"Time": models, code, and papers
Explaining random forest prediction through diverse rulesets
Mar 29, 2022

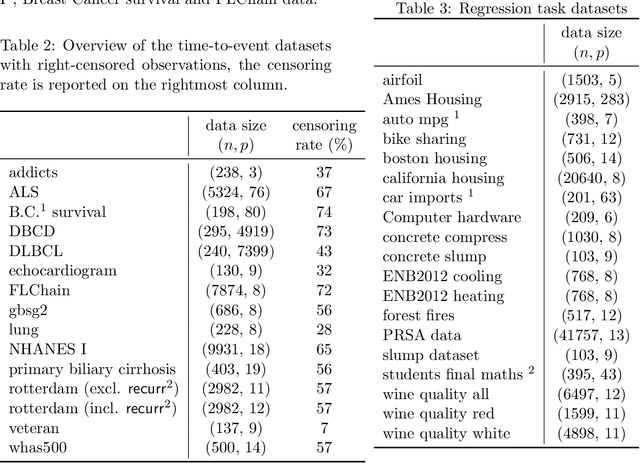
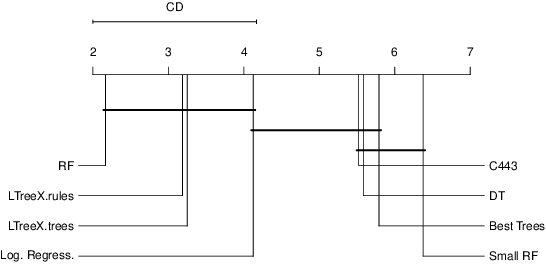
Tree-ensemble algorithms, such as random forest, are effective machine learning methods popular for their flexibility, high performance, and robustness to overfitting. However, since multiple learners are combined,they are not as interpretable as a single decision tree. In this work we propose a methodology, called Local Tree eXtractor (LTreeX) which is able to explain the forest prediction for a given test instance with a few diverse rules. Starting from the decision trees generated by a random forest, our method 1) pre-selects a subset of them, 2) creates a vector representation, and 3) eventually clusters such a representation. Each cluster prototype results in a rule that explains the test instance prediction. We test the effectiveness of LTreeX on 71 real-world datasets and we demonstrate the validity of our approach for binary classification, regression, multi-label classification and time-to-event tasks. In all set-ups, we show that our extracted surrogate model manages to approximate the performance of the corresponding ensemble model, while selecting only few trees from the whole forest.We also show that our proposed approach substantially outperforms other explainable methods in terms of predictive performance.
Fruit Mapping with Shape Completion for Autonomous Crop Monitoring
Mar 29, 2022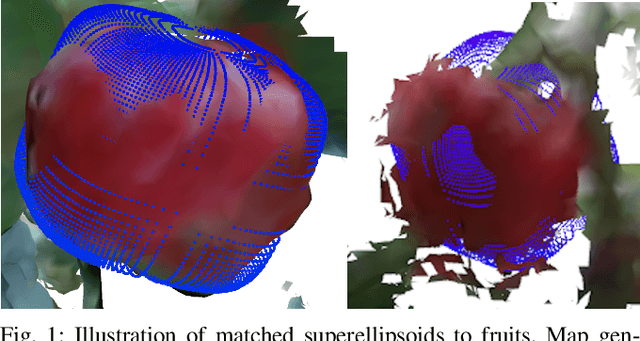
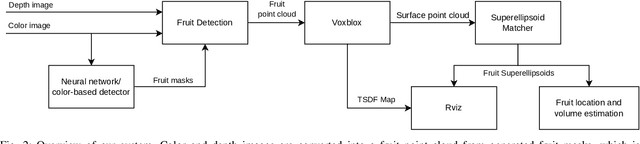
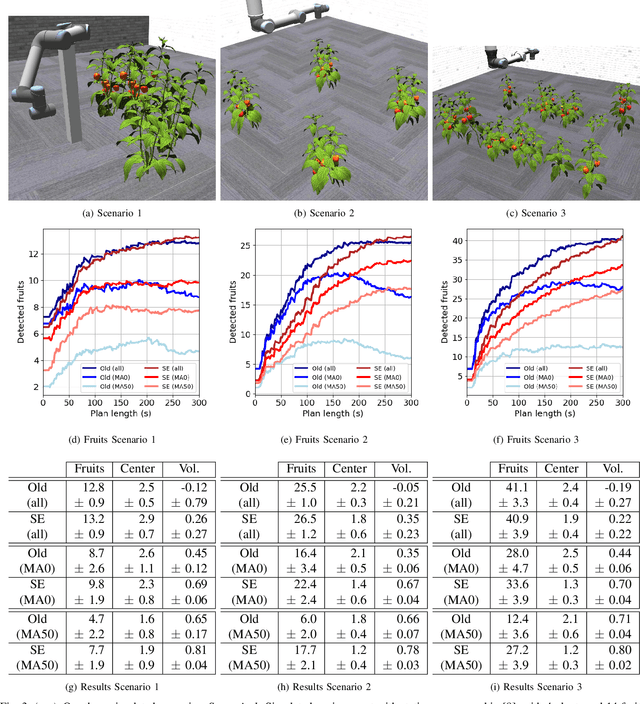
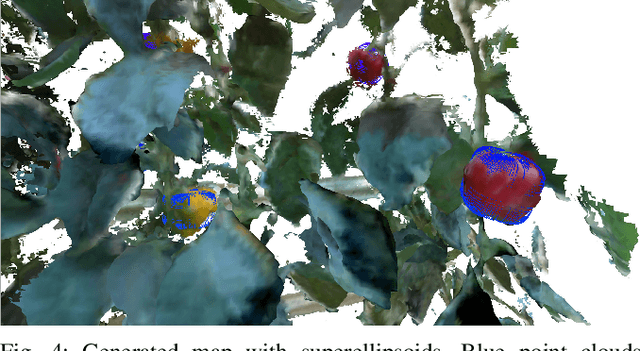
Autonomous crop monitoring is a difficult task due to the complex structure of plants. Occlusions from leaves can make it impossible to obtain complete views about all fruits of, e.g., pepper plants. Therefore, accurately estimating the shape and volume of fruits from partial information is crucial to enable further advanced automation tasks such as yield estimation and automated fruit picking. In this paper, we present an approach for mapping fruits on plants and estimating their shape by matching superellipsoids. Our system segments fruits in images and uses their masks to generate point clouds of the fruits. To combine sequences of acquired point clouds, we utilize a real-time 3D mapping framework and build up a fruit map based on truncated signed distance fields. We cluster fruits from this map and use optimized superellipsoids for matching to obtain accurate shape estimates. In our experiments, we show in various simulated scenarios with a robotic arm equipped with an RGB-D camera that our approach can accurately estimate fruit volumes. Additionally, we provide qualitative results of estimated fruit shapes from data recorded in a commercial glasshouse environment.
Learning Structured Gaussians to Approximate Deep Ensembles
Mar 29, 2022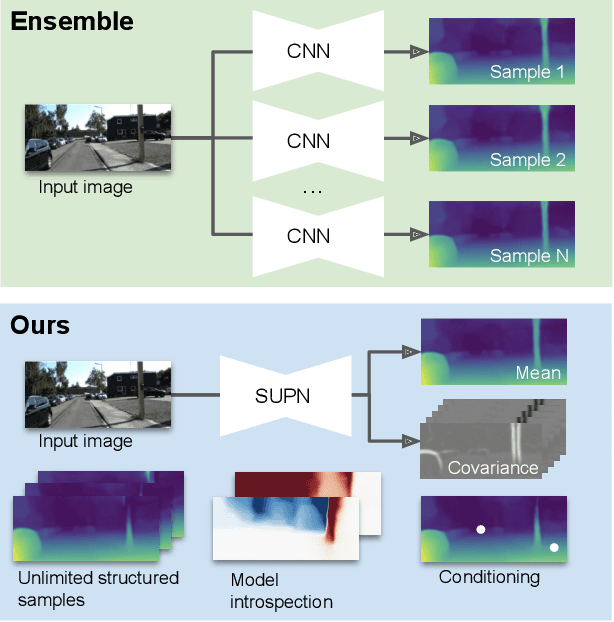
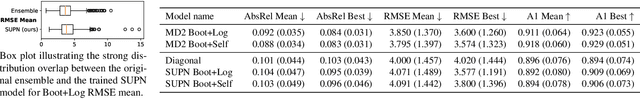
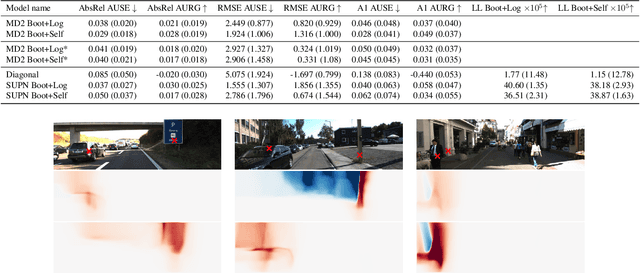
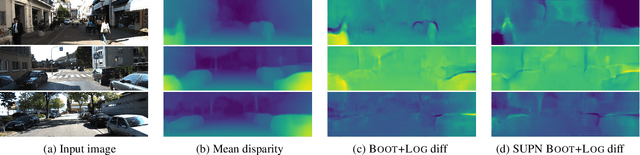
This paper proposes using a sparse-structured multivariate Gaussian to provide a closed-form approximator for the output of probabilistic ensemble models used for dense image prediction tasks. This is achieved through a convolutional neural network that predicts the mean and covariance of the distribution, where the inverse covariance is parameterised by a sparsely structured Cholesky matrix. Similarly to distillation approaches, our single network is trained to maximise the probability of samples from pre-trained probabilistic models, in this work we use a fixed ensemble of networks. Once trained, our compact representation can be used to efficiently draw spatially correlated samples from the approximated output distribution. Importantly, this approach captures the uncertainty and structured correlations in the predictions explicitly in a formal distribution, rather than implicitly through sampling alone. This allows direct introspection of the model, enabling visualisation of the learned structure. Moreover, this formulation provides two further benefits: estimation of a sample probability, and the introduction of arbitrary spatial conditioning at test time. We demonstrate the merits of our approach on monocular depth estimation and show that the advantages of our approach are obtained with comparable quantitative performance.
Identifying Suitable Tasks for Inductive Transfer Through the Analysis of Feature Attributions
Feb 02, 2022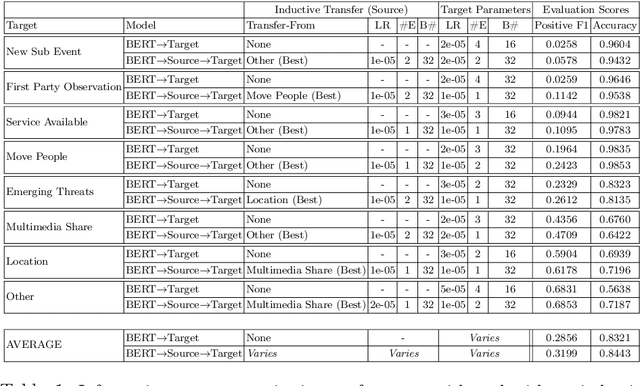
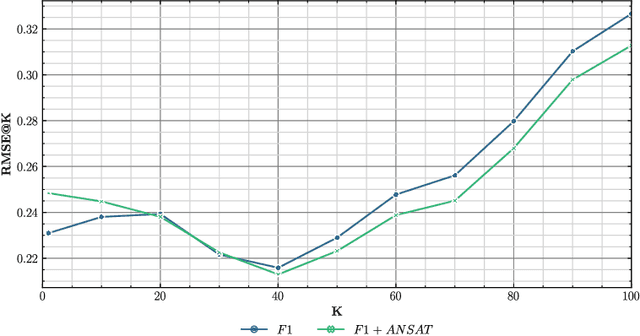
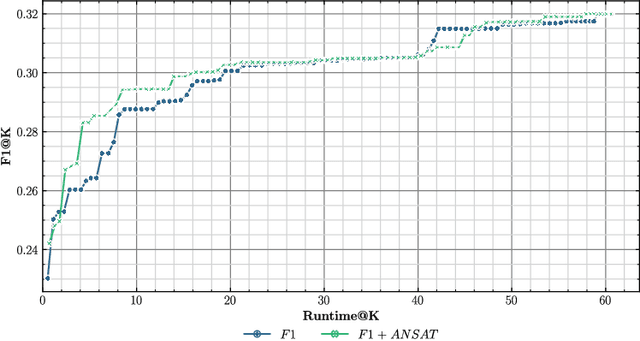
Transfer learning approaches have shown to significantly improve performance on downstream tasks. However, it is common for prior works to only report where transfer learning was beneficial, ignoring the significant trial-and-error required to find effective settings for transfer. Indeed, not all task combinations lead to performance benefits, and brute-force searching rapidly becomes computationally infeasible. Hence the question arises, can we predict whether transfer between two tasks will be beneficial without actually performing the experiment? In this paper, we leverage explainability techniques to effectively predict whether task pairs will be complementary, through comparison of neural network activation between single-task models. In this way, we can avoid grid-searches over all task and hyperparameter combinations, dramatically reducing the time needed to find effective task pairs. Our results show that, through this approach, it is possible to reduce training time by up to 83.5% at a cost of only 0.034 reduction in positive-class F1 on the TREC-IS 2020-A dataset.
Parameterized Intractability for Multi-Winner Election under the Chamberlin-Courant Rule and the Monroe Rule
Feb 24, 2022Answering an open question by Betzler et al. [Betzler et al., JAIR'13], we resolve the parameterized complexity of the multi-winner determination problem under two famous representation voting rules: the Chamberlin-Courant (in short CC) rule [Chamberlin and Courant, APSR'83] and the Monroe rule [Monroe, APSR'95]. We show that under both rules, the problem is W[1]-hard with respect to the sum $\beta$ of misrepresentations, thereby precluding the existence of any $f(\beta) \cdot |I|^{O(1)}$ -time algorithm, where $|I|$ denotes the size of the input instance.
SRP-DNN: Learning Direct-Path Phase Difference for Multiple Moving Sound Source Localization
Feb 16, 2022
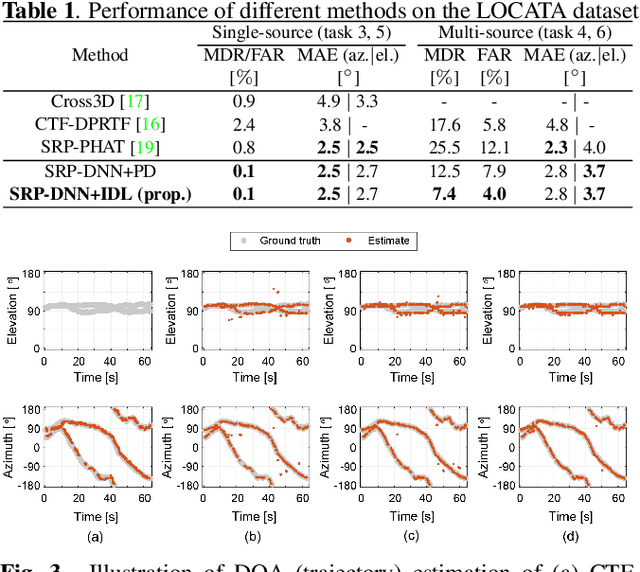
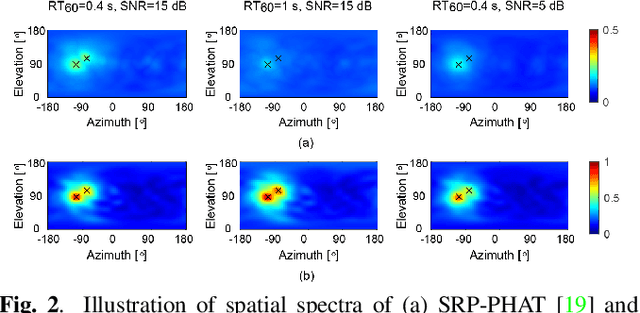
Multiple moving sound source localization in real-world scenarios remains a challenging issue due to interaction between sources, time-varying trajectories, distorted spatial cues, etc. In this work, we propose to use deep learning techniques to learn competing and time-varying direct-path phase differences for localizing multiple moving sound sources. A causal convolutional recurrent neural network is designed to extract the direct-path phase difference sequence from signals of each microphone pair. To avoid the assignment ambiguity and the problem of uncertain output-dimension encountered when simultaneously predicting multiple targets, the learning target is designed in a weighted sum format, which encodes source activity in the weight and direct-path phase differences in the summed value. The learned direct-path phase differences for all microphone pairs can be directly used to construct the spatial spectrum according to the formulation of steered response power (SRP). This deep neural network (DNN) based SRP method is referred to as SRP-DNN. The locations of sources are estimated by iteratively detecting and removing the dominant source from the spatial spectrum, in which way the interaction between sources is reduced. Experimental results on both simulated and real-world data show the superiority of the proposed method in the presence of noise and reverberation.
PGNet: Real-time Arbitrarily-Shaped Text Spotting with Point Gathering Network
Apr 12, 2021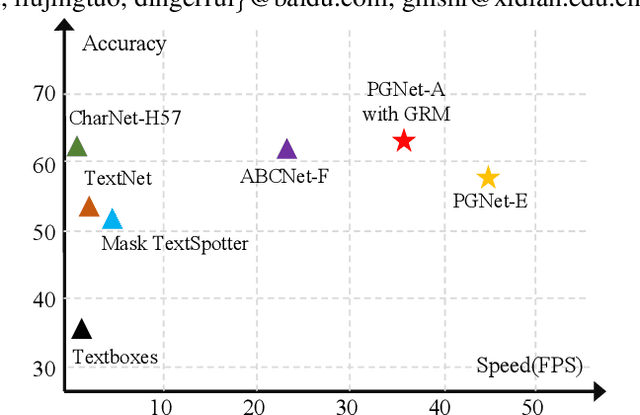
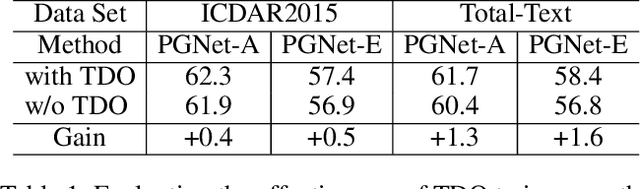
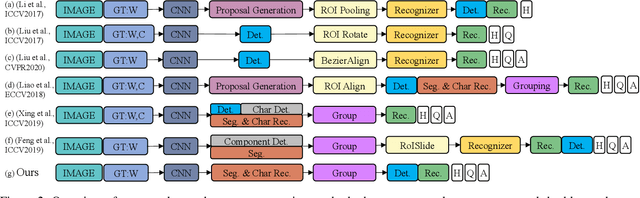
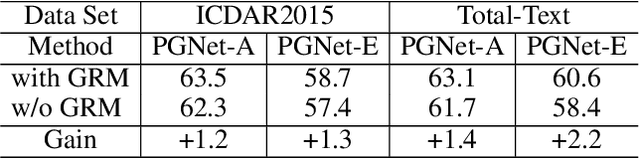
The reading of arbitrarily-shaped text has received increasing research attention. However, existing text spotters are mostly built on two-stage frameworks or character-based methods, which suffer from either Non-Maximum Suppression (NMS), Region-of-Interest (RoI) operations, or character-level annotations. In this paper, to address the above problems, we propose a novel fully convolutional Point Gathering Network (PGNet) for reading arbitrarily-shaped text in real-time. The PGNet is a single-shot text spotter, where the pixel-level character classification map is learned with proposed PG-CTC loss avoiding the usage of character-level annotations. With a PG-CTC decoder, we gather high-level character classification vectors from two-dimensional space and decode them into text symbols without NMS and RoI operations involved, which guarantees high efficiency. Additionally, reasoning the relations between each character and its neighbors, a graph refinement module (GRM) is proposed to optimize the coarse recognition and improve the end-to-end performance. Experiments prove that the proposed method achieves competitive accuracy, meanwhile significantly improving the running speed. In particular, in Total-Text, it runs at 46.7 FPS, surpassing the previous spotters with a large margin.
Giga-scale Kernel Matrix Vector Multiplication on GPU
Feb 02, 2022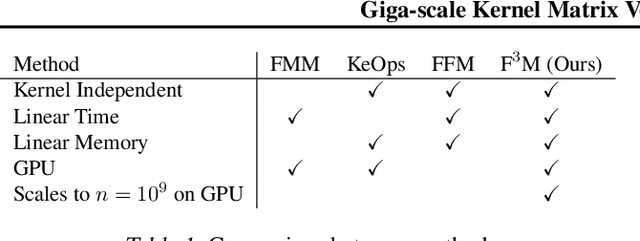
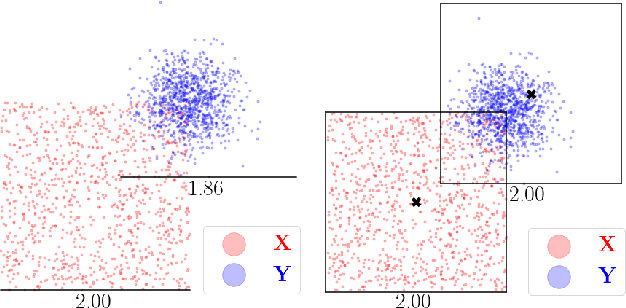
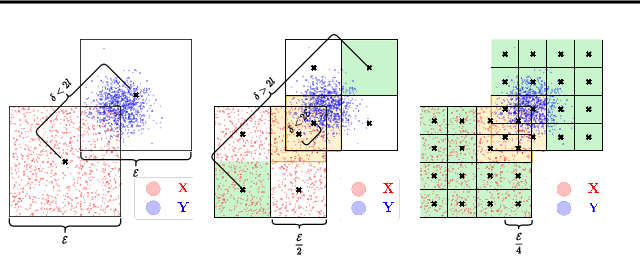
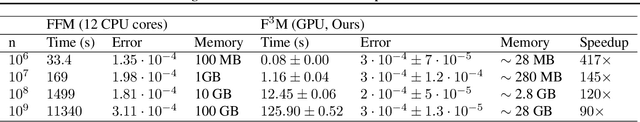
Kernel matrix vector multiplication (KMVM) is a ubiquitous operation in machine learning and scientific computing, spanning from the kernel literature to signal processing. As kernel matrix vector multiplication tends to scale quadratically in both memory and time, applications are often limited by these computational scaling constraints. We propose a novel approximation procedure coined Faster-Fast and Free Memory Method ($\text{F}^3$M) to address these scaling issues for KMVM. Extensive experiments demonstrate that $\text{F}^3$M has empirical \emph{linear time and memory} complexity with a relative error of order $10^{-3}$ and can compute a full KMVM for a billion points \emph{in under one minute} on a high-end GPU, leading to a significant speed-up in comparison to existing CPU methods. We further demonstrate the utility of our procedure by applying it as a drop-in for the state-of-the-art GPU-based linear solver FALKON, \emph{improving speed 3-5 times} at the cost of $<$1\% drop in accuracy.
Learning Whole Heart Mesh Generation From Patient Images For Computational Simulations
Mar 20, 2022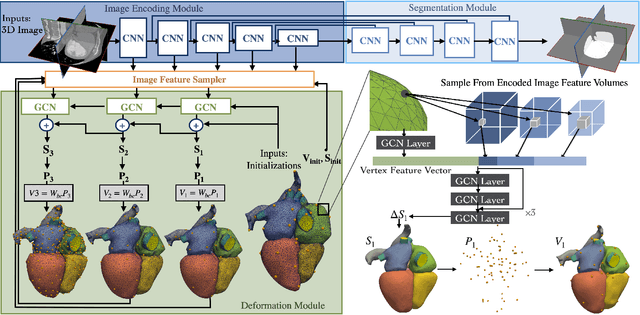
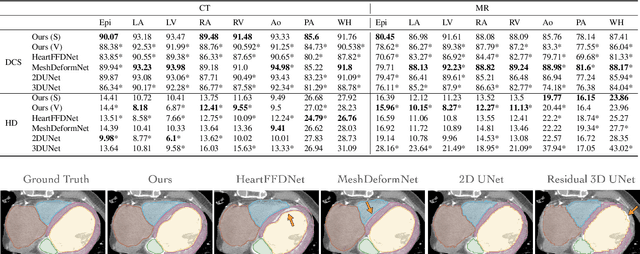
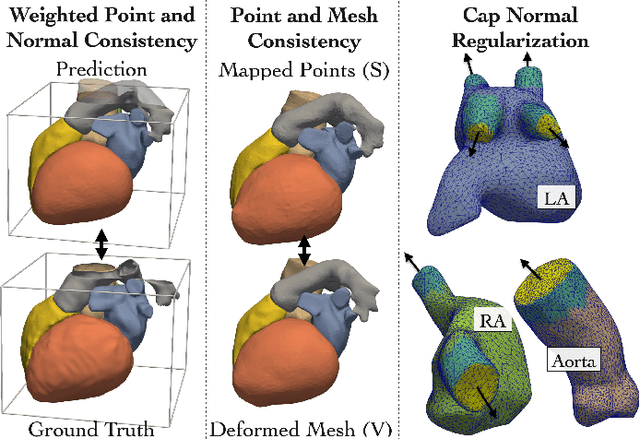
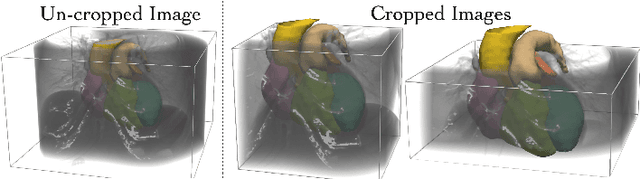
Patient-specific cardiac modeling combines geometries of the heart derived from medical images and biophysical simulations to predict various aspects of cardiac function. However, generating simulation-suitable models of the heart from patient image data often requires complicated procedures and significant human effort. We present a fast and automated deep-learning method to construct simulation-suitable models of the heart from medical images. The approach constructs meshes from 3D patient images by learning to deform a small set of deformation handles on a whole heart template. For both 3D CT and MR data, this method achieves promising accuracy for whole heart reconstruction, consistently outperforming prior methods in constructing simulation-suitable meshes of the heart. When evaluated on time-series CT data, this method produced more anatomically and temporally consistent geometries than prior methods, and was able to produce geometries that better satisfy modeling requirements for cardiac flow simulations. Our source code will be available on GitHub.
Neural Ordinary Differential Equations for Nonlinear System Identification
Mar 15, 2022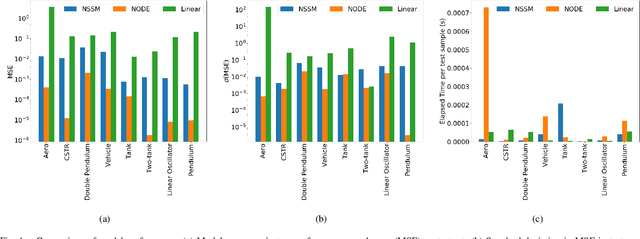
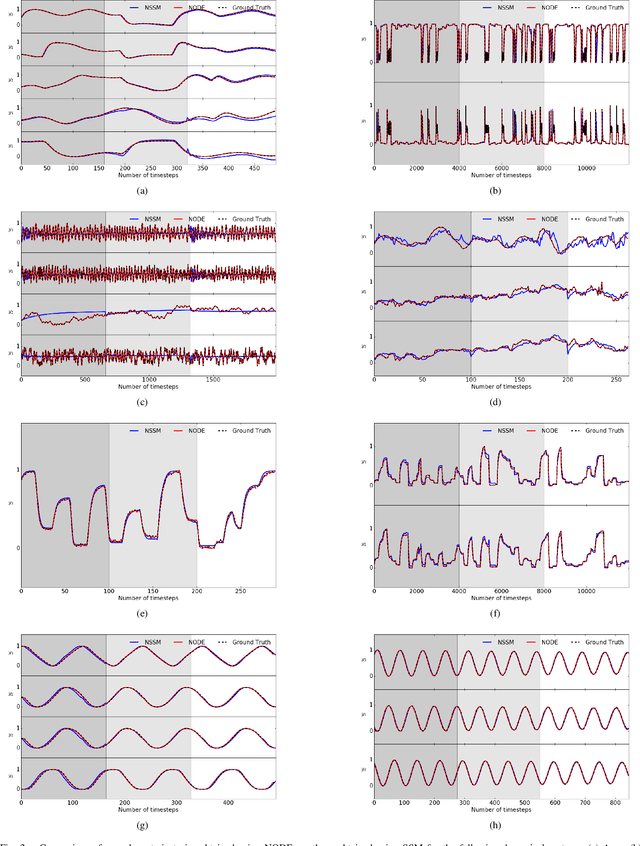
Neural ordinary differential equations (NODE) have been recently proposed as a promising approach for nonlinear system identification tasks. In this work, we systematically compare their predictive performance with current state-of-the-art nonlinear and classical linear methods. In particular, we present a quantitative study comparing NODE's performance against neural state-space models and classical linear system identification methods. We evaluate the inference speed and prediction performance of each method on open-loop errors across eight different dynamical systems. The experiments show that NODEs can consistently improve the prediction accuracy by an order of magnitude compared to benchmark methods. Besides improved accuracy, we also observed that NODEs are less sensitive to hyperparameters compared to neural state-space models. On the other hand, these performance gains come with a slight increase of computation at the inference time.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge