Bernhard Kainz
Biomedical Image Analysis Group, Department of Computing, Imperial College London
Adnexal Mass Segmentation with Ultrasound Data Synthesis
Sep 25, 2022Abstract:Ovarian cancer is the most lethal gynaecological malignancy. The disease is most commonly asymptomatic at its early stages and its diagnosis relies on expert evaluation of transvaginal ultrasound images. Ultrasound is the first-line imaging modality for characterising adnexal masses, it requires significant expertise and its analysis is subjective and labour-intensive, therefore open to error. Hence, automating processes to facilitate and standardise the evaluation of scans is desired in clinical practice. Using supervised learning, we have demonstrated that segmentation of adnexal masses is possible, however, prevalence and label imbalance restricts the performance on under-represented classes. To mitigate this we apply a novel pathology-specific data synthesiser. We create synthetic medical images with their corresponding ground truth segmentations by using Poisson image editing to integrate less common masses into other samples. Our approach achieves the best performance across all classes, including an improvement of up to 8% when compared with nnU-Net baseline approaches.
nnOOD: A Framework for Benchmarking Self-supervised Anomaly Localisation Methods
Sep 02, 2022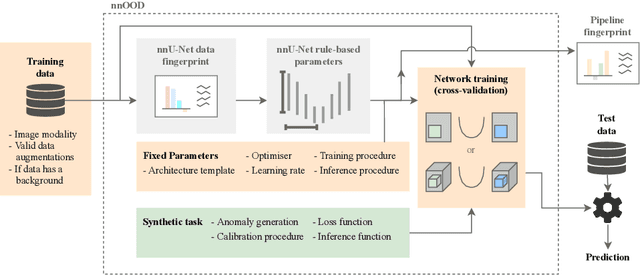
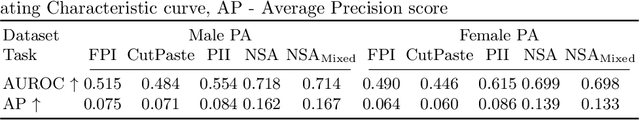
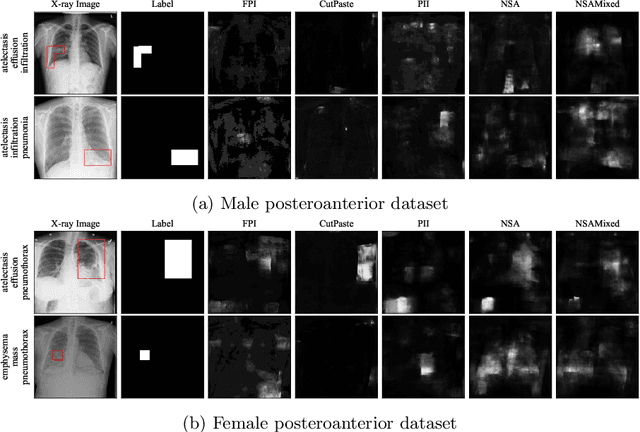
Abstract:The wide variety of in-distribution and out-of-distribution data in medical imaging makes universal anomaly detection a challenging task. Recently a number of self-supervised methods have been developed that train end-to-end models on healthy data augmented with synthetic anomalies. However, it is difficult to compare these methods as it is not clear whether gains in performance are from the task itself or the training pipeline around it. It is also difficult to assess whether a task generalises well for universal anomaly detection, as they are often only tested on a limited range of anomalies. To assist with this we have developed nnOOD, a framework that adapts nnU-Net to allow for comparison of self-supervised anomaly localisation methods. By isolating the synthetic, self-supervised task from the rest of the training process we perform a more faithful comparison of the tasks, whilst also making the workflow for evaluating over a given dataset quick and easy. Using this we have implemented the current state-of-the-art tasks and evaluated them on a challenging X-ray dataset.
Improved post-hoc probability calibration for out-of-domain MRI segmentation
Aug 04, 2022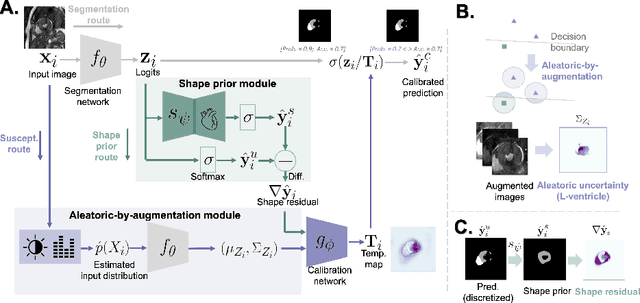
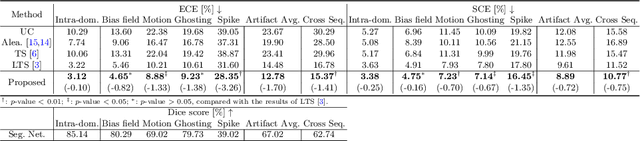
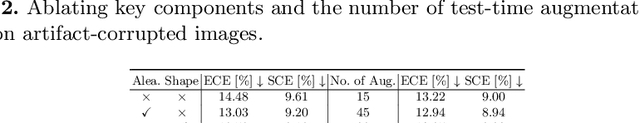
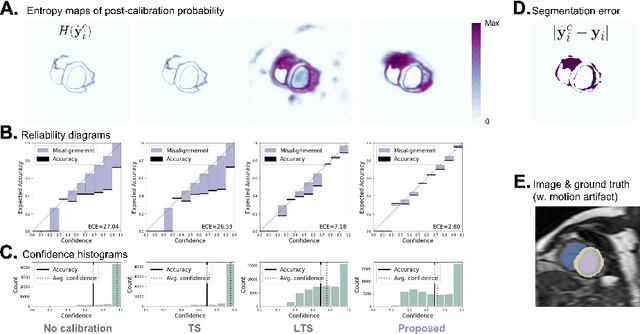
Abstract:Probability calibration for deep models is highly desirable in safety-critical applications such as medical imaging. It makes output probabilities of deep networks interpretable, by aligning prediction probabilities with the actual accuracy in test data. In image segmentation, well-calibrated probabilities allow radiologists to identify regions where model-predicted segmentations are unreliable. These unreliable predictions often occur to out-of-domain (OOD) images that are caused by imaging artifacts or unseen imaging protocols. Unfortunately, most previous calibration methods for image segmentation perform sub-optimally on OOD images. To reduce the calibration error when confronted with OOD images, we propose a novel post-hoc calibration model. Our model leverages the pixel susceptibility against perturbations at the local level, and the shape prior information at the global level. The model is tested on cardiac MRI segmentation datasets that contain unseen imaging artifacts and images from an unseen imaging protocol. We demonstrate reduced calibration errors compared with the state-of-the-art calibration algorithm.
Placenta Segmentation in Ultrasound Imaging: Addressing Sources of Uncertainty and Limited Field-of-View
Jun 29, 2022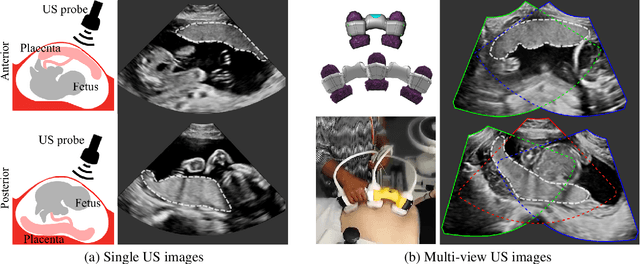
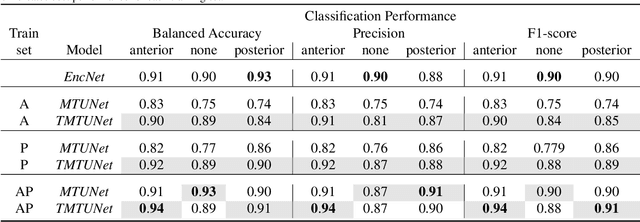
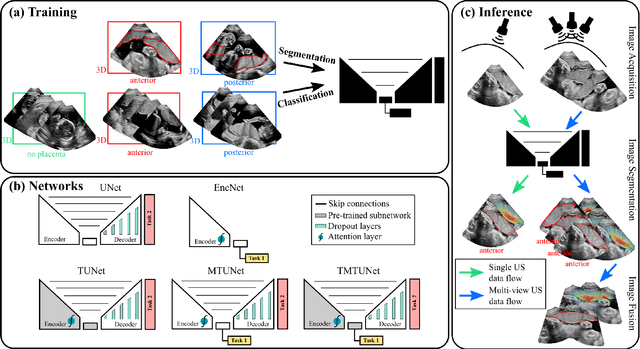
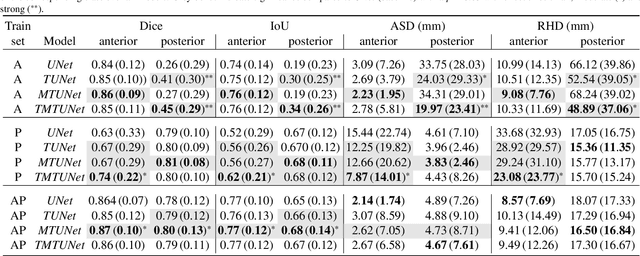
Abstract:Automatic segmentation of the placenta in fetal ultrasound (US) is challenging due to the (i) high diversity of placenta appearance, (ii) the restricted quality in US resulting in highly variable reference annotations, and (iii) the limited field-of-view of US prohibiting whole placenta assessment at late gestation. In this work, we address these three challenges with a multi-task learning approach that combines the classification of placental location (e.g., anterior, posterior) and semantic placenta segmentation in a single convolutional neural network. Through the classification task the model can learn from larger and more diverse datasets while improving the accuracy of the segmentation task in particular in limited training set conditions. With this approach we investigate the variability in annotations from multiple raters and show that our automatic segmentations (Dice of 0.86 for anterior and 0.83 for posterior placentas) achieve human-level performance as compared to intra- and inter-observer variability. Lastly, our approach can deliver whole placenta segmentation using a multi-view US acquisition pipeline consisting of three stages: multi-probe image acquisition, image fusion and image segmentation. This results in high quality segmentation of larger structures such as the placenta in US with reduced image artifacts which are beyond the field-of-view of single probes.
A Review of Causality for Learning Algorithms in Medical Image Analysis
Jun 11, 2022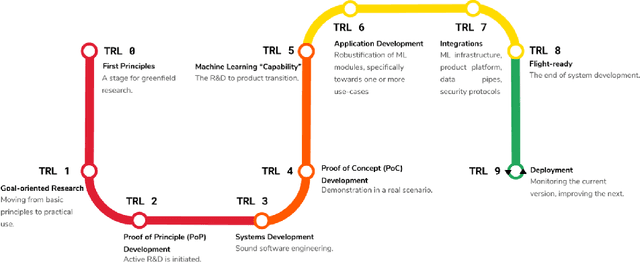
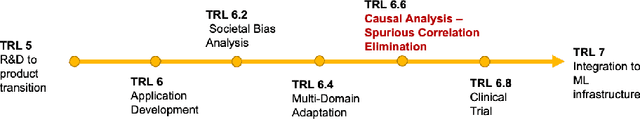

Abstract:Medical image analysis is a vibrant research area that offers doctors and medical practitioners invaluable insight and the ability to accurately diagnose and monitor disease. Machine learning provides an additional boost for this area. However, machine learning for medical image analysis is particularly vulnerable to natural biases like domain shifts that affect algorithmic performance and robustness. In this paper we analyze machine learning for medical image analysis within the framework of Technology Readiness Levels and review how causal analysis methods can fill a gap when creating robust and adaptable medical image analysis algorithms. We review methods using causality in medical imaging AI/ML and find that causal analysis has the potential to mitigate critical problems for clinical translation but that uptake and clinical downstream research has been limited so far.
Is More Data All You Need? A Causal Exploration
Jun 06, 2022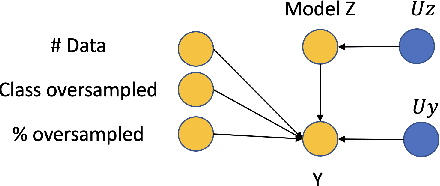
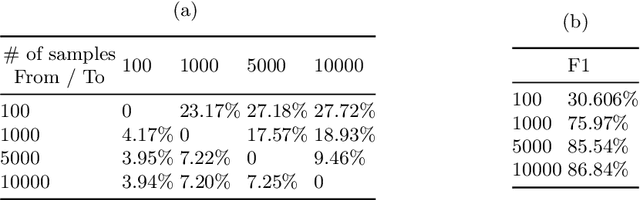

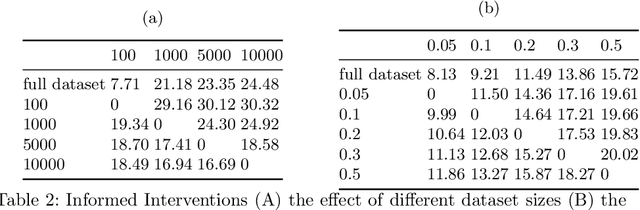
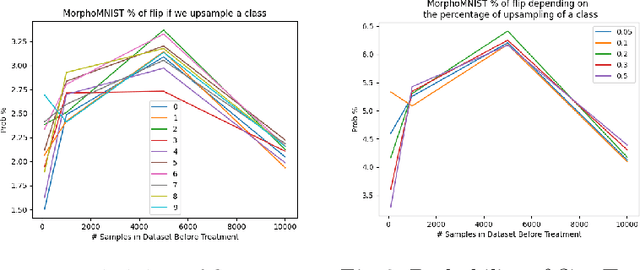
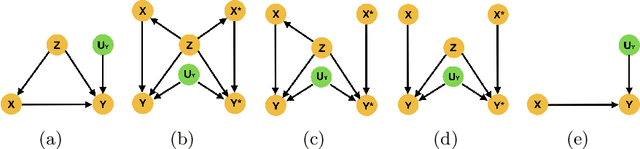
Abstract:Curating a large scale medical imaging dataset for machine learning applications is both time consuming and expensive. Balancing the workload between model development, data collection and annotations is difficult for machine learning practitioners, especially under time constraints. Causal analysis is often used in medicine and economics to gain insights about the effects of actions and policies. In this paper we explore the effect of dataset interventions on the output of image classification models. Through a causal approach we investigate the effects of the quantity and type of data we need to incorporate in a dataset to achieve better performance for specific subtasks. The main goal of this paper is to highlight the potential of causal analysis as a tool for resource optimization for developing medical imaging ML applications. We explore this concept with a synthetic dataset and an exemplary use-case for Diabetic Retinopathy image analysis.
D'ARTAGNAN: Counterfactual Video Generation
Jun 03, 2022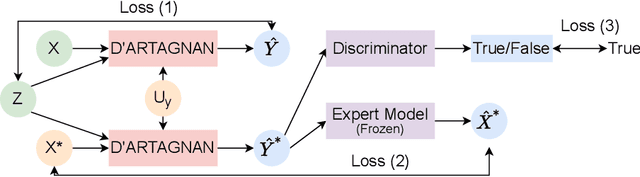
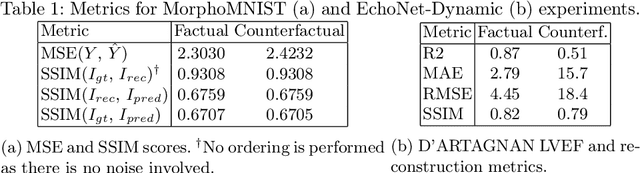


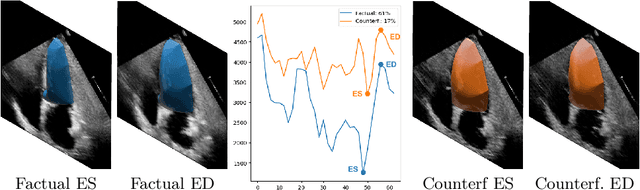
Abstract:Causally-enabled machine learning frameworks could help clinicians to identify the best course of treatments by answering counterfactual questions. We explore this path for the case of echocardiograms by looking into the variation of the Left Ventricle Ejection Fraction, the most essential clinical metric gained from these examinations. We combine deep neural networks, twin causal networks and generative adversarial methods for the first time to build D'ARTAGNAN (Deep ARtificial Twin-Architecture GeNerAtive Networks), a novel causal generative model. We demonstrate the soundness of our approach on a synthetic dataset before applying it to cardiac ultrasound videos by answering the question: "What would this echocardiogram look like if the patient had a different ejection fraction?". To do so, we generate new ultrasound videos, retaining the video style and anatomy of the original patient, with variations of the Ejection Fraction conditioned on a given input. We achieve an SSIM score of 0.79 and an R2 score of 0.51 on the counterfactual videos. Code and models are available at https://github.com/HReynaud/dartagnan.
CAS-Net: Conditional Atlas Generation and Brain Segmentation for Fetal MRI
May 17, 2022Abstract:Fetal Magnetic Resonance Imaging (MRI) is used in prenatal diagnosis and to assess early brain development. Accurate segmentation of the different brain tissues is a vital step in several brain analysis tasks, such as cortical surface reconstruction and tissue thickness measurements. Fetal MRI scans, however, are prone to motion artifacts that can affect the correctness of both manual and automatic segmentation techniques. In this paper, we propose a novel network structure that can simultaneously generate conditional atlases and predict brain tissue segmentation, called CAS-Net. The conditional atlases provide anatomical priors that can constrain the segmentation connectivity, despite the heterogeneity of intensity values caused by motion or partial volume effects. The proposed method is trained and evaluated on 253 subjects from the developing Human Connectome Project (dHCP). The results demonstrate that the proposed method can generate conditional age-specific atlas with sharp boundary and shape variance. It also segment multi-category brain tissues for fetal MRI with a high overall Dice similarity coefficient (DSC) of $85.2\%$ for the selected 9 tissue labels.
CortexODE: Learning Cortical Surface Reconstruction by Neural ODEs
Feb 16, 2022
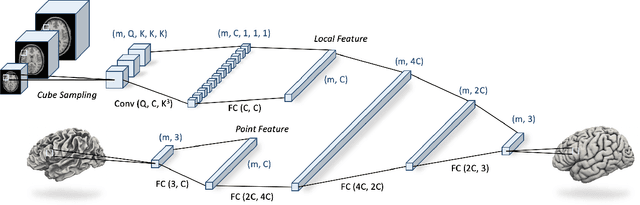
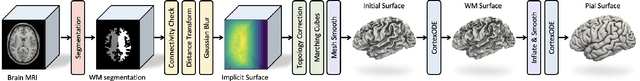
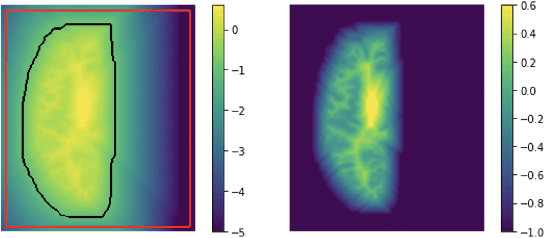
Abstract:We present CortexODE, a deep learning framework for cortical surface reconstruction. CortexODE leverages neural ordinary different equations (ODEs) to deform an input surface into a target shape by learning a diffeomorphic flow. The trajectories of the points on the surface are modeled as ODEs, where the derivatives of their coordinates are parameterized via a learnable Lipschitz-continuous deformation network. This provides theoretical guarantees for the prevention of self-intersections. CortexODE can be integrated to an automatic learning-based pipeline, which reconstructs cortical surfaces efficiently in less than 6 seconds. The pipeline utilizes a 3D U-Net to predict a white matter segmentation from brain Magnetic Resonance Imaging (MRI) scans, and further generates a signed distance function that represents an initial surface. Fast topology correction is introduced to guarantee homeomorphism to a sphere. Following the isosurface extraction step, two CortexODE models are trained to deform the initial surface to white matter and pial surfaces respectively. The proposed pipeline is evaluated on large-scale neuroimage datasets in various age groups including neonates (25-45 weeks), young adults (22-36 years) and elderly subjects (55-90 years). Our experiments demonstrate that the CortexODE-based pipeline can achieve less than 0.2mm average geometric error while being orders of magnitude faster compared to conventional processing pipelines.
Cross Modality 3D Navigation Using Reinforcement Learning and Neural Style Transfer
Nov 05, 2021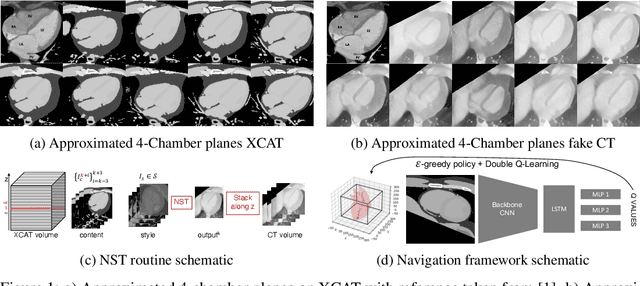
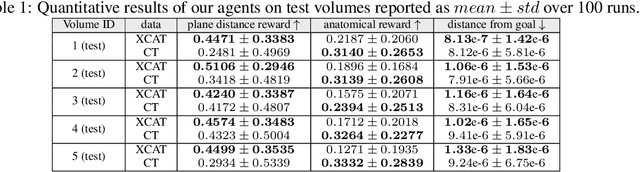
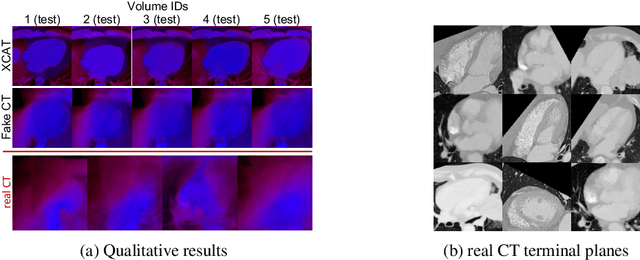
Abstract:This paper presents the use of Multi-Agent Reinforcement Learning (MARL) to perform navigation in 3D anatomical volumes from medical imaging. We utilize Neural Style Transfer to create synthetic Computed Tomography (CT) agent gym environments and assess the generalization capabilities of our agents to clinical CT volumes. Our framework does not require any labelled clinical data and integrates easily with several image translation techniques, enabling cross modality applications. Further, we solely condition our agents on 2D slices, breaking grounds for 3D guidance in much more difficult imaging modalities, such as ultrasound imaging. This is an important step towards user guidance during the acquisition of standardised diagnostic view planes, improving diagnostic consistency and facilitating better case comparison.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge