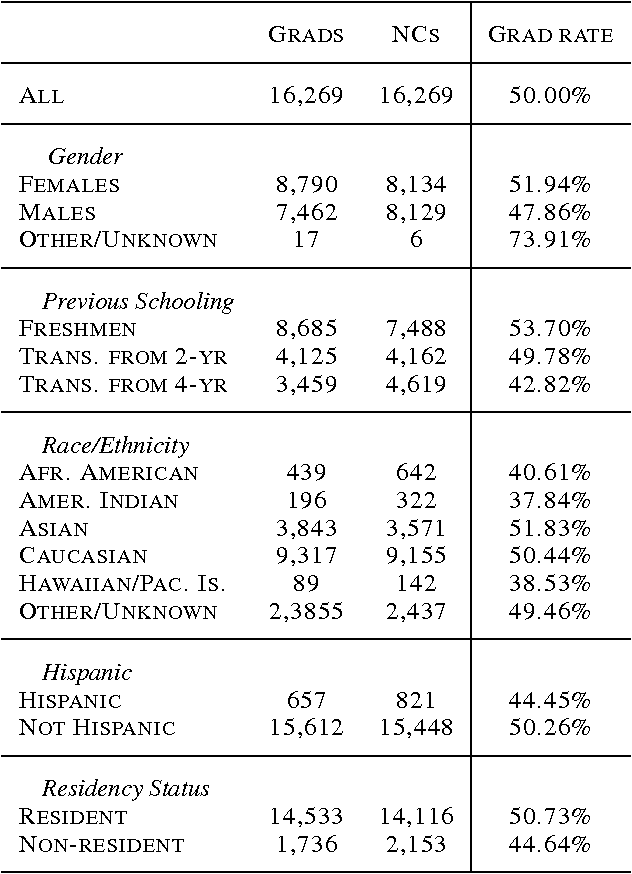
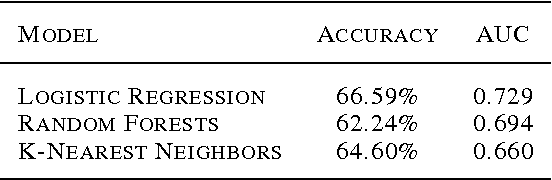
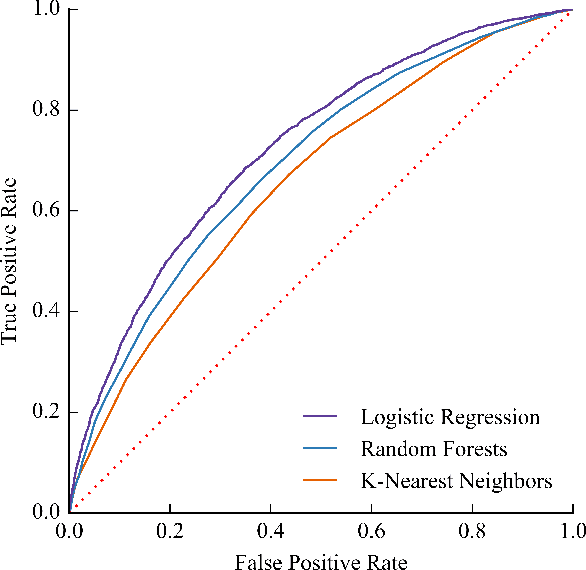

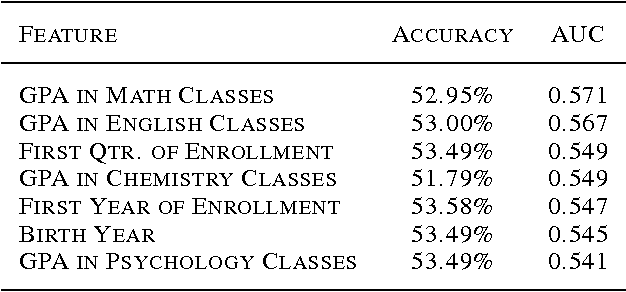
Abstract:Each year, roughly 30% of first-year students at US baccalaureate institutions do not return for their second year and over $9 billion is spent educating these students. Yet, little quantitative research has analyzed the causes and possible remedies for student attrition. Here, we describe initial efforts to model student dropout using the largest known dataset on higher education attrition, which tracks over 32,500 students' demographics and transcript records at one of the nation's largest public universities. Our results highlight several early indicators of student attrition and show that dropout can be accurately predicted even when predictions are based on a single term of academic transcript data. These results highlight the potential for machine learning to have an impact on student retention and success while pointing to several promising directions for future work.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge