"Image": models, code, and papers
Neighborhood Contrastive Learning Applied to Online Patient Monitoring
Jun 09, 2021
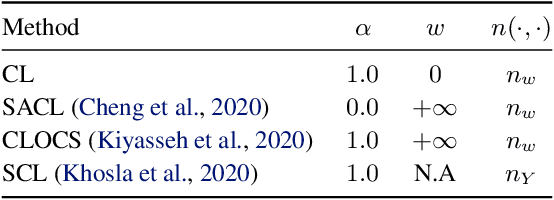
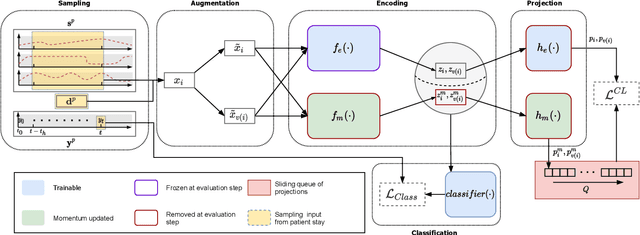
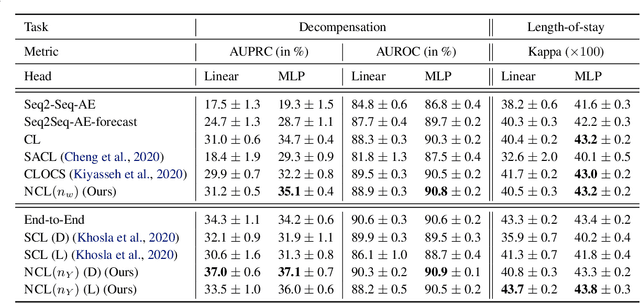
Intensive care units (ICU) are increasingly looking towards machine learning for methods to provide online monitoring of critically ill patients. In machine learning, online monitoring is often formulated as a supervised learning problem. Recently, contrastive learning approaches have demonstrated promising improvements over competitive supervised benchmarks. These methods rely on well-understood data augmentation techniques developed for image data which do not apply to online monitoring. In this work, we overcome this limitation by supplementing time-series data augmentation techniques with a novel contrastive learning objective which we call neighborhood contrastive learning (NCL). Our objective explicitly groups together contiguous time segments from each patient while maintaining state-specific information. Our experiments demonstrate a marked improvement over existing work applying contrastive methods to medical time-series.
External Prior Guided Internal Prior Learning for Real-World Noisy Image Denoising
Oct 15, 2018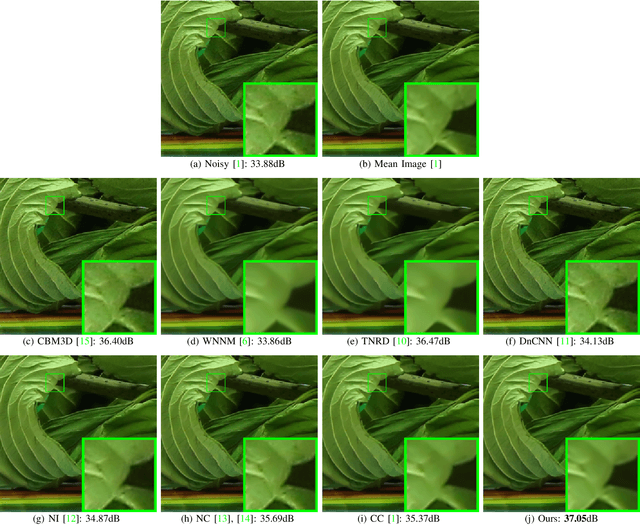
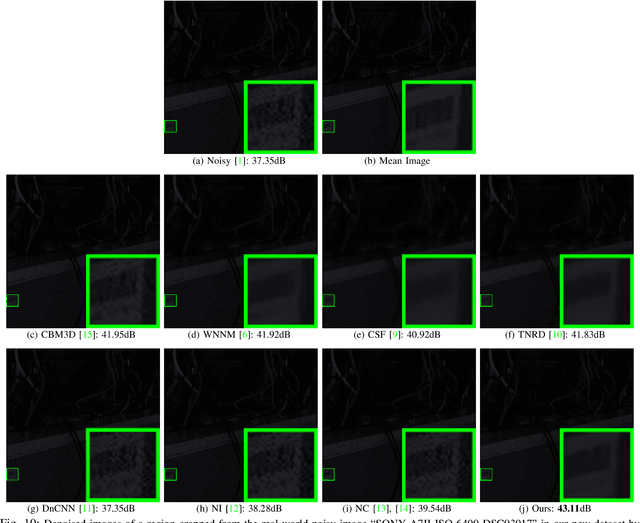
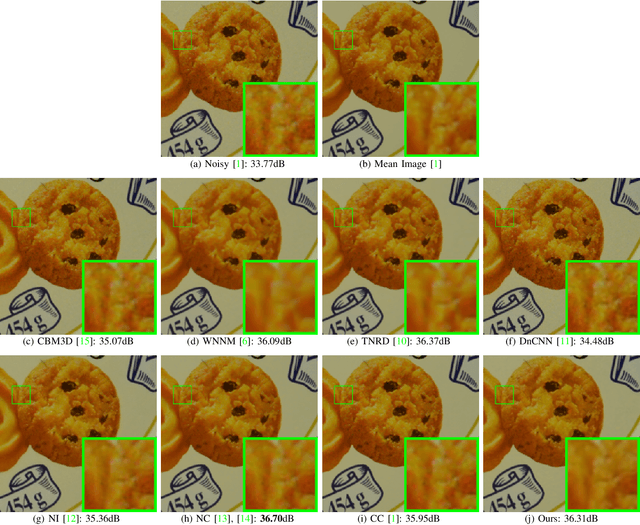



Most of existing image denoising methods learn image priors from either external data or the noisy image itself to remove noise. However, priors learned from external data may not be adaptive to the image to be denoised, while priors learned from the given noisy image may not be accurate due to the interference of corrupted noise. Meanwhile, the noise in real-world noisy images is very complex, which is hard to be described by simple distributions such as Gaussian distribution, making real-world noisy image denoising a very challenging problem. We propose to exploit the information in both external data and the given noisy image, and develop an external prior guided internal prior learning method for real-world noisy image denoising. We first learn external priors from an independent set of clean natural images. With the aid of learned external priors, we then learn internal priors from the given noisy image to refine the prior model. The external and internal priors are formulated as a set of orthogonal dictionaries to efficiently reconstruct the desired image. Extensive experiments are performed on several real-world noisy image datasets. The proposed method demonstrates highly competitive denoising performance, outperforming state-of-the-art denoising methods including those designed for real-world noisy images.
Regularized Evolution for Image Classifier Architecture Search
Oct 26, 2018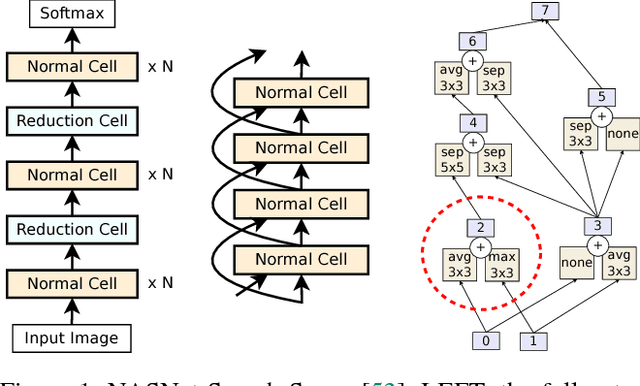
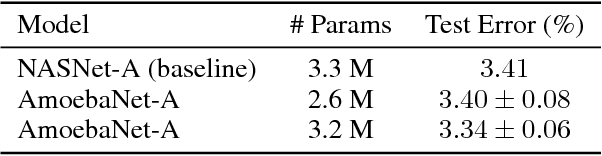
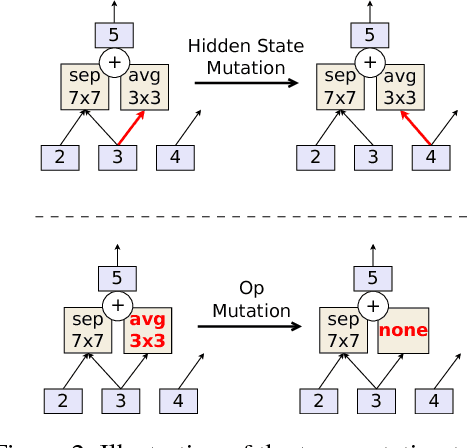
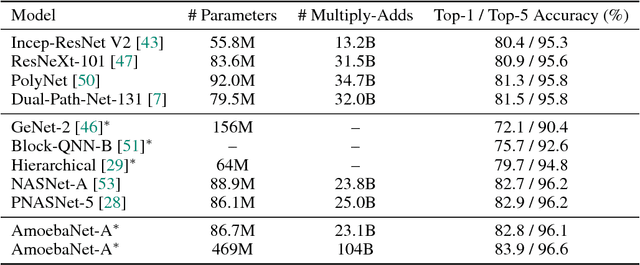
The effort devoted to hand-crafting neural network image classifiers has motivated the use of architecture search to discover them automatically. Although evolutionary algorithms have been repeatedly applied to neural network topologies, the image classifiers thus discovered have remained inferior to human-crafted ones. Here, we evolve an image classifier---AmoebaNet-A---that surpasses hand-designs for the first time. To do this, we modify the tournament selection evolutionary algorithm by introducing an age property to favor the younger genotypes. Matching size, AmoebaNet-A has comparable accuracy to current state-of-the-art ImageNet models discovered with more complex architecture-search methods. Scaled to larger size, AmoebaNet-A sets a new state-of-the-art 83.9% top-1 / 96.6% top-5 ImageNet accuracy. In a controlled comparison against a well known reinforcement learning algorithm, we give evidence that evolution can obtain results faster with the same hardware, especially at the earlier stages of the search. This is relevant when fewer compute resources are available. Evolution is, thus, a simple method to effectively discover high-quality architectures.
Generative Modeling for Multi-task Visual Learning
Jun 25, 2021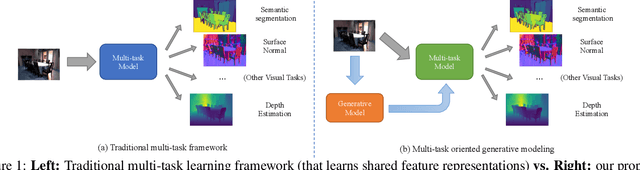
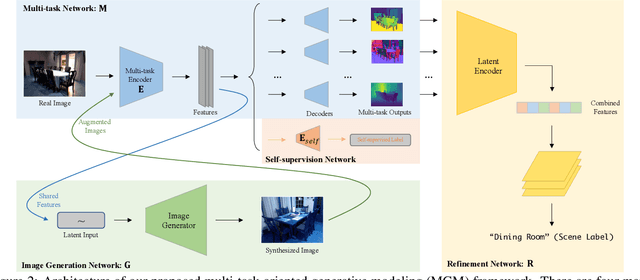
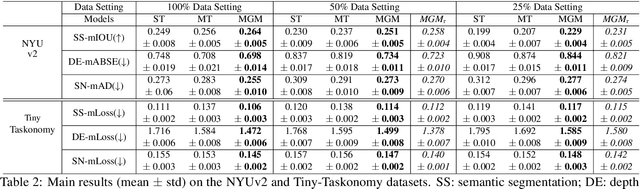
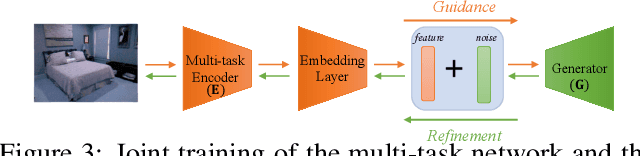
Generative modeling has recently shown great promise in computer vision, but it has mostly focused on synthesizing visually realistic images. In this paper, motivated by multi-task learning of shareable feature representations, we consider a novel problem of learning a shared generative model that is useful across various visual perception tasks. Correspondingly, we propose a general multi-task oriented generative modeling (MGM) framework, by coupling a discriminative multi-task network with a generative network. While it is challenging to synthesize both RGB images and pixel-level annotations in multi-task scenarios, our framework enables us to use synthesized images paired with only weak annotations (i.e., image-level scene labels) to facilitate multiple visual tasks. Experimental evaluation on challenging multi-task benchmarks, including NYUv2 and Taskonomy, demonstrates that our MGM framework improves the performance of all the tasks by large margins, consistently outperforming state-of-the-art multi-task approaches.
ICMSC: Intra- and Cross-modality Semantic Consistency for Unsupervised Domain Adaptation on Hip Joint Bone Segmentation
Dec 23, 2020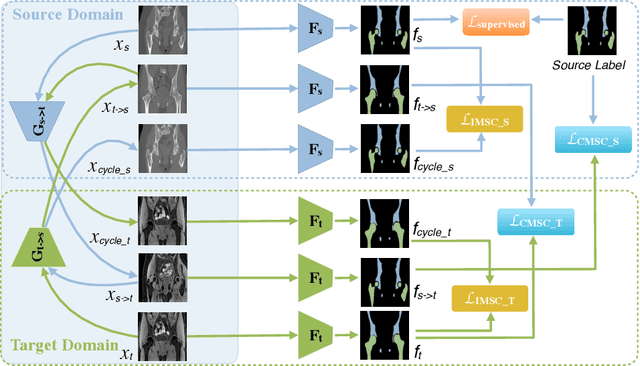
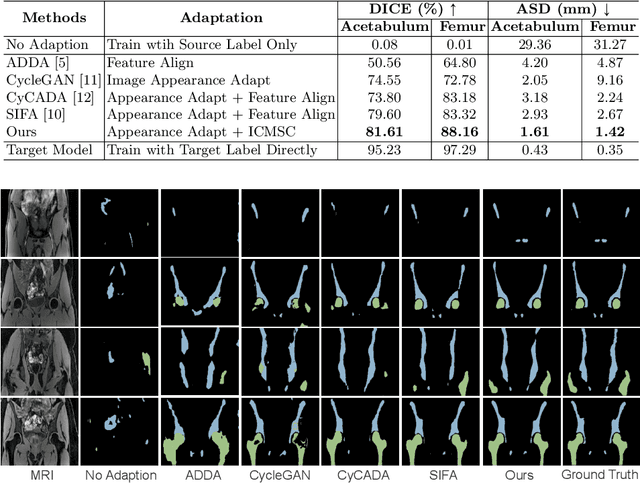
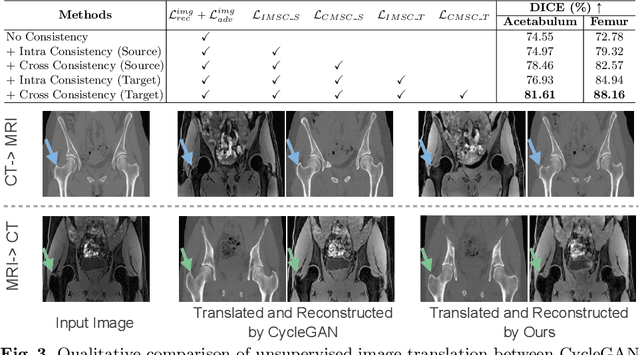
Unsupervised domain adaptation (UDA) for cross-modality medical image segmentation has shown great progress by domain-invariant feature learning or image appearance translation. Adapted feature learning usually cannot detect domain shifts at the pixel level and is not able to achieve good results in dense semantic segmentation tasks. Image appearance translation, e.g. CycleGAN, translates images into different styles with good appearance, despite its population, its semantic consistency is hardly to maintain and results in poor cross-modality segmentation. In this paper, we propose intra- and cross-modality semantic consistency (ICMSC) for UDA and our key insight is that the segmentation of synthesised images in different styles should be consistent. Specifically, our model consists of an image translation module and a domain-specific segmentation module. The image translation module is a standard CycleGAN, while the segmentation module contains two domain-specific segmentation networks. The intra-modality semantic consistency (IMSC) forces the reconstructed image after a cycle to be segmented in the same way as the original input image, while the cross-modality semantic consistency (CMSC) encourages the synthesized images after translation to be segmented exactly the same as before translation. Comprehensive experimental results on cross-modality hip joint bone segmentation show the effectiveness of our proposed method, which achieves an average DICE of 81.61% on the acetabulum and 88.16% on the proximal femur, outperforming other state-of-the-art methods. It is worth to note that without UDA, a model trained on CT for hip joint bone segmentation is non-transferable to MRI and has almost zero-DICE segmentation.
Exploiting Raw Images for Real-Scene Super-Resolution
Feb 02, 2021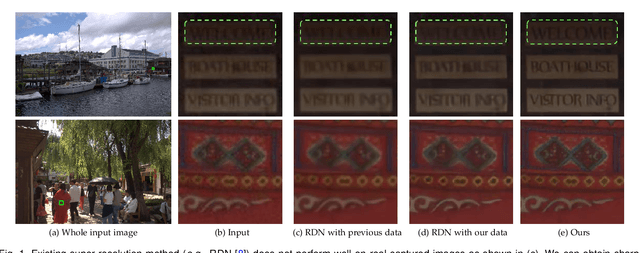
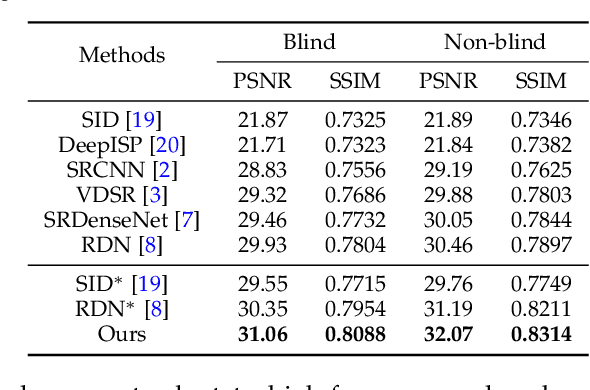
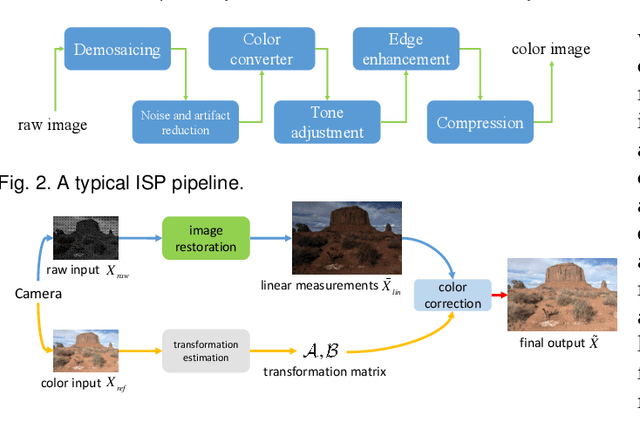
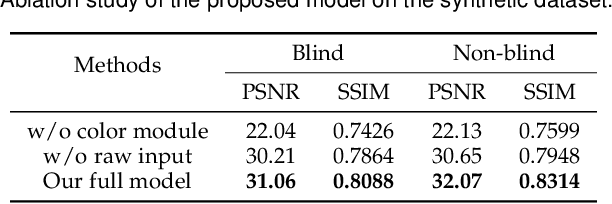
Super-resolution is a fundamental problem in computer vision which aims to overcome the spatial limitation of camera sensors. While significant progress has been made in single image super-resolution, most algorithms only perform well on synthetic data, which limits their applications in real scenarios. In this paper, we study the problem of real-scene single image super-resolution to bridge the gap between synthetic data and real captured images. We focus on two issues of existing super-resolution algorithms: lack of realistic training data and insufficient utilization of visual information obtained from cameras. To address the first issue, we propose a method to generate more realistic training data by mimicking the imaging process of digital cameras. For the second issue, we develop a two-branch convolutional neural network to exploit the radiance information originally-recorded in raw images. In addition, we propose a dense channel-attention block for better image restoration as well as a learning-based guided filter network for effective color correction. Our model is able to generalize to different cameras without deliberately training on images from specific camera types. Extensive experiments demonstrate that the proposed algorithm can recover fine details and clear structures, and achieve high-quality results for single image super-resolution in real scenes.
* A larger version with higher-resolution figures is available at: https://sites.google.com/view/xiangyuxu. arXiv admin note: text overlap with arXiv:1905.12156
Multi-Task Attention-Based Semi-Supervised Learning for Medical Image Segmentation
Jul 29, 2019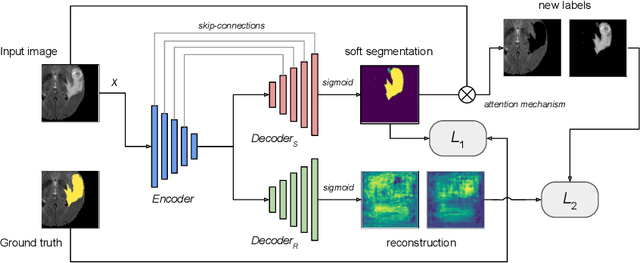
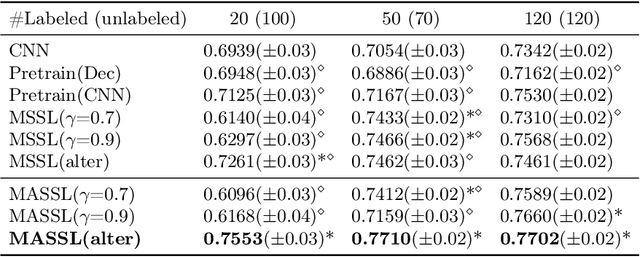
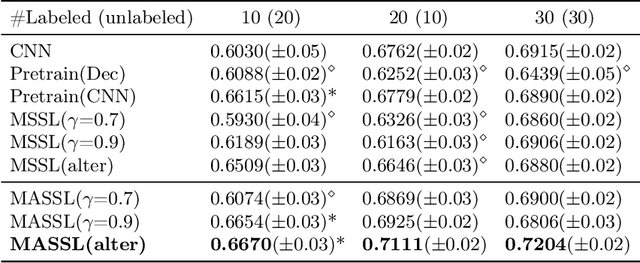
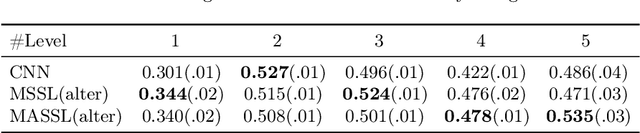
We propose a novel semi-supervised image segmentation method that simultaneously optimizes a supervised segmentation and an unsupervised reconstruction objectives. The reconstruction objective uses an attention mechanism that separates the reconstruction of image areas corresponding to different classes. The proposed approach was evaluated on two applications: brain tumor and white matter hyperintensities segmentation. Our method, trained on unlabeled and a small number of labeled images, outperformed supervised CNNs trained with the same number of images and CNNs pre-trained on unlabeled data. In ablation experiments, we observed that the proposed attention mechanism substantially improves segmentation performance. We explore two multi-task training strategies: joint training and alternating training. Alternating training requires fewer hyperparameters and achieves a better, more stable performance than joint training. Finally, we analyze the features learned by different methods and find that the attention mechanism helps to learn more discriminative features in the deeper layers of encoders.
SynthRef: Generation of Synthetic Referring Expressions for Object Segmentation
Jun 09, 2021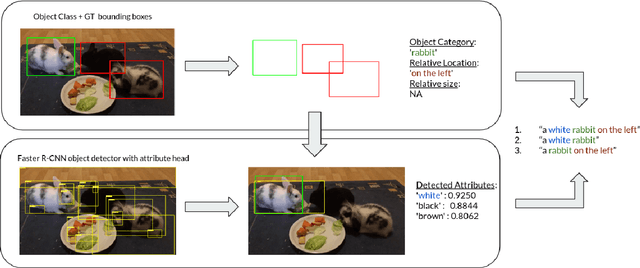
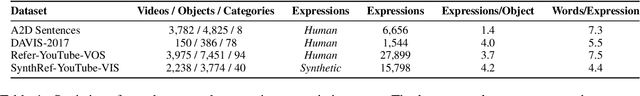
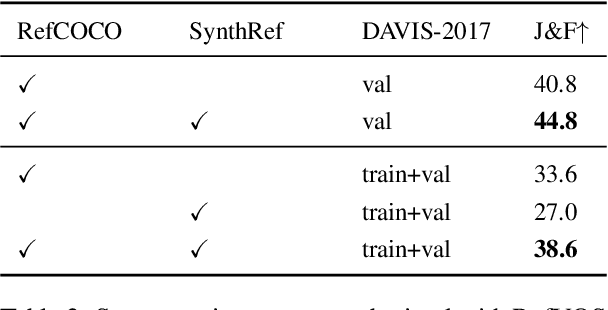



Recent advances in deep learning have brought significant progress in visual grounding tasks such as language-guided video object segmentation. However, collecting large datasets for these tasks is expensive in terms of annotation time, which represents a bottleneck. To this end, we propose a novel method, namely SynthRef, for generating synthetic referring expressions for target objects in an image (or video frame), and we also present and disseminate the first large-scale dataset with synthetic referring expressions for video object segmentation. Our experiments demonstrate that by training with our synthetic referring expressions one can improve the ability of a model to generalize across different datasets, without any additional annotation cost. Moreover, our formulation allows its application to any object detection or segmentation dataset.
Leaf Image-based Plant Disease Identification using Color and Texture Features
Feb 08, 2021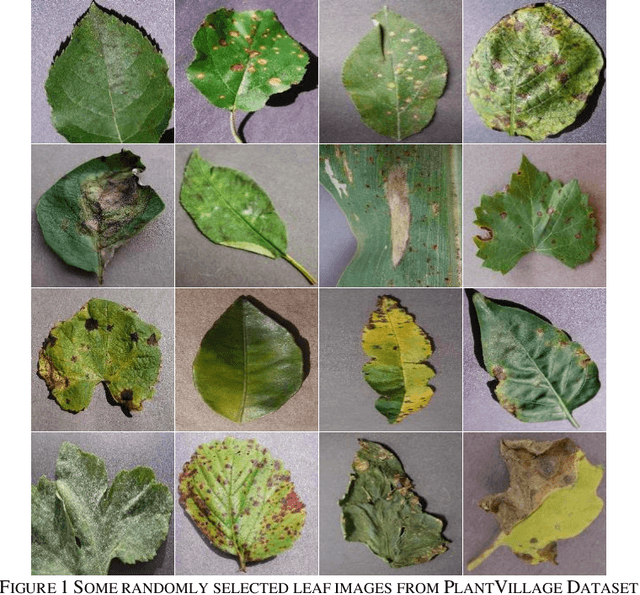
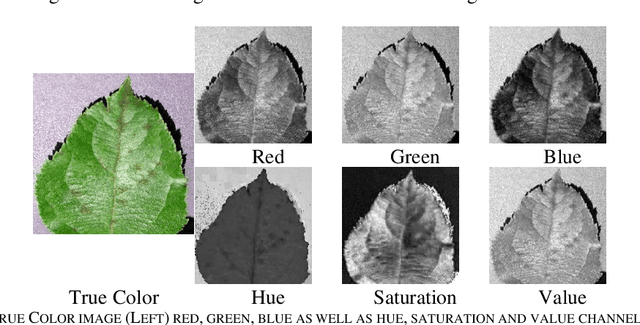
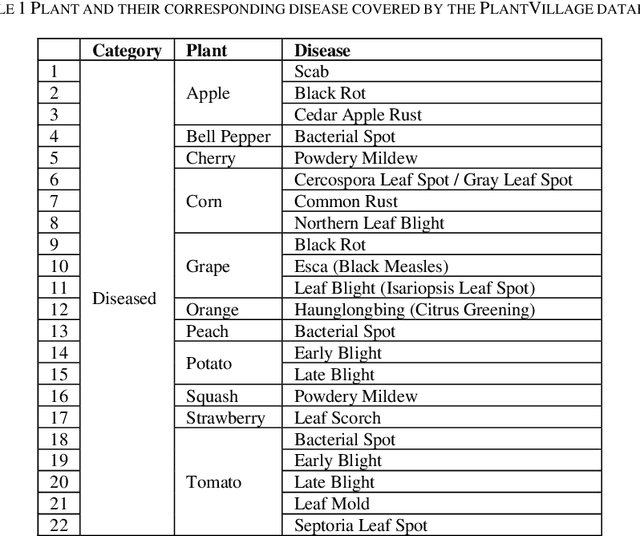

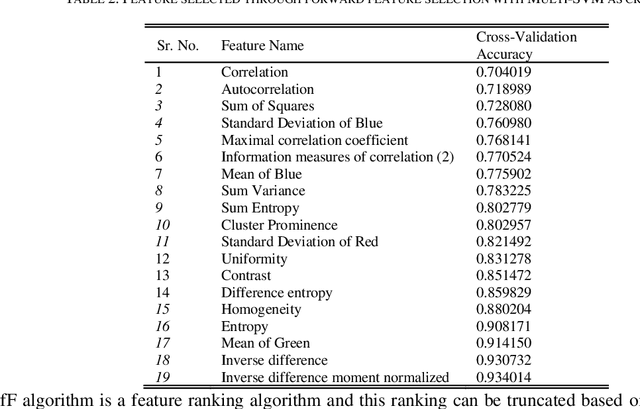
Identification of plant disease is usually done through visual inspection or during laboratory examination which causes delays resulting in yield loss by the time identification is complete. On the other hand, complex deep learning models perform the task with reasonable performance but due to their large size and high computational requirements, they are not suited to mobile and handheld devices. Our proposed approach contributes automated identification of plant diseases which follows a sequence of steps involving pre-processing, segmentation of diseased leaf area, calculation of features based on the Gray-Level Co-occurrence Matrix (GLCM), feature selection and classification. In this study, six color features and twenty-two texture features have been calculated. Support vector machines is used to perform one-vs-one classification of plant disease. The proposed model of disease identification provides an accuracy of 98.79% with a standard deviation of 0.57 on 10-fold cross-validation. The accuracy on a self-collected dataset is 82.47% for disease identification and 91.40% for healthy and diseased classification. The reported performance measures are better or comparable to the existing approaches and highest among the feature-based methods, presenting it as the most suitable method to automated leaf-based plant disease identification. This prototype system can be extended by adding more disease categories or targeting specific crop or disease categories.
Counterfactual Depth from a Single RGB Image
Sep 03, 2019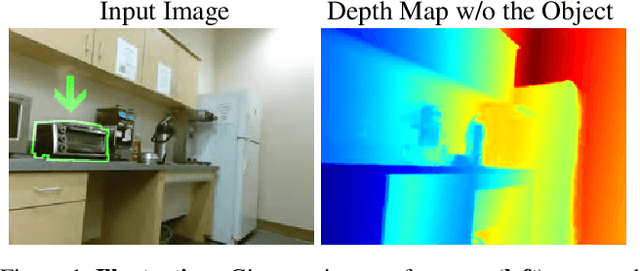
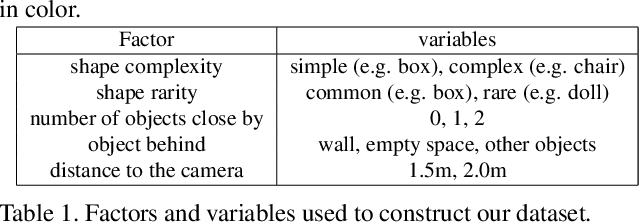

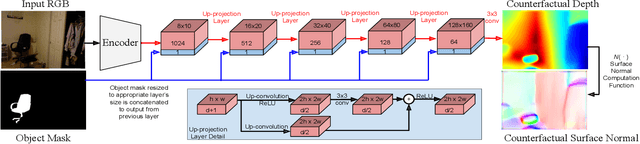
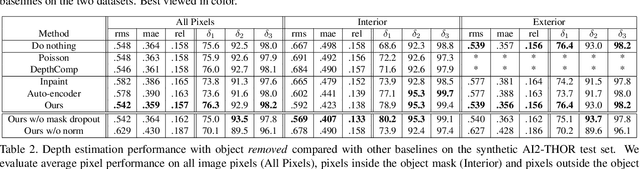
We describe a method that predicts, from a single RGB image, a depth map that describes the scene when a masked object is removed - we call this "counterfactual depth" that models hidden scene geometry together with the observations. Our method works for the same reason that scene completion works: the spatial structure of objects is simple. But we offer a much higher resolution representation of space than current scene completion methods, as we operate at pixel-level precision and do not rely on a voxel representation. Furthermore, we do not require RGBD inputs. Our method uses a standard encoder-decoder architecture, and with a decoder modified to accept an object mask. We describe a small evaluation dataset that we have collected, which allows inference about what factors affect reconstruction most strongly. Using this dataset, we show that our depth predictions for masked objects are better than other baselines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge