cancer detection
Cancer detection using Artificial Intelligence (AI) involves leveraging advanced machine learning algorithms and techniques to identify and diagnose cancer from various medical data sources. The goal is to enhance early detection, improve diagnostic accuracy, and potentially reduce the need for invasive procedures.
Papers and Code
Bluish Veil Detection and Lesion Classification using Custom Deep Learnable Layers with Explainable Artificial Intelligence (XAI)
Jul 10, 2025Melanoma, one of the deadliest types of skin cancer, accounts for thousands of fatalities globally. The bluish, blue-whitish, or blue-white veil (BWV) is a critical feature for diagnosing melanoma, yet research into detecting BWV in dermatological images is limited. This study utilizes a non-annotated skin lesion dataset, which is converted into an annotated dataset using a proposed imaging algorithm based on color threshold techniques on lesion patches and color palettes. A Deep Convolutional Neural Network (DCNN) is designed and trained separately on three individual and combined dermoscopic datasets, using custom layers instead of standard activation function layers. The model is developed to categorize skin lesions based on the presence of BWV. The proposed DCNN demonstrates superior performance compared to conventional BWV detection models across different datasets. The model achieves a testing accuracy of 85.71% on the augmented PH2 dataset, 95.00% on the augmented ISIC archive dataset, 95.05% on the combined augmented (PH2+ISIC archive) dataset, and 90.00% on the Derm7pt dataset. An explainable artificial intelligence (XAI) algorithm is subsequently applied to interpret the DCNN's decision-making process regarding BWV detection. The proposed approach, coupled with XAI, significantly improves the detection of BWV in skin lesions, outperforming existing models and providing a robust tool for early melanoma diagnosis.
* Accepted version. Published in Computers in Biology and Medicine, 14 June 2024. DOI: 10.1016/j.compbiomed.2024.108758
GuidedMorph: Two-Stage Deformable Registration for Breast MRI
May 19, 2025



Accurately registering breast MR images from different time points enables the alignment of anatomical structures and tracking of tumor progression, supporting more effective breast cancer detection, diagnosis, and treatment planning. However, the complexity of dense tissue and its highly non-rigid nature pose challenges for conventional registration methods, which primarily focus on aligning general structures while overlooking intricate internal details. To address this, we propose \textbf{GuidedMorph}, a novel two-stage registration framework designed to better align dense tissue. In addition to a single-scale network for global structure alignment, we introduce a framework that utilizes dense tissue information to track breast movement. The learned transformation fields are fused by introducing the Dual Spatial Transformer Network (DSTN), improving overall alignment accuracy. A novel warping method based on the Euclidean distance transform (EDT) is also proposed to accurately warp the registered dense tissue and breast masks, preserving fine structural details during deformation. The framework supports paradigms that require external segmentation models and with image data only. It also operates effectively with the VoxelMorph and TransMorph backbones, offering a versatile solution for breast registration. We validate our method on ISPY2 and internal dataset, demonstrating superior performance in dense tissue, overall breast alignment, and breast structural similarity index measure (SSIM), with notable improvements by over 13.01% in dense tissue Dice, 3.13% in breast Dice, and 1.21% in breast SSIM compared to the best learning-based baseline.
Vision-Language Model-Based Semantic-Guided Imaging Biomarker for Early Lung Cancer Detection
Apr 30, 2025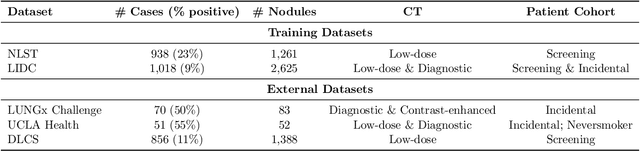
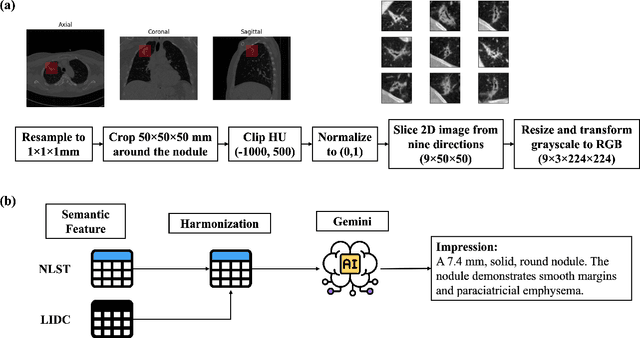
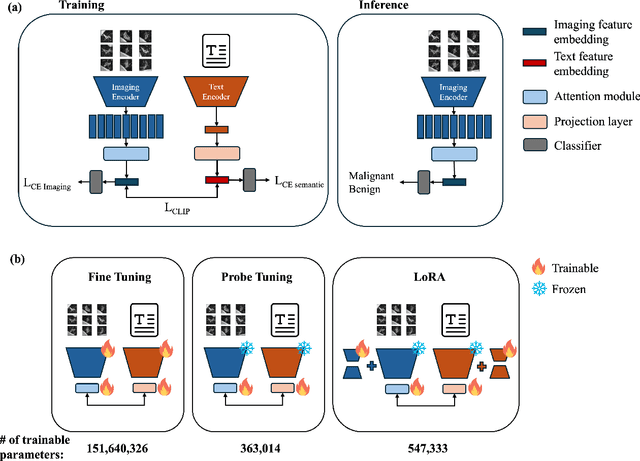
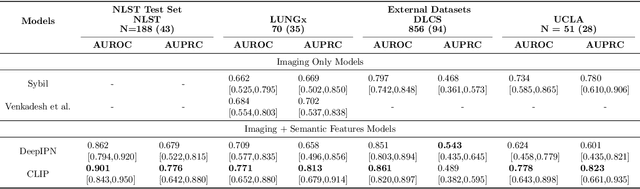
Objective: A number of machine learning models have utilized semantic features, deep features, or both to assess lung nodule malignancy. However, their reliance on manual annotation during inference, limited interpretability, and sensitivity to imaging variations hinder their application in real-world clinical settings. Thus, this research aims to integrate semantic features derived from radiologists' assessments of nodules, allowing the model to learn clinically relevant, robust, and explainable features for predicting lung cancer. Methods: We obtained 938 low-dose CT scans from the National Lung Screening Trial with 1,246 nodules and semantic features. The Lung Image Database Consortium dataset contains 1,018 CT scans, with 2,625 lesions annotated for nodule characteristics. Three external datasets were obtained from UCLA Health, the LUNGx Challenge, and the Duke Lung Cancer Screening. We finetuned a pretrained Contrastive Language-Image Pretraining model with a parameter-efficient fine-tuning approach to align imaging and semantic features and predict the one-year lung cancer diagnosis. Results: We evaluated the performance of the one-year diagnosis of lung cancer with AUROC and AUPRC and compared it to three state-of-the-art models. Our model demonstrated an AUROC of 0.90 and AUPRC of 0.78, outperforming baseline state-of-the-art models on external datasets. Using CLIP, we also obtained predictions on semantic features, such as nodule margin (AUROC: 0.81), nodule consistency (0.81), and pleural attachment (0.84), that can be used to explain model predictions. Conclusion: Our approach accurately classifies lung nodules as benign or malignant, providing explainable outputs, aiding clinicians in comprehending the underlying meaning of model predictions. This approach also prevents the model from learning shortcuts and generalizes across clinical settings.
Prostate Cancer Screening with Artificial Intelligence-Enhanced Micro-Ultrasound: A Comparative Study with Traditional Methods
May 27, 2025Background and objective: Micro-ultrasound (micro-US) is a novel imaging modality with diagnostic accuracy comparable to MRI for detecting clinically significant prostate cancer (csPCa). We investigated whether artificial intelligence (AI) interpretation of micro-US can outperform clinical screening methods using PSA and digital rectal examination (DRE). Methods: We retrospectively studied 145 men who underwent micro-US guided biopsy (79 with csPCa, 66 without). A self-supervised convolutional autoencoder was used to extract deep image features from 2D micro-US slices. Random forest classifiers were trained using five-fold cross-validation to predict csPCa at the slice level. Patients were classified as csPCa-positive if 88 or more consecutive slices were predicted positive. Model performance was compared with a classifier using PSA, DRE, prostate volume, and age. Key findings and limitations: The AI-based micro-US model and clinical screening model achieved AUROCs of 0.871 and 0.753, respectively. At a fixed threshold, the micro-US model achieved 92.5% sensitivity and 68.1% specificity, while the clinical model showed 96.2% sensitivity but only 27.3% specificity. Limitations include a retrospective single-center design and lack of external validation. Conclusions and clinical implications: AI-interpreted micro-US improves specificity while maintaining high sensitivity for csPCa detection. This method may reduce unnecessary biopsies and serve as a low-cost alternative to PSA-based screening. Patient summary: We developed an AI system to analyze prostate micro-ultrasound images. It outperformed PSA and DRE in detecting aggressive cancer and may help avoid unnecessary biopsies.
Devising a solution to the problems of Cancer awareness in Telangana
Jun 26, 2025



According to the data, the percent of women who underwent screening for cervical cancer, breast and oral cancer in Telangana in the year 2020 was 3.3 percent, 0.3 percent and 2.3 percent respectively. Although early detection is the only way to reduce morbidity and mortality, people have very low awareness about cervical and breast cancer signs and symptoms and screening practices. We developed an ML classification model to predict if a person is susceptible to breast or cervical cancer based on demographic factors. We devised a system to provide suggestions for the nearest hospital or Cancer treatment centres based on the users location or address. In addition to this, we can integrate the health card to maintain medical records of all individuals and conduct awareness drives and campaigns. For ML classification models, we used decision tree classification and support vector classification algorithms for cervical cancer susceptibility and breast cancer susceptibility respectively. Thus, by devising this solution we come one step closer to our goal which is spreading cancer awareness, thereby, decreasing the cancer mortality and increasing cancer literacy among the people of Telangana.
White Light Specular Reflection Data Augmentation for Deep Learning Polyp Detection
May 08, 2025Colorectal cancer is one of the deadliest cancers today, but it can be prevented through early detection of malignant polyps in the colon, primarily via colonoscopies. While this method has saved many lives, human error remains a significant challenge, as missing a polyp could have fatal consequences for the patient. Deep learning (DL) polyp detectors offer a promising solution. However, existing DL polyp detectors often mistake white light reflections from the endoscope for polyps, which can lead to false positives.To address this challenge, in this paper, we propose a novel data augmentation approach that artificially adds more white light reflections to create harder training scenarios. Specifically, we first generate a bank of artificial lights using the training dataset. Then we find the regions of the training images that we should not add these artificial lights on. Finally, we propose a sliding window method to add the artificial light to the areas that fit of the training images, resulting in augmented images. By providing the model with more opportunities to make mistakes, we hypothesize that it will also have more chances to learn from those mistakes, ultimately improving its performance in polyp detection. Experimental results demonstrate the effectiveness of our new data augmentation method.
Lightweight Relational Embedding in Task-Interpolated Few-Shot Networks for Enhanced Gastrointestinal Disease Classification
May 30, 2025Traditional diagnostic methods like colonoscopy are invasive yet critical tools necessary for accurately diagnosing colorectal cancer (CRC). Detection of CRC at early stages is crucial for increasing patient survival rates. However, colonoscopy is dependent on obtaining adequate and high-quality endoscopic images. Prolonged invasive procedures are inherently risky for patients, while suboptimal or insufficient images hamper diagnostic accuracy. These images, typically derived from video frames, often exhibit similar patterns, posing challenges in discrimination. To overcome these challenges, we propose a novel Deep Learning network built on a Few-Shot Learning architecture, which includes a tailored feature extractor, task interpolation, relational embedding, and a bi-level routing attention mechanism. The Few-Shot Learning paradigm enables our model to rapidly adapt to unseen fine-grained endoscopic image patterns, and the task interpolation augments the insufficient images artificially from varied instrument viewpoints. Our relational embedding approach discerns critical intra-image features and captures inter-image transitions between consecutive endoscopic frames, overcoming the limitations of Convolutional Neural Networks (CNNs). The integration of a light-weight attention mechanism ensures a concentrated analysis of pertinent image regions. By training on diverse datasets, the model's generalizability and robustness are notably improved for handling endoscopic images. Evaluated on Kvasir dataset, our model demonstrated superior performance, achieving an accuracy of 90.1\%, precision of 0.845, recall of 0.942, and an F1 score of 0.891. This surpasses current state-of-the-art methods, presenting a promising solution to the challenges of invasive colonoscopy by optimizing CRC detection through advanced image analysis.
* 6 pages, 15 figures
Mammo-Mamba: A Hybrid State-Space and Transformer Architecture with Sequential Mixture of Experts for Multi-View Mammography
Jul 23, 2025Breast cancer (BC) remains one of the leading causes of cancer-related mortality among women, despite recent advances in Computer-Aided Diagnosis (CAD) systems. Accurate and efficient interpretation of multi-view mammograms is essential for early detection, driving a surge of interest in Artificial Intelligence (AI)-powered CAD models. While state-of-the-art multi-view mammogram classification models are largely based on Transformer architectures, their computational complexity scales quadratically with the number of image patches, highlighting the need for more efficient alternatives. To address this challenge, we propose Mammo-Mamba, a novel framework that integrates Selective State-Space Models (SSMs), transformer-based attention, and expert-driven feature refinement into a unified architecture. Mammo-Mamba extends the MambaVision backbone by introducing the Sequential Mixture of Experts (SeqMoE) mechanism through its customized SecMamba block. The SecMamba is a modified MambaVision block that enhances representation learning in high-resolution mammographic images by enabling content-adaptive feature refinement. These blocks are integrated into the deeper stages of MambaVision, allowing the model to progressively adjust feature emphasis through dynamic expert gating, effectively mitigating the limitations of traditional Transformer models. Evaluated on the CBIS-DDSM benchmark dataset, Mammo-Mamba achieves superior classification performance across all key metrics while maintaining computational efficiency.
Advanced cervical cancer classification: enhancing pap smear images with hybrid PMD Filter-CLAHE
Jun 18, 2025



Cervical cancer remains a significant health problem, especially in developing countries. Early detection is critical for effective treatment. Convolutional neural networks (CNN) have shown promise in automated cervical cancer screening, but their performance depends on Pap smear image quality. This study investigates the impact of various image preprocessing techniques on CNN performance for cervical cancer classification using the SIPaKMeD dataset. Three preprocessing techniques were evaluated: perona-malik diffusion (PMD) filter for noise reduction, contrast-limited adaptive histogram equalization (CLAHE) for image contrast enhancement, and the proposed hybrid PMD filter-CLAHE approach. The enhanced image datasets were evaluated on pretrained models, such as ResNet-34, ResNet-50, SqueezeNet-1.0, MobileNet-V2, EfficientNet-B0, EfficientNet-B1, DenseNet-121, and DenseNet-201. The results show that hybrid preprocessing PMD filter-CLAHE can improve the Pap smear image quality and CNN architecture performance compared to the original images. The maximum metric improvements are 13.62% for accuracy, 10.04% for precision, 13.08% for recall, and 14.34% for F1-score. The proposed hybrid PMD filter-CLAHE technique offers a new perspective in improving cervical cancer classification performance using CNN architectures.
MAMBO: High-Resolution Generative Approach for Mammography Images
Jun 10, 2025Mammography is the gold standard for the detection and diagnosis of breast cancer. This procedure can be significantly enhanced with Artificial Intelligence (AI)-based software, which assists radiologists in identifying abnormalities. However, training AI systems requires large and diverse datasets, which are often difficult to obtain due to privacy and ethical constraints. To address this issue, the paper introduces MAMmography ensemBle mOdel (MAMBO), a novel patch-based diffusion approach designed to generate full-resolution mammograms. Diffusion models have shown breakthrough results in realistic image generation, yet few studies have focused on mammograms, and none have successfully generated high-resolution outputs required to capture fine-grained features of small lesions. To achieve this, MAMBO integrates separate diffusion models to capture both local and global (image-level) contexts. The contextual information is then fed into the final patch-based model, significantly aiding the noise removal process. This thoughtful design enables MAMBO to generate highly realistic mammograms of up to 3840x3840 pixels. Importantly, this approach can be used to enhance the training of classification models and extended to anomaly detection. Experiments, both numerical and radiologist validation, assess MAMBO's capabilities in image generation, super-resolution, and anomaly detection, highlighting its potential to enhance mammography analysis for more accurate diagnoses and earlier lesion detection.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge