Dazhou Guo
Comprehensive and Clinically Accurate Head and Neck Organs at Risk Delineation via Stratified Deep Learning: A Large-scale Multi-Institutional Study
Nov 01, 2021
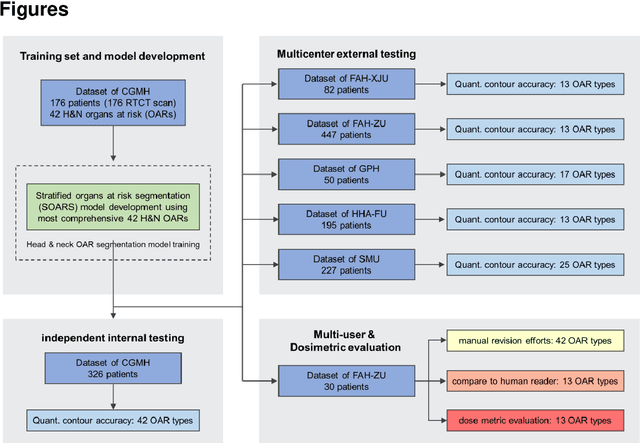
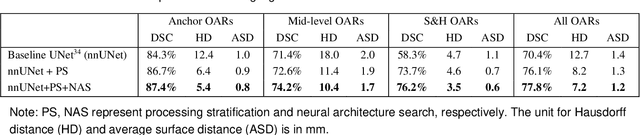
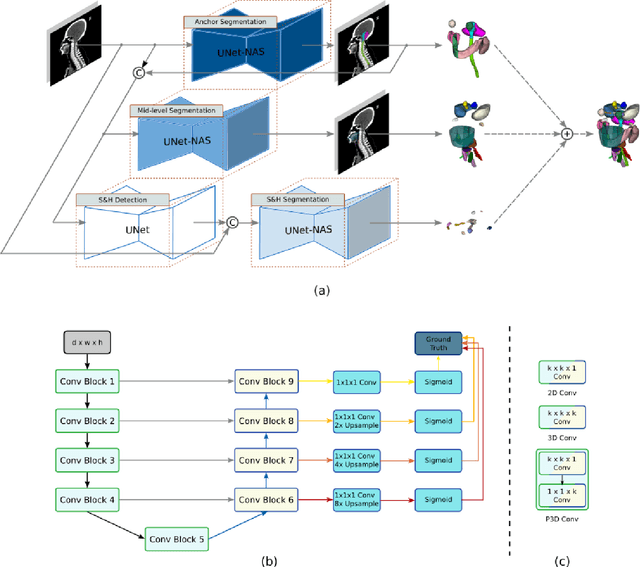
Abstract:Accurate organ at risk (OAR) segmentation is critical to reduce the radiotherapy post-treatment complications. Consensus guidelines recommend a set of more than 40 OARs in the head and neck (H&N) region, however, due to the predictable prohibitive labor-cost of this task, most institutions choose a substantially simplified protocol by delineating a smaller subset of OARs and neglecting the dose distributions associated with other OARs. In this work we propose a novel, automated and highly effective stratified OAR segmentation (SOARS) system using deep learning to precisely delineate a comprehensive set of 42 H&N OARs. SOARS stratifies 42 OARs into anchor, mid-level, and small & hard subcategories, with specifically derived neural network architectures for each category by neural architecture search (NAS) principles. We built SOARS models using 176 training patients in an internal institution and independently evaluated on 1327 external patients across six different institutions. It consistently outperformed other state-of-the-art methods by at least 3-5% in Dice score for each institutional evaluation (up to 36% relative error reduction in other metrics). More importantly, extensive multi-user studies evidently demonstrated that 98% of the SOARS predictions need only very minor or no revisions for direct clinical acceptance (saving 90% radiation oncologists workload), and their segmentation and dosimetric accuracy are within or smaller than the inter-user variation. These findings confirmed the strong clinical applicability of SOARS for the OAR delineation process in H&N cancer radiotherapy workflows, with improved efficiency, comprehensiveness, and quality.
Multi-institutional Validation of Two-Streamed Deep Learning Method for Automated Delineation of Esophageal Gross Tumor Volume using planning-CT and FDG-PETCT
Oct 11, 2021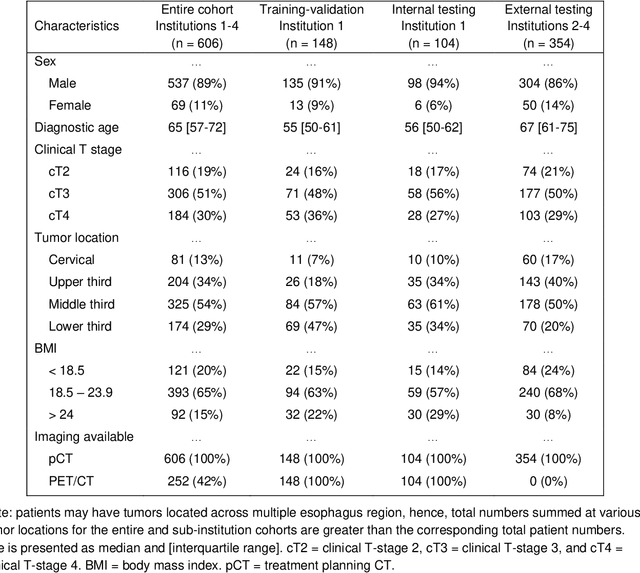
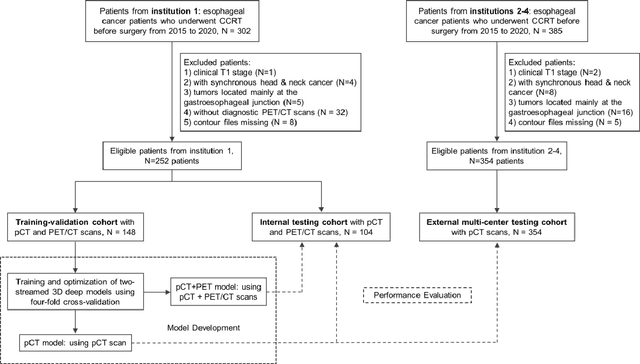
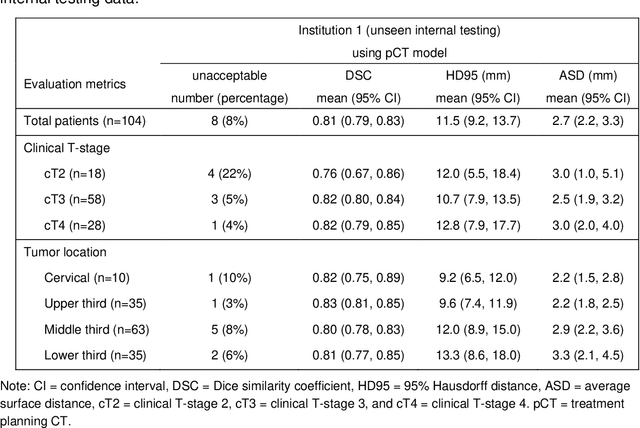
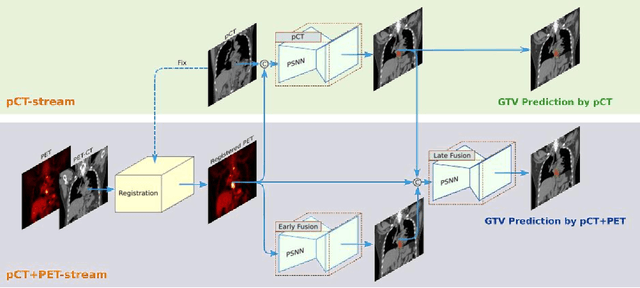
Abstract:Background: The current clinical workflow for esophageal gross tumor volume (GTV) contouring relies on manual delineation of high labor-costs and interuser variability. Purpose: To validate the clinical applicability of a deep learning (DL) multi-modality esophageal GTV contouring model, developed at 1 institution whereas tested at multiple ones. Methods and Materials: We collected 606 esophageal cancer patients from four institutions. 252 institution-1 patients had a treatment planning-CT (pCT) and a pair of diagnostic FDG-PETCT; 354 patients from other 3 institutions had only pCT. A two-streamed DL model for GTV segmentation was developed using pCT and PETCT scans of a 148 patient institution-1 subset. This built model had the flexibility of segmenting GTVs via only pCT or pCT+PETCT combined. For independent evaluation, the rest 104 institution-1 patients behaved as unseen internal testing, and 354 institutions 2-4 patients were used for external testing. We evaluated manual revision degrees by human experts to assess the contour-editing effort. The performance of the deep model was compared against 4 radiation oncologists in a multiuser study with 20 random external patients. Contouring accuracy and time were recorded for the pre-and post-DL assisted delineation process. Results: Our model achieved high segmentation accuracy in internal testing (mean Dice score: 0.81 using pCT and 0.83 using pCT+PET) and generalized well to external evaluation (mean DSC: 0.80). Expert assessment showed that the predicted contours of 88% patients need only minor or no revision. In multi-user evaluation, with the assistance of a deep model, inter-observer variation and required contouring time were reduced by 37.6% and 48.0%, respectively. Conclusions: Deep learning predicted GTV contours were in close agreement with the ground truth and could be adopted clinically with mostly minor or no changes.
SAME: Deformable Image Registration based on Self-supervised Anatomical Embeddings
Sep 23, 2021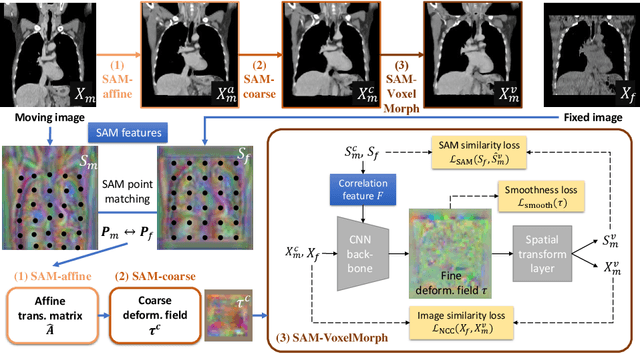
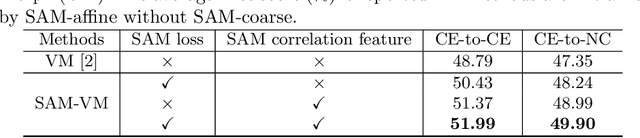
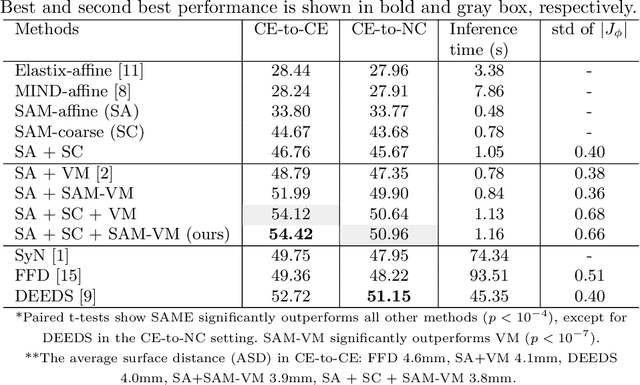

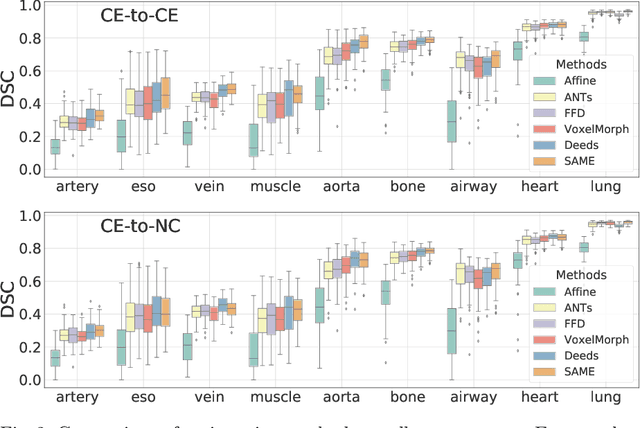
Abstract:In this work, we introduce a fast and accurate method for unsupervised 3D medical image registration. This work is built on top of a recent algorithm SAM, which is capable of computing dense anatomical/semantic correspondences between two images at the pixel level. Our method is named SAME, which breaks down image registration into three steps: affine transformation, coarse deformation, and deep deformable registration. Using SAM embeddings, we enhance these steps by finding more coherent correspondences, and providing features and a loss function with better semantic guidance. We collect a multi-phase chest computed tomography dataset with 35 annotated organs for each patient and conduct inter-subject registration for quantitative evaluation. Results show that SAME outperforms widely-used traditional registration techniques (Elastix FFD, ANTs SyN) and learning based VoxelMorph method by at least 4.7% and 2.7% in Dice scores for two separate tasks of within-contrast-phase and across-contrast-phase registration, respectively. SAME achieves the comparable performance to the best traditional registration method, DEEDS (from our evaluation), while being orders of magnitude faster (from 45 seconds to 1.2 seconds).
DeepStationing: Thoracic Lymph Node Station Parsing in CT Scans using Anatomical Context Encoding and Key Organ Auto-Search
Sep 20, 2021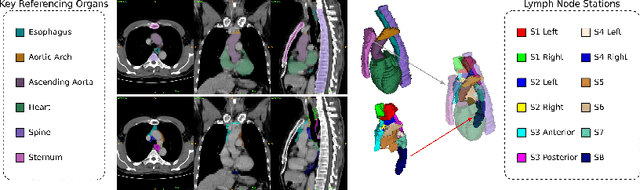
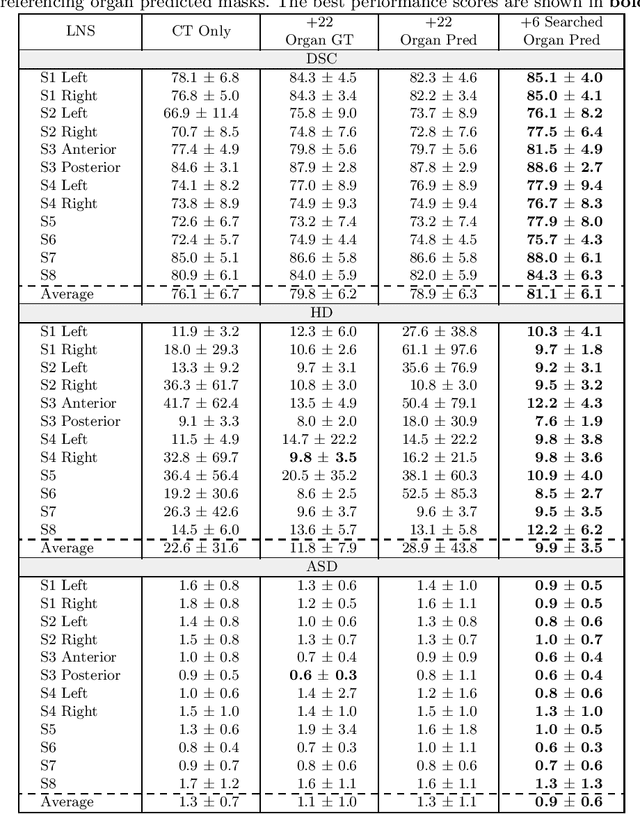
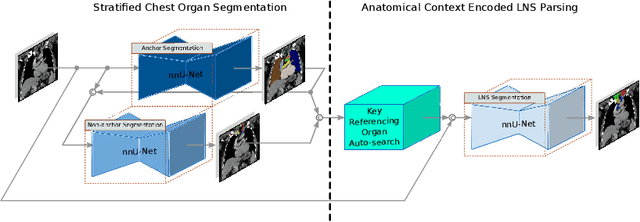
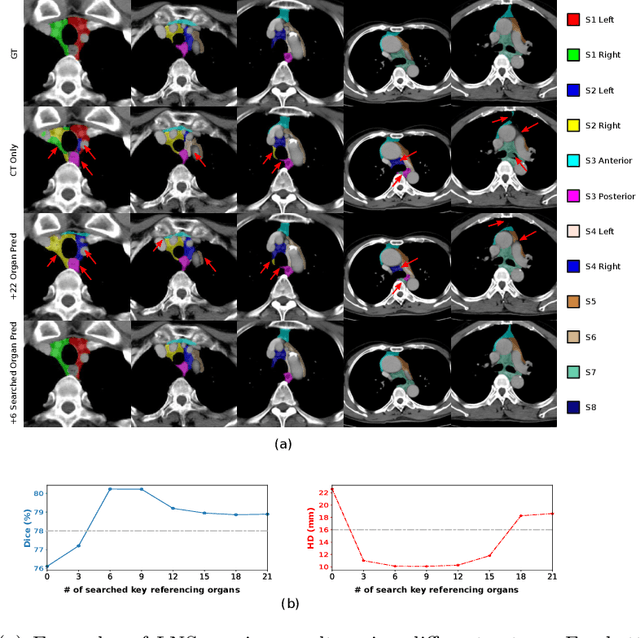
Abstract:Lymph node station (LNS) delineation from computed tomography (CT) scans is an indispensable step in radiation oncology workflow. High inter-user variabilities across oncologists and prohibitive laboring costs motivated the automated approach. Previous works exploit anatomical priors to infer LNS based on predefined ad-hoc margins. However, without voxel-level supervision, the performance is severely limited. LNS is highly context-dependent - LNS boundaries are constrained by anatomical organs - we formulate it as a deep spatial and contextual parsing problem via encoded anatomical organs. This permits the deep network to better learn from both CT appearance and organ context. We develop a stratified referencing organ segmentation protocol that divides the organs into anchor and non-anchor categories and uses the former's predictions to guide the later segmentation. We further develop an auto-search module to identify the key organs that opt for the optimal LNS parsing performance. Extensive four-fold cross-validation experiments on a dataset of 98 esophageal cancer patients (with the most comprehensive set of 12 LNSs + 22 organs in thoracic region to date) are conducted. Our LNS parsing model produces significant performance improvements, with an average Dice score of 81.1% +/- 6.1%, which is 5.0% and 19.2% higher over the pure CT-based deep model and the previous representative approach, respectively.
Self-supervised Learning of Pixel-wise Anatomical Embeddings in Radiological Images
Dec 04, 2020
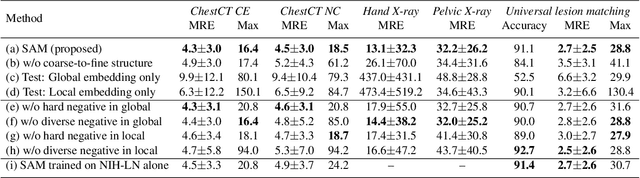
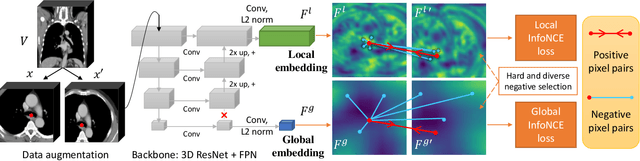
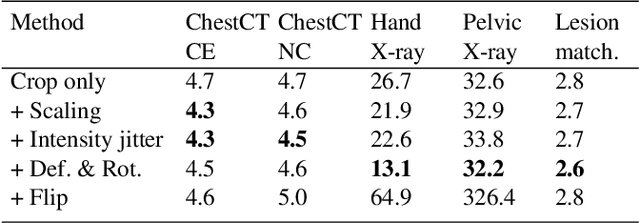
Abstract:Radiological images such as computed tomography (CT) and X-rays render anatomy with intrinsic structures. Being able to reliably locate the same anatomical or semantic structure across varying images is a fundamental task in medical image analysis. In principle it is possible to use landmark detection or semantic segmentation for this task, but to work well these require large numbers of labeled data for each anatomical structure and sub-structure of interest. A more universal approach would discover the intrinsic structure from unlabeled images. We introduce such an approach, called Self-supervised Anatomical eMbedding (SAM). SAM generates semantic embeddings for each image pixel that describes its anatomical location or body part. To produce such embeddings, we propose a pixel-level contrastive learning framework. A coarse-to-fine strategy ensures both global and local anatomical information are encoded. Negative sample selection strategies are designed to enhance the discriminability among different body parts. Using SAM, one can label any point of interest on a template image, and then locate the same body part in other images by simple nearest neighbor searching. We demonstrate the effectiveness of SAM in multiple tasks with 2D and 3D image modalities. On a chest CT dataset with 19 landmarks, SAM outperforms widely-used registration algorithms while being 200 times faster. On two X-ray datasets, SAM, with only one labeled template image, outperforms supervised methods trained on 50 labeled images. We also apply SAM on whole-body follow-up lesion matching in CT and obtain an accuracy of 91%.
Lymph Node Gross Tumor Volume Detection in Oncology Imaging via Relationship Learning Using Graph Neural Network
Aug 29, 2020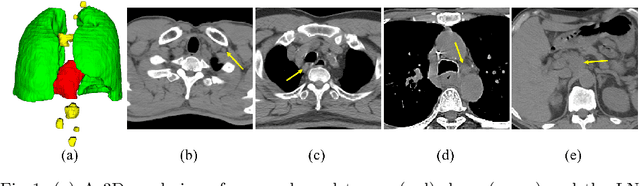
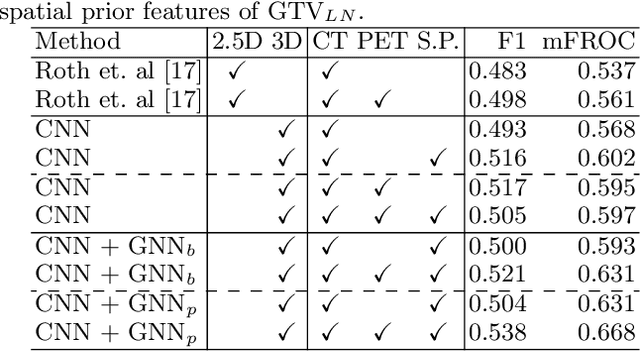
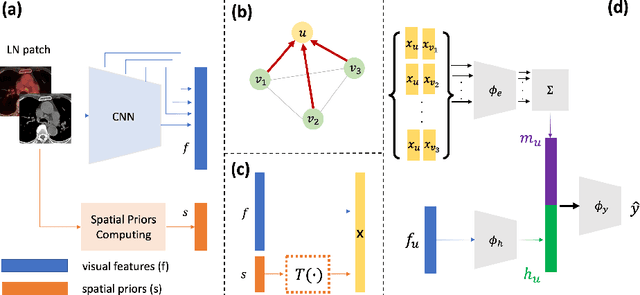
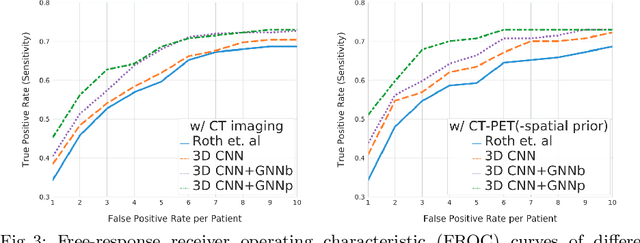
Abstract:Determining the spread of GTV$_{LN}$ is essential in defining the respective resection or irradiating regions for the downstream workflows of surgical resection and radiotherapy for many cancers. Different from the more common enlarged lymph node (LN), GTV$_{LN}$ also includes smaller ones if associated with high positron emission tomography signals and/or any metastasis signs in CT. This is a daunting task. In this work, we propose a unified LN appearance and inter-LN relationship learning framework to detect the true GTV$_{LN}$. This is motivated by the prior clinical knowledge that LNs form a connected lymphatic system, and the spread of cancer cells among LNs often follows certain pathways. Specifically, we first utilize a 3D convolutional neural network with ROI-pooling to extract the GTV$_{LN}$'s instance-wise appearance features. Next, we introduce a graph neural network to further model the inter-LN relationships where the global LN-tumor spatial priors are included in the learning process. This leads to an end-to-end trainable network to detect by classifying GTV$_{LN}$. We operate our model on a set of GTV$_{LN}$ candidates generated by a preliminary 1st-stage method, which has a sensitivity of $>85\%$ at the cost of high false positive (FP) ($>15$ FPs per patient). We validate our approach on a radiotherapy dataset with 142 paired PET/RTCT scans containing the chest and upper abdominal body parts. The proposed method significantly improves over the state-of-the-art (SOTA) LN classification method by $5.5\%$ and $13.1\%$ in F1 score and the averaged sensitivity value at $2, 3, 4, 6$ FPs per patient, respectively.
Lymph Node Gross Tumor Volume Detection and Segmentation via Distance-based Gating using 3D CT/PET Imaging in Radiotherapy
Aug 27, 2020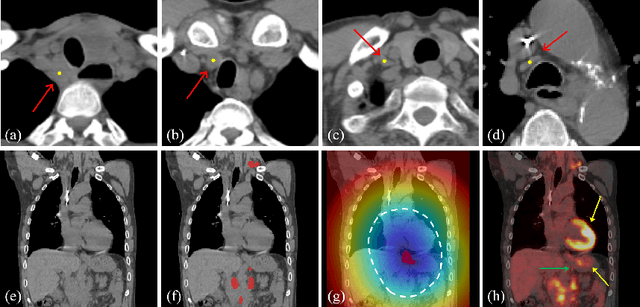
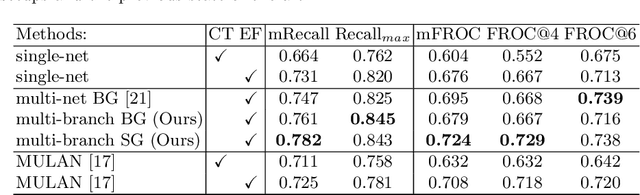
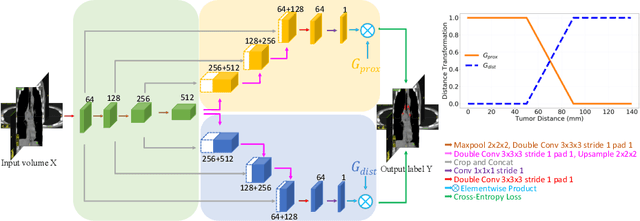
Abstract:Finding, identifying and segmenting suspicious cancer metastasized lymph nodes from 3D multi-modality imaging is a clinical task of paramount importance. In radiotherapy, they are referred to as Lymph Node Gross Tumor Volume (GTVLN). Determining and delineating the spread of GTVLN is essential in defining the corresponding resection and irradiating regions for the downstream workflows of surgical resection and radiotherapy of various cancers. In this work, we propose an effective distance-based gating approach to simulate and simplify the high-level reasoning protocols conducted by radiation oncologists, in a divide-and-conquer manner. GTVLN is divided into two subgroups of tumor-proximal and tumor-distal, respectively, by means of binary or soft distance gating. This is motivated by the observation that each category can have distinct though overlapping distributions of appearance, size and other LN characteristics. A novel multi-branch detection-by-segmentation network is trained with each branch specializing on learning one GTVLN category features, and outputs from multi-branch are fused in inference. The proposed method is evaluated on an in-house dataset of $141$ esophageal cancer patients with both PET and CT imaging modalities. Our results validate significant improvements on the mean recall from $72.5\%$ to $78.2\%$, as compared to previous state-of-the-art work. The highest achieved GTVLN recall of $82.5\%$ at $20\%$ precision is clinically relevant and valuable since human observers tend to have low sensitivity (around $80\%$ for the most experienced radiation oncologists, as reported by literature).
Detecting Scatteredly-Distributed, Small, andCritically Important Objects in 3D OncologyImaging via Decision Stratification
May 27, 2020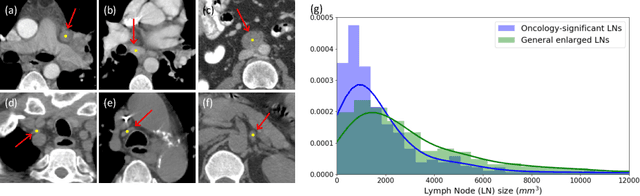
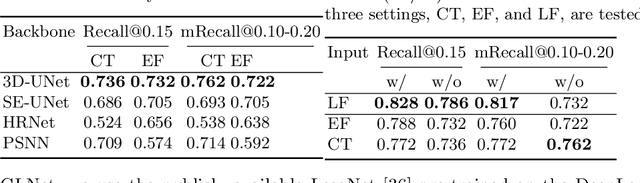
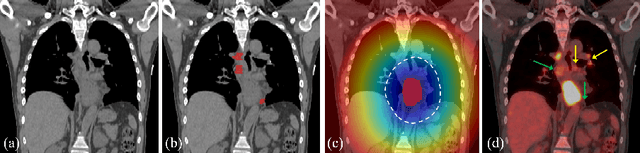
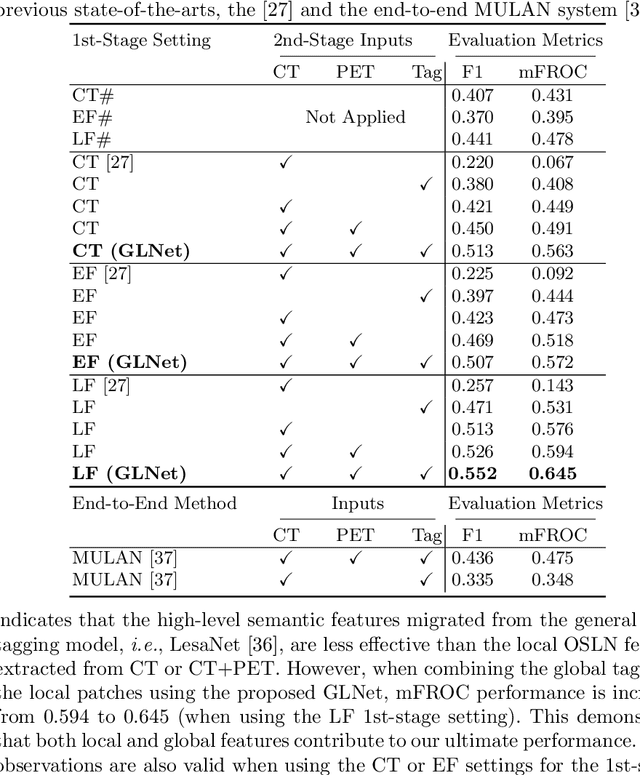
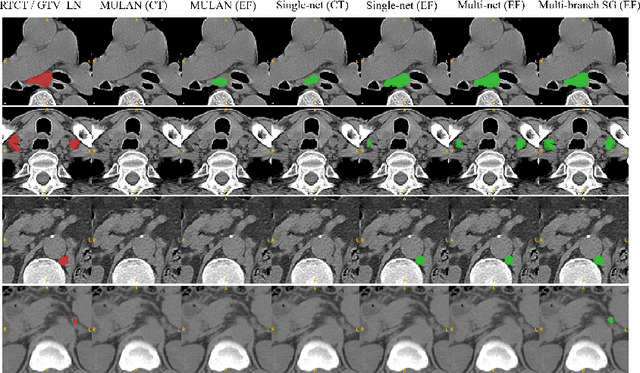
Abstract:Finding and identifying scatteredly-distributed, small, and critically important objects in 3D oncology images is very challenging. We focus on the detection and segmentation of oncology-significant (or suspicious cancer metastasized) lymph nodes (OSLNs), which has not been studied before as a computational task. Determining and delineating the spread of OSLNs is essential in defining the corresponding resection/irradiating regions for the downstream workflows of surgical resection and radiotherapy of various cancers. For patients who are treated with radiotherapy, this task is performed by experienced radiation oncologists that involves high-level reasoning on whether LNs are metastasized, which is subject to high inter-observer variations. In this work, we propose a divide-and-conquer decision stratification approach that divides OSLNs into tumor-proximal and tumor-distal categories. This is motivated by the observation that each category has its own different underlying distributions in appearance, size and other characteristics. Two separate detection-by-segmentation networks are trained per category and fused. To further reduce false positives (FP), we present a novel global-local network (GLNet) that combines high-level lesion characteristics with features learned from localized 3D image patches. Our method is evaluated on a dataset of 141 esophageal cancer patients with PET and CT modalities (the largest to-date). Our results significantly improve the recall from $45\%$ to $67\%$ at $3$ FPs per patient as compared to previous state-of-the-art methods. The highest achieved OSLN recall of $0.828$ is clinically relevant and valuable.
Organ at Risk Segmentation for Head and Neck Cancer using Stratified Learning and Neural Architecture Search
Apr 17, 2020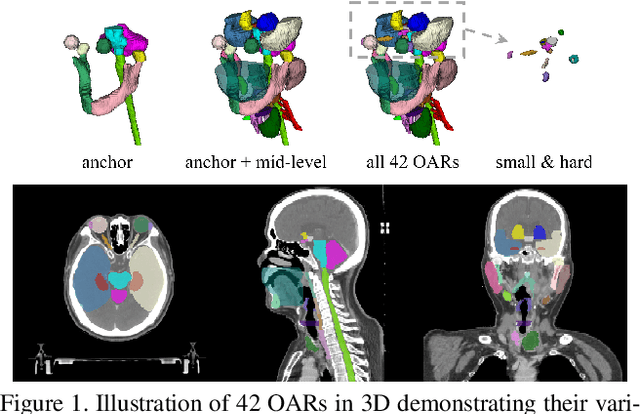
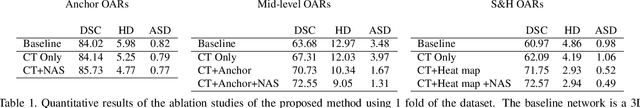
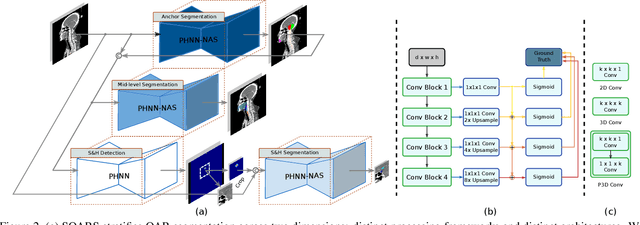
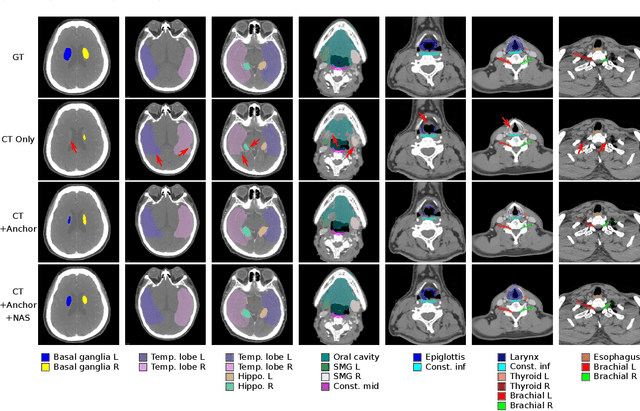
Abstract:OAR segmentation is a critical step in radiotherapy of head and neck (H&N) cancer, where inconsistencies across radiation oncologists and prohibitive labor costs motivate automated approaches. However, leading methods using standard fully convolutional network workflows that are challenged when the number of OARs becomes large, e.g. > 40. For such scenarios, insights can be gained from the stratification approaches seen in manual clinical OAR delineation. This is the goal of our work, where we introduce stratified organ at risk segmentation (SOARS), an approach that stratifies OARs into anchor, mid-level, and small & hard (S&H) categories. SOARS stratifies across two dimensions. The first dimension is that distinct processing pipelines are used for each OAR category. In particular, inspired by clinical practices, anchor OARs are used to guide the mid-level and S&H categories. The second dimension is that distinct network architectures are used to manage the significant contrast, size, and anatomy variations between different OARs. We use differentiable neural architecture search (NAS), allowing the network to choose among 2D, 3D or Pseudo-3D convolutions. Extensive 4-fold cross-validation on 142 H&N cancer patients with 42 manually labeled OARs, the most comprehensive OAR dataset to date, demonstrates that both pipeline- and NAS-stratification significantly improves quantitative performance over the state-of-the-art (from 69.52% to 73.68% in absolute Dice scores). Thus, SOARS provides a powerful and principled means to manage the highly complex segmentation space of OARs.
Deep Esophageal Clinical Target Volume Delineation using Encoded 3D Spatial Context of Tumors, Lymph Nodes, and Organs At Risk
Sep 06, 2019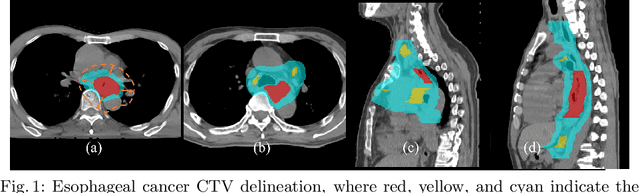

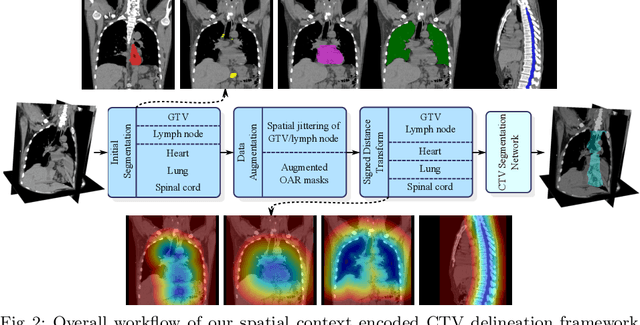
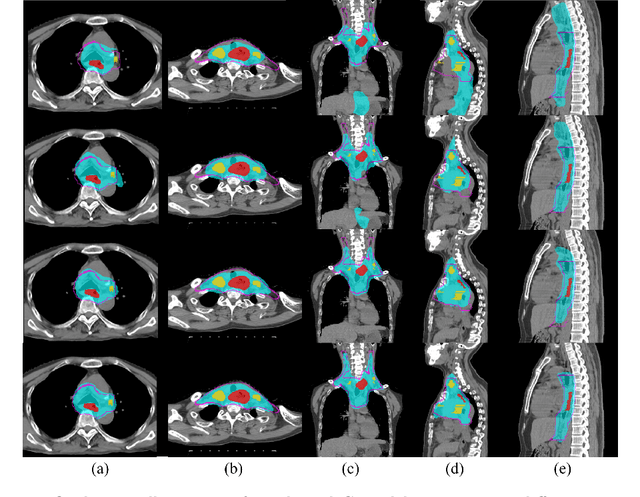
Abstract:Clinical target volume (CTV) delineation from radiotherapy computed tomography (RTCT) images is used to define the treatment areas containing the gross tumor volume (GTV) and/or sub-clinical malignant disease for radiotherapy (RT). High intra- and inter-user variability makes this a particularly difficult task for esophageal cancer. This motivates automated solutions, which is the aim of our work. Because CTV delineation is highly context-dependent--it must encompass the GTV and regional lymph nodes (LNs) while also avoiding excessive exposure to the organs at risk (OARs)--we formulate it as a deep contextual appearance-based problem using encoded spatial contexts of these anatomical structures. This allows the deep network to better learn from and emulate the margin- and appearance-based delineation performed by human physicians. Additionally, we develop domain-specific data augmentation to inject robustness to our system. Finally, we show that a simple 3D progressive holistically nested network (PHNN), which avoids computationally heavy decoding paths while still aggregating features at different levels of context, can outperform more complicated networks. Cross-validated experiments on a dataset of 135 esophageal cancer patients demonstrate that our encoded spatial context approach can produce concrete performance improvements, with an average Dice score of 83.9% and an average surface distance of 4.2 mm, representing improvements of 3.8% and 2.4 mm, respectively, over the state-of-the-art approach.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge