Arthur Gretton
E3S
Learning Deep Features in Instrumental Variable Regression
Nov 01, 2020
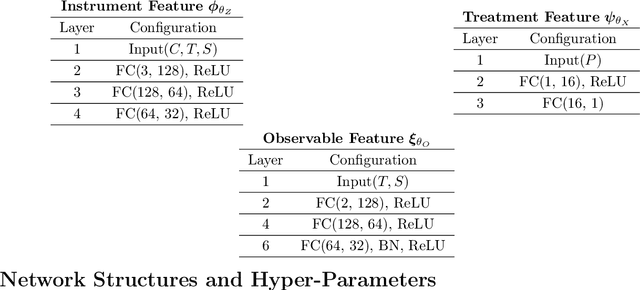
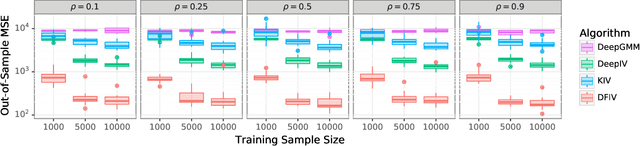

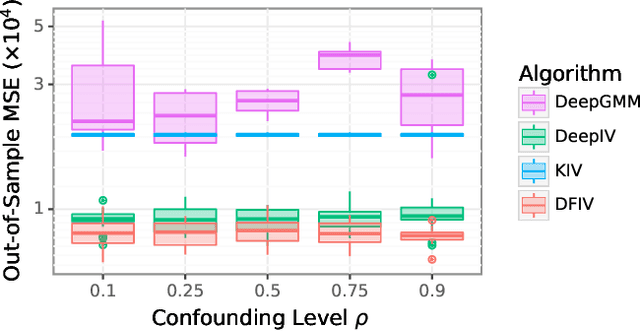
Abstract:Instrumental variable (IV) regression is a standard strategy for learning causal relationships between confounded treatment and outcome variables from observational data by utilizing an instrumental variable, which affects the outcome only through the treatment. In classical IV regression, learning proceeds in two stages: stage 1 performs linear regression from the instrument to the treatment; and stage 2 performs linear regression from the treatment to the outcome, conditioned on the instrument. We propose a novel method, deep feature instrumental variable regression (DFIV), to address the case where relations between instruments, treatments, and outcomes may be nonlinear. In this case, deep neural nets are trained to define informative nonlinear features on the instruments and treatments. We propose an alternating training regime for these features to ensure good end-to-end performance when composing stages 1 and 2, thus obtaining highly flexible feature maps in a computationally efficient manner. DFIV outperforms recent state-of-the-art methods on challenging IV benchmarks, including settings involving high dimensional image data. DFIV also exhibits competitive performance in off-policy policy evaluation for reinforcement learning, which can be understood as an IV regression task.
A Weaker Faithfulness Assumption based on Triple Interactions
Oct 27, 2020
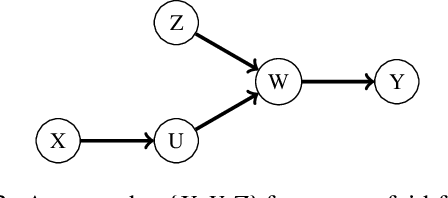
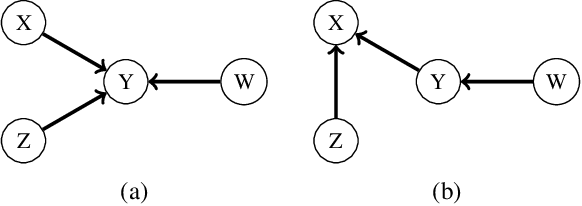
Abstract:One of the core assumptions in causal discovery is the faithfulness assumption---i.e. assuming that independencies found in the data are due to separations in the true causal graph. This assumption can, however, be violated in many ways, including xor connections, deterministic functions or cancelling paths. In this work, we propose a weaker assumption that we call 2-adjacency faithfulness. In contrast to adjacency faithfulness, which assumes that there is no conditional independence between each pair of variables that are connected in the causal graph, we only require no conditional independence between a node and a subset of its Markov blanket that can contain up to two nodes. Equivalently, we adapt orientation faithfulness to this setting. We further propose a sound orientation rule for causal discovery that applies under weaker assumptions. As a proof of concept, we derive a modified Grow and Shrink algorithm that recovers the Markov blanket of a target node and prove its correctness under strictly weaker assumptions than the standard faithfulness assumption.
Kernel Methods for Policy Evaluation: Treatment Effects, Mediation Analysis, and Off-Policy Planning
Oct 13, 2020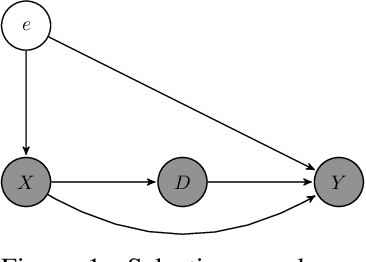
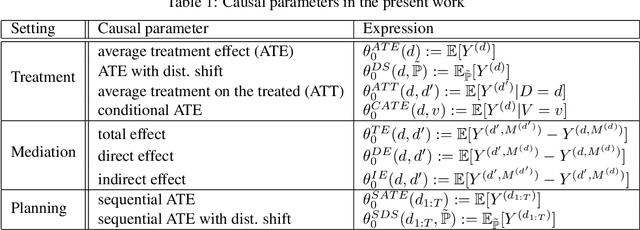
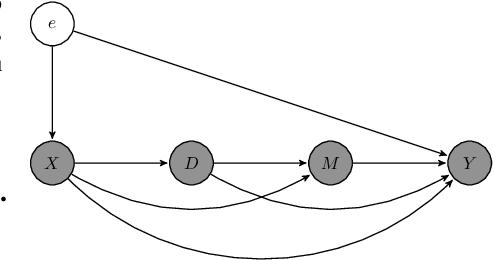
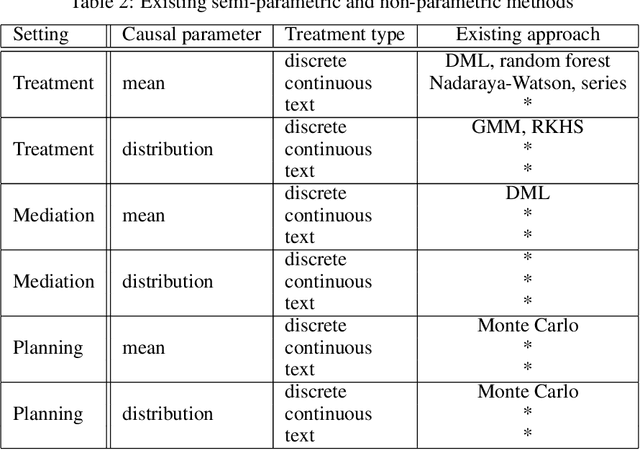
Abstract:We propose a novel framework for non-parametric policy evaluation in static and dynamic settings. Under the assumption of selection on observables, we consider treatment effects of the population, of sub-populations, and of alternative populations that may have alternative covariate distributions. We further consider the decomposition of a total effect into a direct effect and an indirect effect (as mediated by a particular mechanism). Under the assumption of sequential selection on observables, we consider the effects of sequences of treatments. Across settings, we allow for treatments that may be discrete, continuous, or even text. Across settings, we allow for estimation of not only counterfactual mean outcomes but also counterfactual distributions of outcomes. We unify analyses across settings by showing that all of these causal learning problems reduce to the re-weighting of a prediction, i.e. causal adjustment. We implement the re-weighting as an inner product in a function space called a reproducing kernel Hilbert space (RKHS), with a closed form solution that can be computed in one line of code. We prove uniform consistency and provide finite sample rates of convergence. We evaluate our estimators in simulations devised by other authors. We use our new estimators to evaluate continuous and heterogeneous treatment effects of the US Jobs Corps training program for disadvantaged youth.
Efficient Wasserstein Natural Gradients for Reinforcement Learning
Oct 12, 2020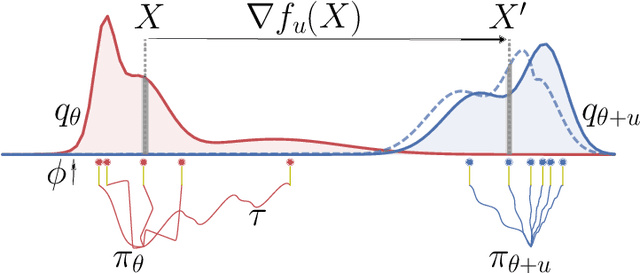
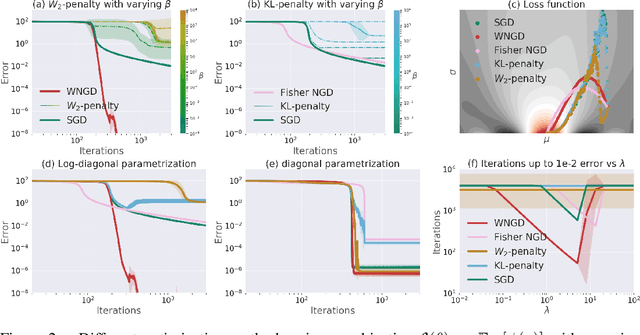
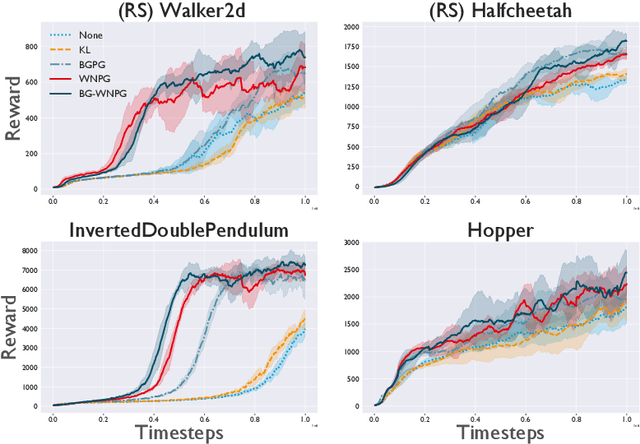
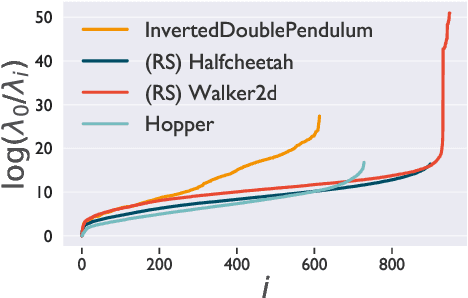
Abstract:A novel optimization approach is proposed for application to policy gradient methods and evolution strategies for reinforcement learning (RL). The procedure uses a computationally efficient Wasserstein natural gradient (WNG) descent that takes advantage of the geometry induced by a Wasserstein penalty to speed optimization. This method follows the recent theme in RL of including a divergence penalty in the objective to establish a trust region. Experiments on challenging tasks demonstrate improvements in both computational cost and performance over advanced baselines.
Kernelized Stein Discrepancy Tests of Goodness-of-fit for Time-to-Event Data
Aug 26, 2020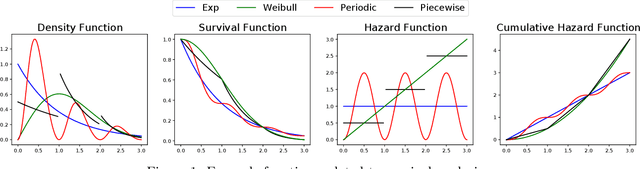

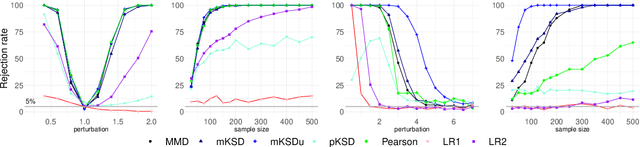

Abstract:Survival Analysis and Reliability Theory are concerned with the analysis of time-to-event data, in which observations correspond to waiting times until an event of interest such as death from a particular disease or failure of a component in a mechanical system. This type of data is unique due to the presence of censoring, a type of missing data that occurs when we do not observe the actual time of the event of interest but, instead, we have access to an approximation for it given by random interval in which the observation is known to belong. Most traditional methods are not designed to deal with censoring, and thus we need to adapt them to censored time-to-event data. In this paper, we focus on non-parametric goodness-of-fit testing procedures based on combining the Stein's method and kernelized discrepancies. While for uncensored data, there is a natural way of implementing a kernelized Stein discrepancy test, for censored data there are several options, each of them with different advantages and disadvantages. In this paper, we propose a collection of kernelized Stein discrepancy tests for time-to-event data, and we study each of them theoretically and empirically; our experimental results show that our proposed methods perform better than existing tests, including previous tests based on a kernelized maximum mean discrepancy.
A Non-Asymptotic Analysis for Stein Variational Gradient Descent
Jun 17, 2020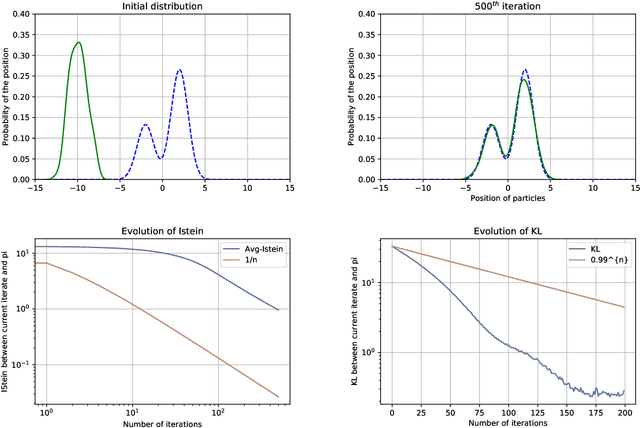
Abstract:We study the Stein Variational Gradient Descent (SVGD) algorithm, which optimises a set of particles to approximate a target probability distribution $\pi\propto e^{-V}$ on $\mathbb{R}^d$. In the population limit, SVGD performs gradient descent in the space of probability distributions on the KL divergence with respect to $\pi$, where the gradient is smoothed through a kernel integral operator. In this paper, we provide a novel finite time analysis for the SVGD algorithm. We obtain a descent lemma establishing that the algorithm decreases the objective at each iteration, and provably converges, with less restrictive assumptions on the step size than required in earlier analyses. We further provide a guarantee on the convergence rate in Kullback-Leibler divergence, assuming $\pi$ satisfies a Stein log-Sobolev inequality as in Duncan et al. (2019), which takes into account the geometry induced by the smoothed KL gradient.
Layer-wise Learning of Kernel Dependence Networks
Jun 15, 2020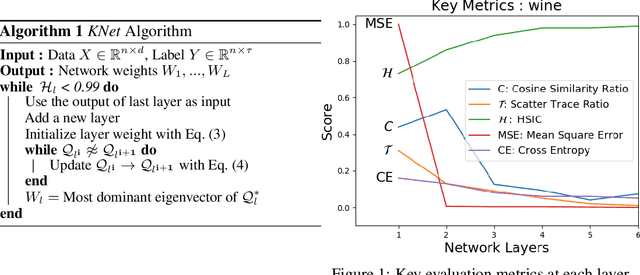
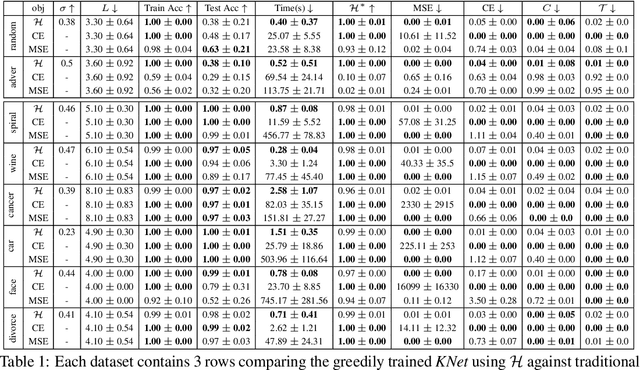
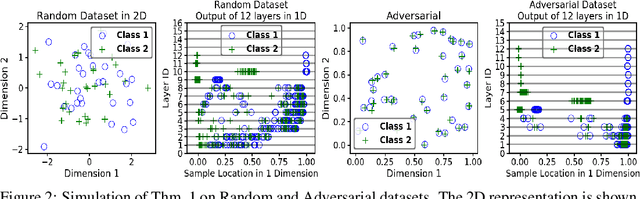
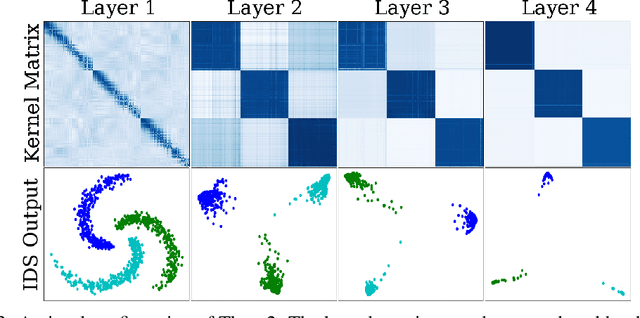
Abstract:We propose a greedy strategy to train a deep network for multi-class classification, where each layer is defined as a composition of a linear projection and a nonlinear mapping. This nonlinear mapping is defined as the feature map of a Gaussian kernel, and the linear projection is learned by maximizing the dependence between the layer output and the labels, using the Hilbert Schmidt Independence Criterion (HSIC) as the dependence measure. Since each layer is trained greedily in sequence, all learning is local, and neither backpropagation nor even gradient descent is needed. The depth and width of the network are determined via natural guidelines, and the procedure regularizes its weights in the linear layer. As the key theoretical result, the function class represented by the network is proved to be sufficiently rich to learn any dataset labeling using a finite number of layers, in the sense of reaching minimum mean-squared error or cross-entropy, as long as no two data points with different labels coincide. Experiments demonstrate good generalization performance of the greedy approach across multiple benchmarks while showing a significant computational advantage against a multilayer perceptron of the same complexity trained globally by backpropagation.
KALE: When Energy-Based Learning Meets Adversarial Training
Mar 10, 2020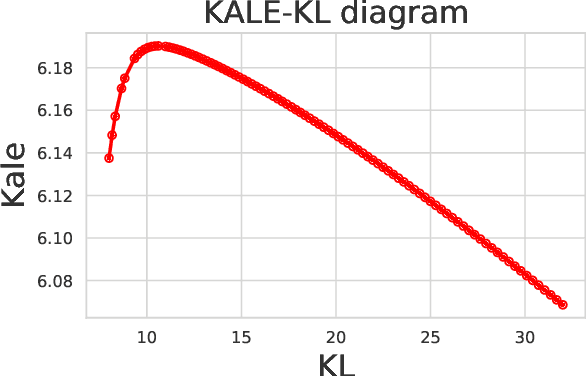
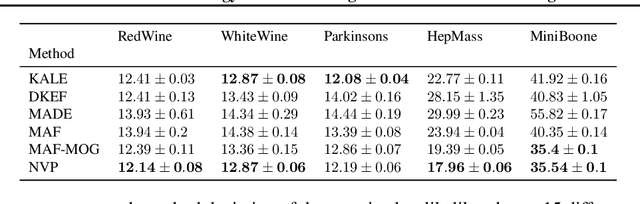
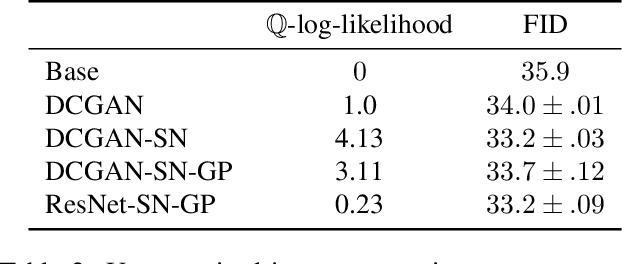
Abstract:Legendre duality provides a variational lower-bound for the Kullback-Leibler divergence (KL) which can be estimated using samples, without explicit knowledge of the density ratio. We use this estimator, the \textit{KL Approximate Lower-bound Estimate} (KALE), in a contrastive setting for learning energy-based models, and show that it provides a maximum likelihood estimate (MLE). We then extend this procedure to adversarial training, where the discriminator represents the energy and the generator is the base measure of the energy-based model. Unlike in standard generative adversarial networks (GANs), the learned model makes use of both generator and discriminator to generate samples. This is achieved using Hamiltonian Monte Carlo in the latent space of the generator, using information from the discriminator, to find regions in that space that produce better quality samples. We also show that, unlike the KL, KALE enjoys smoothness properties that make it suitable for adversarial training, and provide convergence rates for KALE when the negative log density ratio belongs to the variational family. Finally, we demonstrate the effectiveness of this approach on simple datasets.
Learning Deep Kernels for Non-Parametric Two-Sample Tests
Feb 21, 2020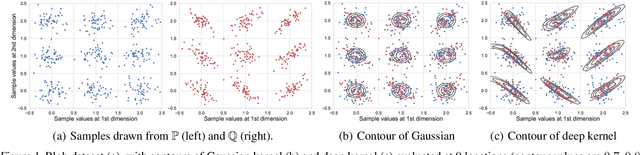
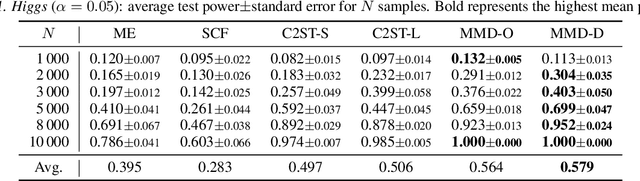
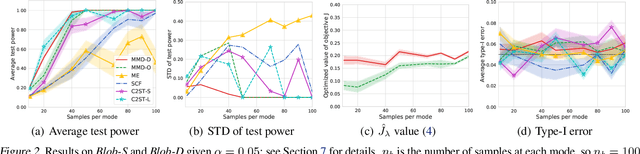
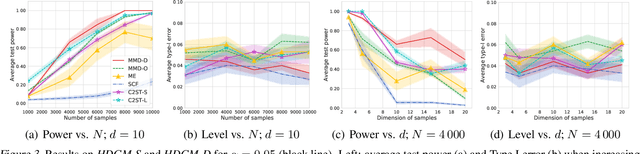
Abstract:We propose a class of kernel-based two-sample tests, which aim to determine whether two sets of samples are drawn from the same distribution. Our tests are constructed from kernels parameterized by deep neural nets, trained to maximize test power. These tests adapt to variations in distribution smoothness and shape over space, and are especially suited to high dimensions and complex data. By contrast, the simpler kernels used in prior kernel testing work are spatially homogeneous, and adaptive only in lengthscale. We explain how this scheme includes popular classifier-based two-sample tests as a special case, but improves on them in general. We provide the first proof of consistency for the proposed adaptation method, which applies both to kernels on deep features and to simpler radial basis kernels or multiple kernel learning. In experiments, we establish the superior performance of our deep kernels in hypothesis testing on benchmark and real-world data. The code of our deep-kernel-based two sample tests is available at https://github.com/fengliu90/DK-for-TST.
A kernel log-rank test of independence for right-censored data
Dec 08, 2019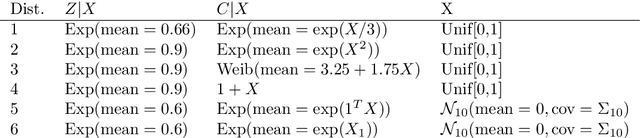
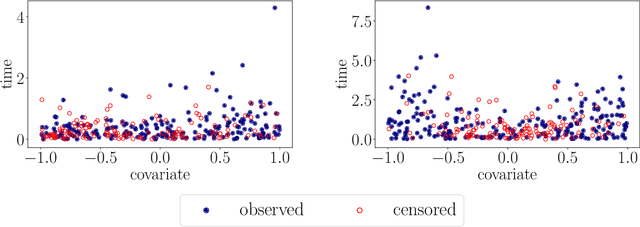
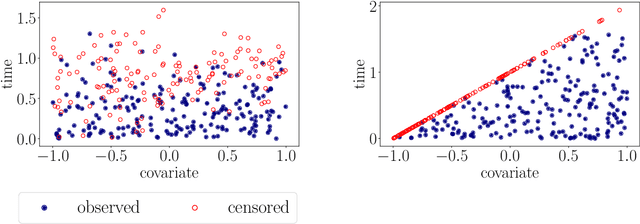
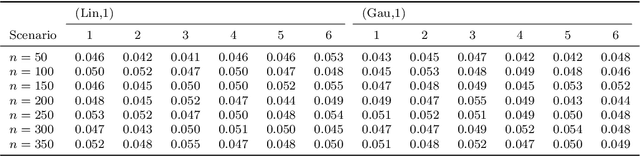
Abstract:With the incorporation of new data gathering methods in clinical research, it becomes fundamental for survival analysis techniques to deal with high-dimensional or/and non-standard covariates. In this paper we introduce a general non-parametric independence test between right-censored survival times and covariates taking values on a general (not necessarily Euclidean) space $\mathcal{X}$. We show that our test statistic has a dual interpretation, first in terms of the supremum of a potentially infinite collection of weight-indexed log-rank tests, with weight functions belonging to a reproducing kernel Hilbert space (RKHS) of functions; and second, as the norm of the difference of embeddings of certain finite measures into the RKHS, similar to the Hilbert-Schmidt Independence Criterion (HSIC) test-statistic. We study the asymptotic properties of the test, finding sufficient conditions to ensure that our test is omnibus. The test statistic can be computed straightforwardly, and the rejection threshold is obtained via an asymptotically consistent Wild-Bootstrap procedure. We perform extensive simulations demonstrating that our testing procedure generally performs better than competing approaches in detecting complex nonlinear dependence.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge