Debdoot Sheet
A Systematic Search over Deep Convolutional Neural Network Architectures for Screening Chest Radiographs
Apr 24, 2020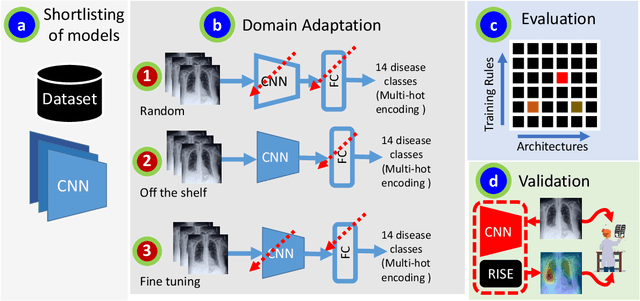
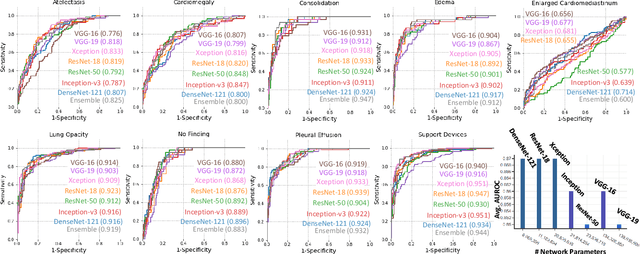
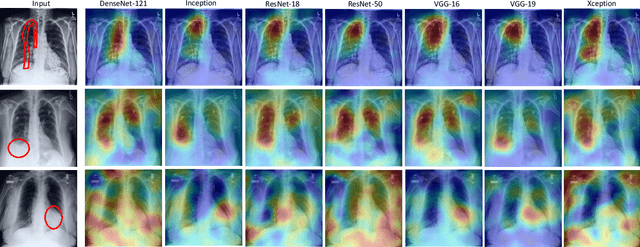
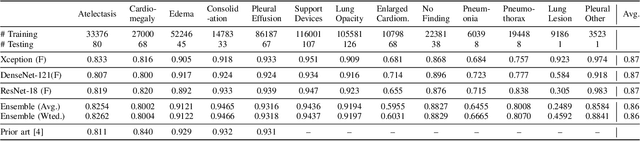
Abstract:Chest radiographs are primarily employed for the screening of pulmonary and cardio-/thoracic conditions. Being undertaken at primary healthcare centers, they require the presence of an on-premise reporting Radiologist, which is a challenge in low and middle income countries. This has inspired the development of machine learning based automation of the screening process. While recent efforts demonstrate a performance benchmark using an ensemble of deep convolutional neural networks (CNN), our systematic search over multiple standard CNN architectures identified single candidate CNN models whose classification performances were found to be at par with ensembles. Over 63 experiments spanning 400 hours, executed on a 11:3 FP32 TensorTFLOPS compute system, we found the Xception and ResNet-18 architectures to be consistent performers in identifying co-existing disease conditions with an average AUC of 0.87 across nine pathologies. We conclude on the reliability of the models by assessing their saliency maps generated using the randomized input sampling for explanation (RISE) method and qualitatively validating them against manual annotations locally sourced from an experienced Radiologist. We also draw a critical note on the limitations of the publicly available CheXpert dataset primarily on account of disparity in class distribution in training vs. testing sets, and unavailability of sufficient samples for few classes, which hampers quantitative reporting due to sample insufficiency.
CHAOS Challenge -- Combined (CT-MR) Healthy Abdominal Organ Segmentation
Jan 17, 2020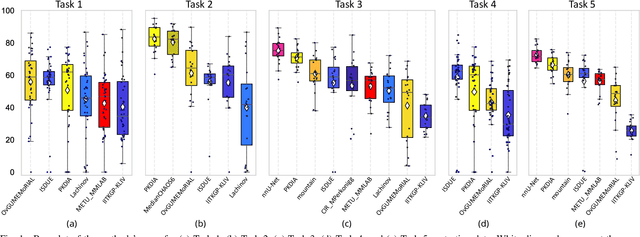
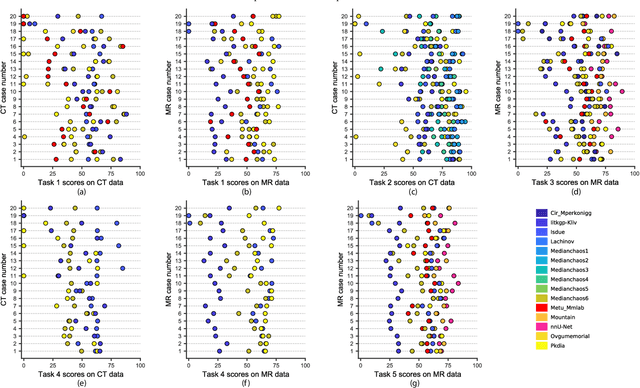
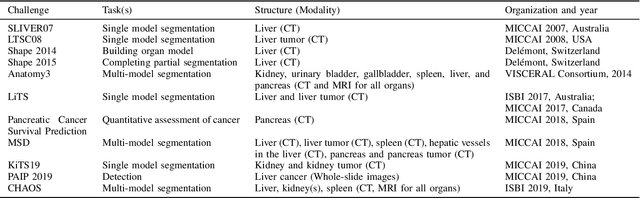
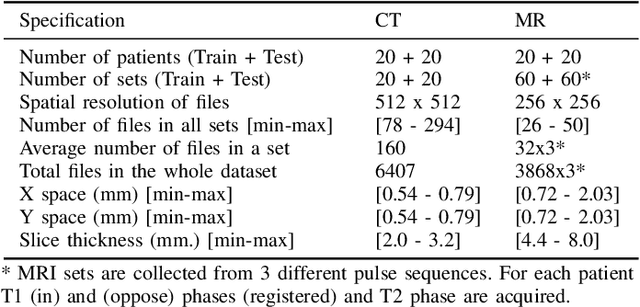
Abstract:Segmentation of abdominal organs has been a comprehensive, yet unresolved, research field for many years. In the last decade, intensive developments in deep learning (DL) have introduced new state-of-the-art segmentation systems. Despite outperforming the overall accuracy of existing systems, the effects of DL model properties and parameters on the performance is hard to interpret. This makes comparative analysis a necessary tool to achieve explainable studies and systems. Moreover, the performance of DL for emerging learning approaches such as cross-modality and multi-modal tasks have been rarely discussed. In order to expand the knowledge in these topics, CHAOS -- Combined (CT-MR) Healthy Abdominal Organ Segmentation challenge has been organized in the IEEE International Symposium on Biomedical Imaging (ISBI), 2019, in Venice, Italy. Despite a large number of the previous abdomen related challenges, the majority of which are focused on tumor/lesion detection and/or classification with a single modality, CHAOS provides both abdominal CT and MR data from healthy subjects. Five different and complementary tasks have been designed to analyze the capabilities of the current approaches from multiple perspectives. The results are investigated thoroughly, compared with manual annotations and interactive methods. The outcomes are reported in detail to reflect the latest advancements in the field. CHAOS challenge and data will be available online to provide a continuous benchmark resource for segmentation.
Adversarially Trained Convolutional Neural Networks for Semantic Segmentation of Ischaemic Stroke Lesion using Multisequence Magnetic Resonance Imaging
Aug 03, 2019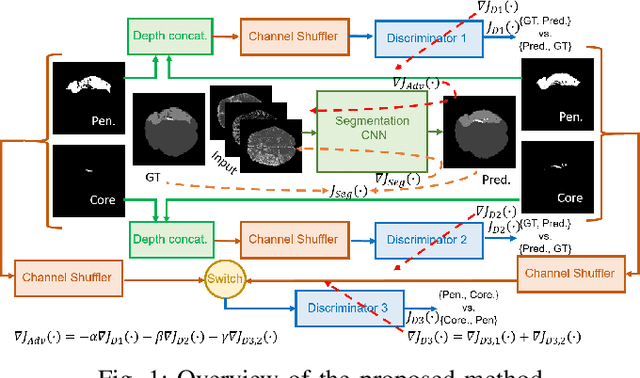
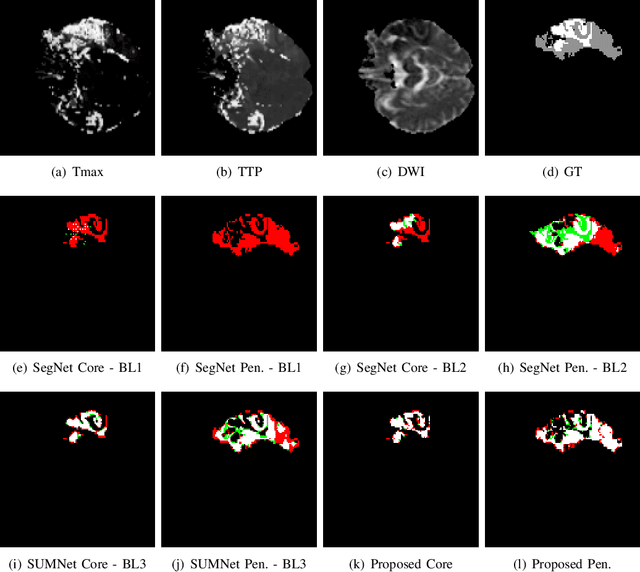
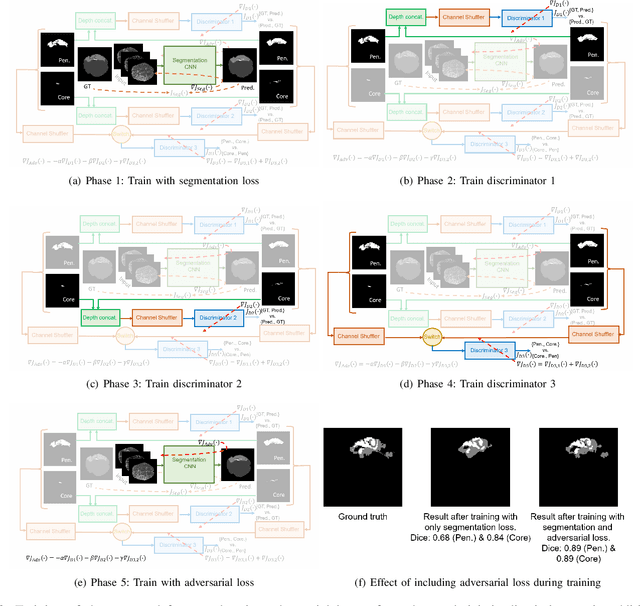

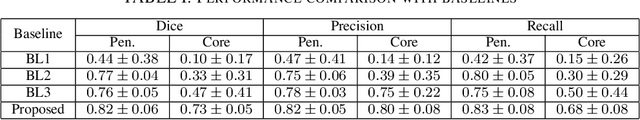
Abstract:Ischaemic stroke is a medical condition caused by occlusion of blood supply to the brain tissue thus forming a lesion. A lesion is zoned into a core associated with irreversible necrosis typically located at the center of the lesion, while reversible hypoxic changes in the outer regions of the lesion are termed as the penumbra. Early estimation of core and penumbra in ischaemic stroke is crucial for timely intervention with thrombolytic therapy to reverse the damage and restore normalcy. Multisequence magnetic resonance imaging (MRI) is commonly employed for clinical diagnosis. However, a sequence singly has not been found to be sufficiently able to differentiate between core and penumbra, while a combination of sequences is required to determine the extent of the damage. The challenge, however, is that with an increase in the number of sequences, it cognitively taxes the clinician to discover symptomatic biomarkers in these images. In this paper, we present a data-driven fully automated method for estimation of core and penumbra in ischaemic lesions using diffusion-weighted imaging (DWI) and perfusion-weighted imaging (PWI) sequence maps of MRI. The method employs recent developments in convolutional neural networks (CNN) for semantic segmentation in medical images. In the absence of availability of a large amount of labeled data, the CNN is trained using an adversarial approach employing cross-entropy as a segmentation loss along with losses aggregated from three discriminators of which two employ relativistic visual Turing test. This method is experimentally validated on the ISLES-2015 dataset through three-fold cross-validation to obtain with an average Dice score of 0.82 and 0.73 for segmentation of penumbra and core respectively.
Adversarially Trained Deep Neural Semantic Hashing Scheme for Subjective Search in Fashion Inventory
Jun 30, 2019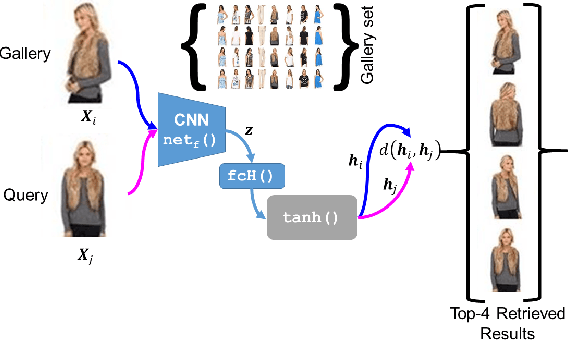
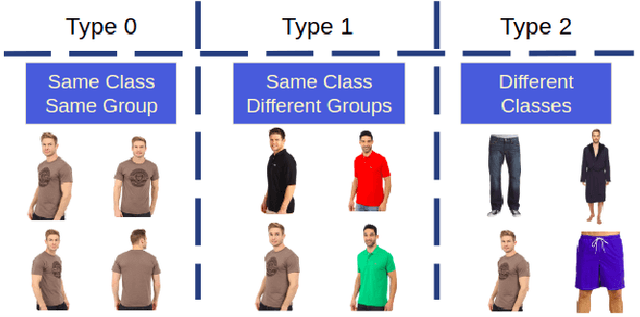



Abstract:The simple approach of retrieving a closest match of a query image from one in the gallery, compares an image pair using sum of absolute difference in pixel or feature space. The process is computationally expensive, ill-posed to illumination, background composition, pose variation, as well as inefficient to be deployed on gallery sets with more than 1000 elements. Hashing is a faster alternative which involves representing images in reduced dimensional simple feature spaces. Encoding images into binary hash codes enables similarity comparison in an image-pair using the Hamming distance measure. The challenge, however, lies in encoding the images using a semantic hashing scheme that lets subjective neighbors lie within the tolerable Hamming radius. This work presents a solution employing adversarial learning of a deep neural semantic hashing network for fashion inventory retrieval. It consists of a feature extracting convolutional neural network (CNN) learned to (i) minimize error in classifying type of clothing, (ii) minimize hamming distance between semantic neighbors and maximize distance between semantically dissimilar images, (iii) maximally scramble a discriminator's ability to identify the corresponding hash code-image pair when processing a semantically similar query-gallery image pair. Experimental validation for fashion inventory search yields a mean average precision (mAP) of 90.65% in finding the closest match as compared to 53.26% obtained by the prior art of deep Cauchy hashing for hamming space retrieval.
Unit Impulse Response as an Explainer of Redundancy in a Deep Convolutional Neural Network
Jun 10, 2019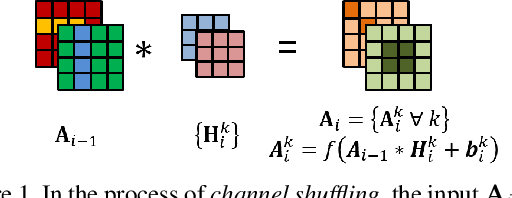
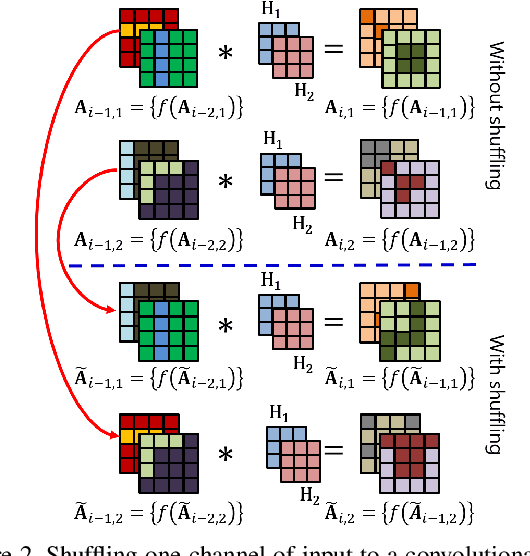
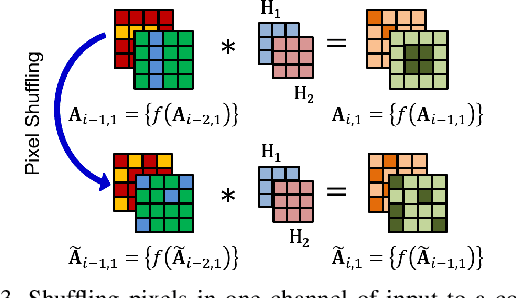
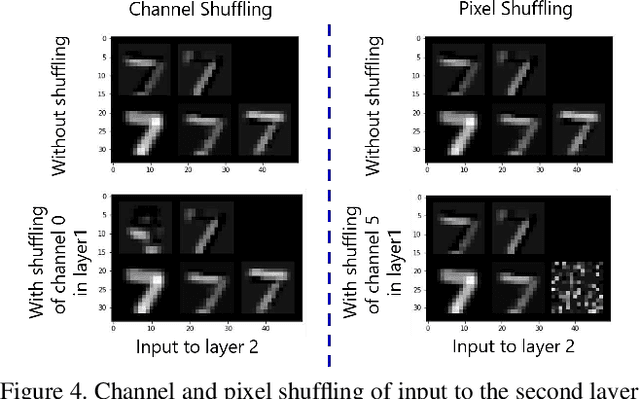
Abstract:Convolutional neural networks (CNN) are generally designed with a heuristic initialization of network architecture and trained for a certain task. This often leads to overparametrization after learning and induces redundancy in the information flow paths within the network. This robustness and reliability is at the increased cost of redundant computations. Several methods have been proposed which leverage metrics that quantify the redundancy in each layer. However, layer-wise evaluation in these methods disregards the long-range redundancy which exists across depth on account of the distributed nature of the features learned by the model. In this paper, we propose (i) a mechanism to empirically demonstrate the robustness in performance of a CNN on account of redundancy across its depth, (ii) a method to identify the systemic redundancy in response of a CNN across depth using the understanding of unit impulse response, we subsequently demonstrate use of these methods to interpret redundancy in few networks as example. These techniques provide better insights into the internal dynamics of a CNN
Multitask Learning of Temporal Connectionism in Convolutional Networks using a Joint Distribution Loss Function to Simultaneously Identify Tools and Phase in Surgical Videos
May 25, 2019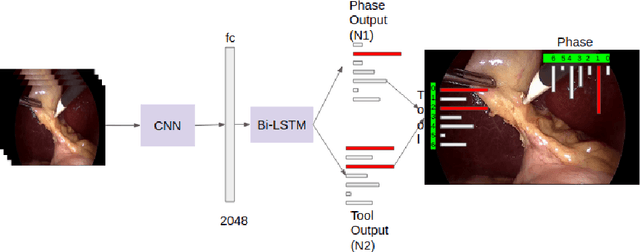
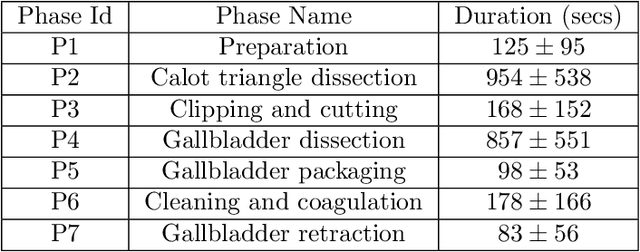
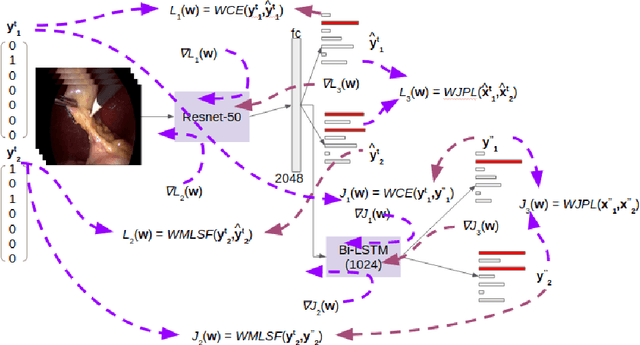
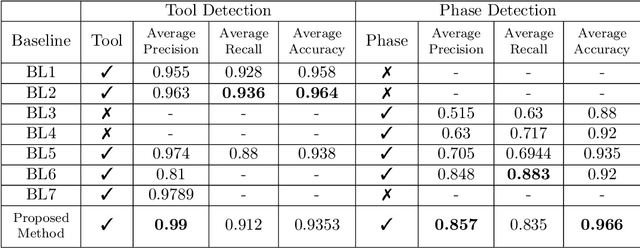
Abstract:Surgical workflow analysis is of importance for understanding onset and persistence of surgical phases and individual tool usage across surgery and in each phase. It is beneficial for clinical quality control and to hospital administrators for understanding surgery planning. Video acquired during surgery typically can be leveraged for this task. Currently, a combination of convolutional neural network (CNN) and recurrent neural networks (RNN) are popularly used for video analysis in general, not only being restricted to surgical videos. In this paper, we propose a multi-task learning framework using CNN followed by a bi-directional long short term memory (Bi-LSTM) to learn to encapsulate both forward and backward temporal dependencies. Further, the joint distribution indicating set of tools associated with a phase is used as an additional loss during learning to correct for their co-occurrence in any predictions. Experimental evaluation is performed using the Cholec80 dataset. We report a mean average precision (mAP) score of 0.99 and 0.86 for tool and phase identification respectively which are higher compared to prior-art in the field.
Fully Convolutional Neural Network for Semantic Segmentation of Anatomical Structure and Pathologies in Colour Fundus Images Associated with Diabetic Retinopathy
Feb 07, 2019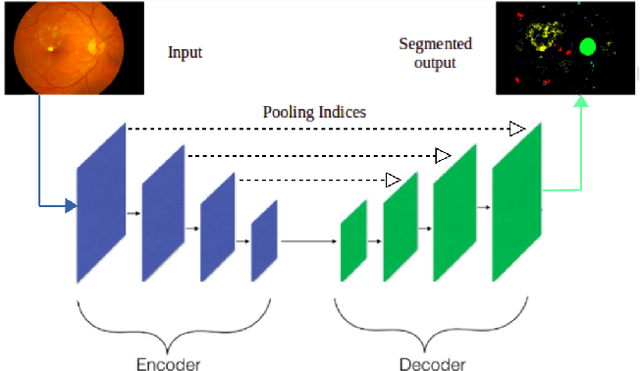
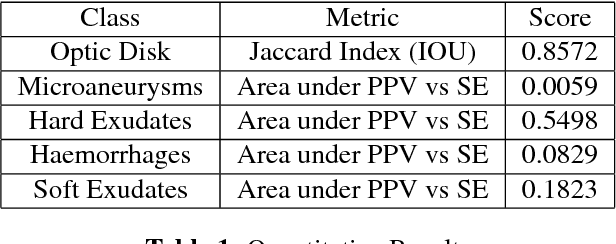
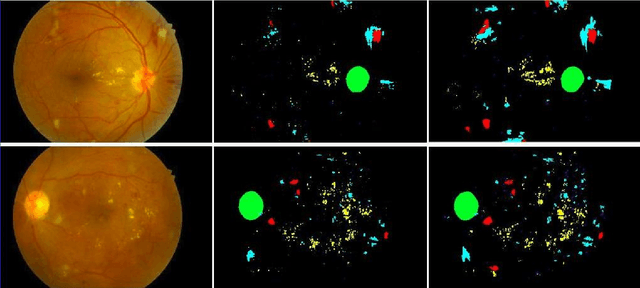
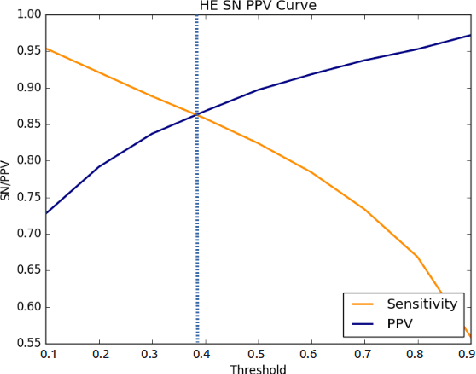
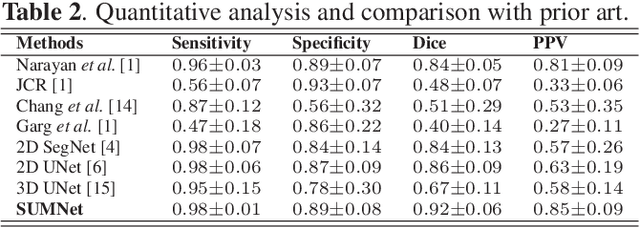
Abstract:Diabetic retinopathy (DR) is the most common form of diabetic eye disease. Retinopathy can affect all diabetic patients and becomes particularly dangerous, increasing the risk of blindness, if it is left untreated. The success rate of its curability solemnly depends on diagnosis at an early stage. The development of automated computer aided disease diagnosis tools could help in faster detection of symptoms with a wider reach and reasonable cost. This paper proposes a method for the automated segmentation of retinal lesions and optic disk in fundus images using a deep fully convolutional neural network for semantic segmentation. This trainable segmentation pipeline consists of an encoder network, a corresponding decoder network followed by pixel-wise classification to segment microaneurysms, hemorrhages, hard exudates, soft exudates, optic disk from background. The network was trained using Binary cross entropy criterion with Sigmoid as the last layer, while during an additional SoftMax layer was used for boosting response of single class. The performance of the proposed method is evaluated using sensitivity, positive prediction value (PPV) and accuracy as the metrices. Further, the position of the Optic disk is localised using the segmented output map.
Segmentation of Lumen and External Elastic Laminae in Intravascular Ultrasound Images using Ultrasonic Backscattering Physics Initialized Multiscale Random Walks
Jan 21, 2019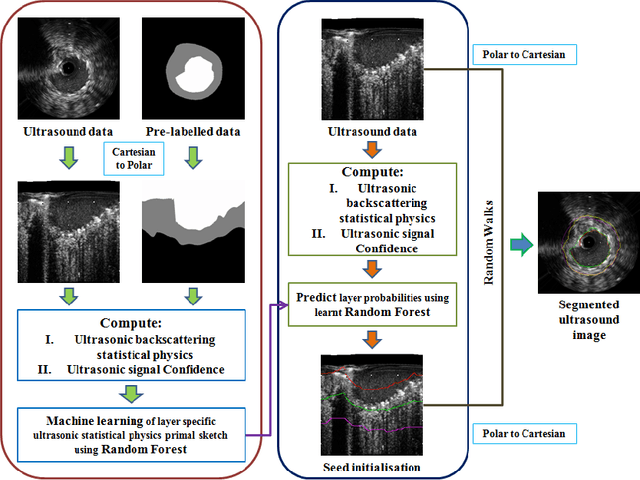
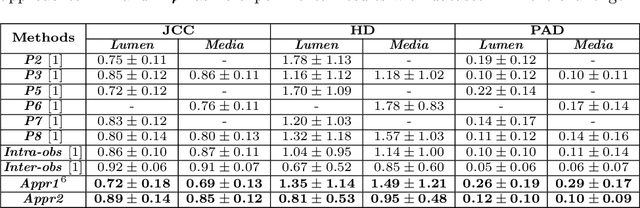
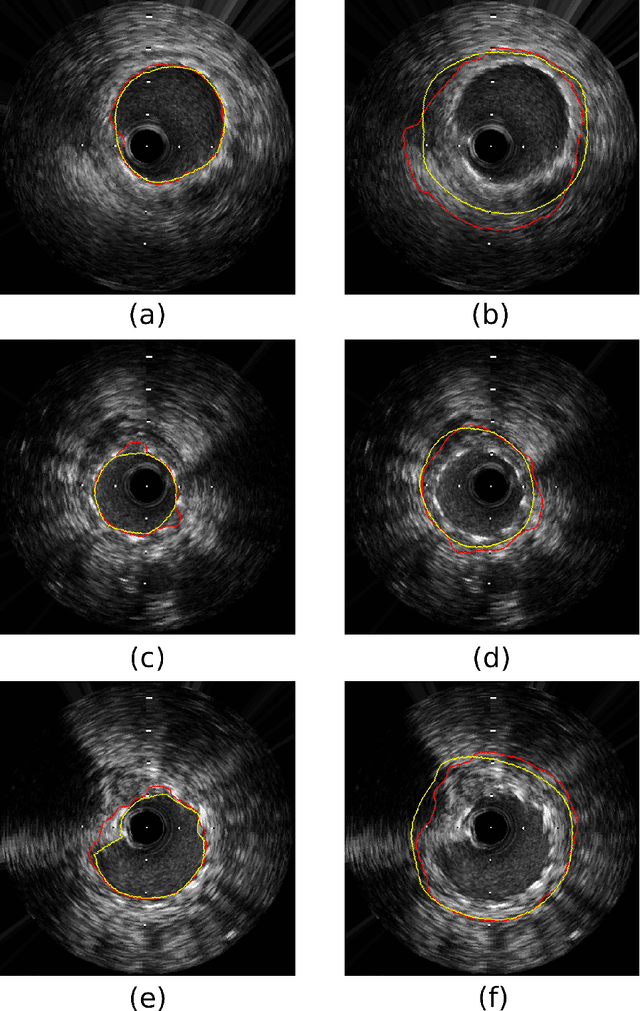
Abstract:Coronary artery disease accounts for a large number of deaths across the world and clinicians generally prefer using x-ray computed tomography or magnetic resonance imaging for localizing vascular pathologies. Interventional imaging modalities like intravascular ultrasound (IVUS) are used to adjunct diagnosis of atherosclerotic plaques in vessels, and help assess morphological state of the vessel and plaque, which play a significant role for treatment planning. Since speckle intensity in IVUS images are inherently stochastic in nature and challenge clinicians with accurate visibility of the vessel wall boundaries, it requires automation. In this paper we present a method for segmenting the lumen and external elastic laminae of the artery wall in IVUS images using random walks over a multiscale pyramid of Gaussian decomposed frames. The seeds for the random walker are initialized by supervised learning of ultrasonic backscattering and attenuation statistical mechanics from labelled training samples. We have experimentally evaluated the performance using $77$ IVUS images acquired at $40$ MHz that are available in the IVUS segmentation challenge dataset\footnote{http://www.cvc.uab.es/IVUSchallenge2011/dataset.html} to obtain a Jaccard score of $0.89 \pm 0.14$ for lumen and $0.85 \pm 0.12$ for external elastic laminae segmentation over a $10$-fold cross-validation study.
SUMNet: Fully Convolutional Model for Fast Segmentation of Anatomical Structures in Ultrasound Volumes
Jan 21, 2019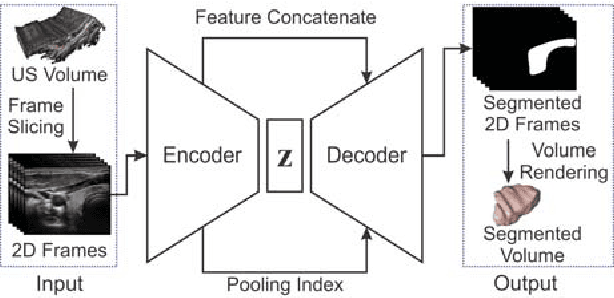
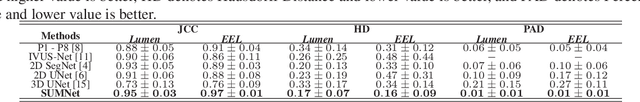
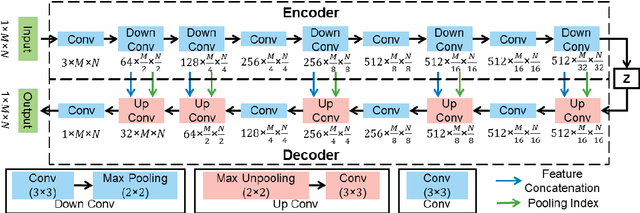
Abstract:Ultrasound imaging is generally employed for real-time investigation of internal anatomy of the human body for disease identification. Delineation of the anatomical boundary of organs and pathological lesions is quite challenging due to the stochastic nature of speckle intensity in the images, which also introduces visual fatigue for the observer. This paper introduces a fully convolutional neural network based method to segment organ and pathologies in ultrasound volume by learning the spatial-relationship between closely related classes in the presence of stochastically varying speckle intensity. We propose a convolutional encoder-decoder like framework with (i) feature concatenation across matched layers in encoder and decoder and (ii) index passing based unpooling at the decoder for semantic segmentation of ultrasound volumes. We have experimentally evaluated the performance on publicly available datasets consisting of $10$ intravascular ultrasound pullback acquired at $20$ MHz and $16$ freehand thyroid ultrasound volumes acquired $11 - 16$ MHz. We have obtained a dice score of $0.93 \pm 0.08$ and $0.92 \pm 0.06$ respectively, following a $10$-fold cross-validation experiment while processing frame of $256 \times 384$ pixel in $0.035$s and a volume of $256 \times 384 \times 384$ voxel in $13.44$s.
Learning a Deep Convolution Network with Turing Test Adversaries for Microscopy Image Super Resolution
Jan 18, 2019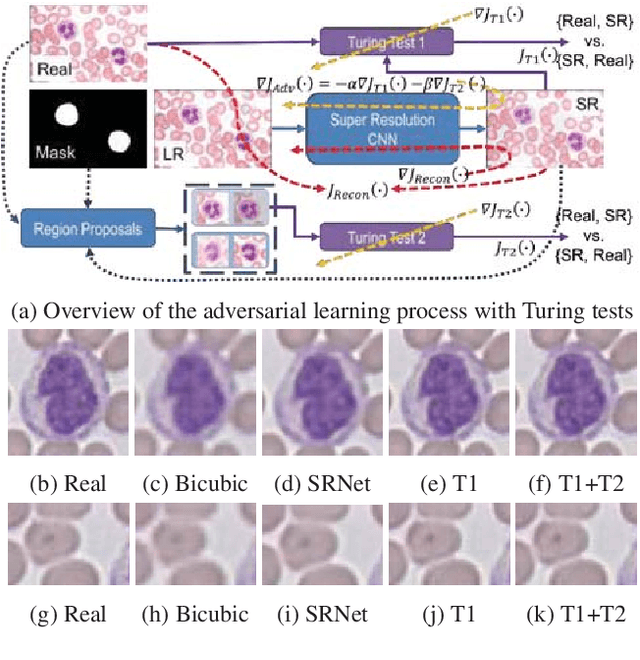
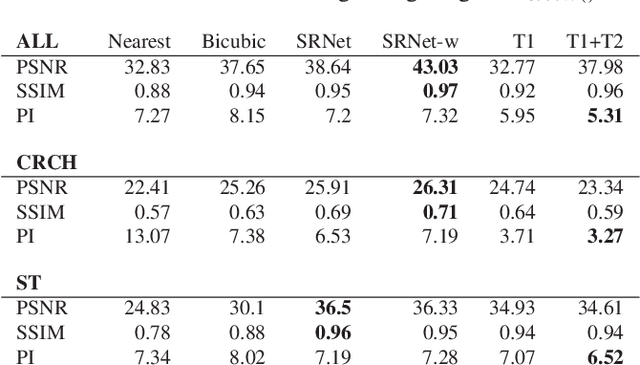
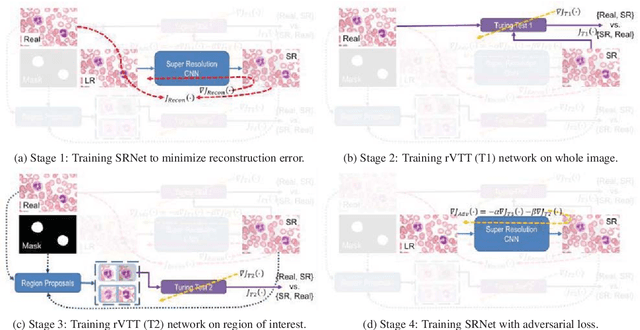

Abstract:Adversarially trained deep neural networks have significantly improved performance of single image super resolution, by hallucinating photorealistic local textures, thereby greatly reducing the perception difference between a real high resolution image and its super resolved (SR) counterpart. However, application to medical imaging requires preservation of diagnostically relevant features while refraining from introducing any diagnostically confusing artifacts. We propose using a deep convolutional super resolution network (SRNet) trained for (i) minimising reconstruction loss between the real and SR images, and (ii) maximally confusing learned relativistic visual Turing test (rVTT) networks to discriminate between (a) pair of real and SR images (T1) and (b) pair of patches in real and SR selected from region of interest (T2). The adversarial loss of T1 and T2 while backpropagated through SRNet helps it learn to reconstruct pathorealism in the regions of interest such as white blood cells (WBC) in peripheral blood smears or epithelial cells in histopathology of cancerous biopsy tissues, which are experimentally demonstrated here. Experiments performed for measuring signal distortion loss using peak signal to noise ratio (pSNR) and structural similarity (SSIM) with variation of SR scale factors, impact of rVTT adversarial losses, and impact on reporting using SR on a commercially available artificial intelligence (AI) digital pathology system substantiate our claims.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge