William Hsu
ViTs for Action Classification in Videos: An Approach to Risky Tackle Detection in American Football Practice Videos
Apr 01, 2026Abstract:Early identification of hazardous actions in contact sports enables timely intervention and improves player safety. We present a method for detecting risky tackles in American football practice videos and introduce a substantially expanded dataset for this task. Our work contains 733 single-athlete-dummy tackle clips, each temporally localized around first point contact and labeled with a strike zone component of the standardized Assessment for Tackling Technique (SATT-3), extending prior work that reported 178 annotated videos. Using a Vision transformer-based model with imbalance-aware training, we obtain risky recall of 0.67 and Risky F1 of 0.59 under crossvalidation. Relative to the previous baseline in a smaller subset (risky recall of 0.58; Risky F1 0.56 ), our approach improves risky recall by more than 8% points on a much larger dataset. These results indicate that the vision transformer-based video analysis, coupled with careful handling of class imbalance, can reliably detect rare but safety-critical tackling patterns, offering a practical pathway toward coach-centered injury prevention tools.
GRAZE: Grounded Refinement and Motion-Aware Zero-Shot Event Localization
Apr 01, 2026Abstract:American football practice generates video at scale, yet the interaction of interest occupies only a brief window of each long, untrimmed clip. Reliable biomechanical analysis, therefore, depends on spatiotemporal localization that identifies both the interacting entities and the onset of contact. We study First Point of Contact (FPOC), defined as the first frame in which a player physically touches a tackle dummy, in unconstrained practice footage with camera motion, clutter, multiple similarly equipped athletes, and rapid pose changes around impact. We present GRAZE, a training-free pipeline for FPOC localization that requires no labeled tackle-contact examples. GRAZE uses Grounding DINO to discover candidate player-dummy interactions, refines them with motion-aware temporal reasoning, and uses SAM2 as an explicit pixel-level verifier of contact rather than relying on detection confidence alone. This separation between candidate discovery and contact confirmation makes the approach robust to cluttered scenes and unstable grounding near impact. On 738 tackle-practice videos, GRAZE produces valid outputs for 97.4% of clips and localizes FPOC within $\pm$ 10 frames on 77.5% of all clips and within $\pm$ 20 frames on 82.7% of all clips. These results show that frame-accurate contact onset localization in real-world practice footage is feasible without task-specific training.
GLIDE-Reg: Global-to-Local Deformable Registration Using Co-Optimized Foundation and Handcrafted Features
Mar 03, 2026Abstract:Deformable registration is crucial in medical imaging. Several existing applications include lesion tracking, probabilistic atlas generation, and treatment response evaluation. However, current methods often lack robustness and generalizability across two key factors: spatial resolution and differences in anatomical coverage. We jointly optimize a registration field and a learnable dimensionality reduction module so that compressed VFM embeddings remain registration-relevant, and fuse these global semantic cues with MIND local descriptors. GLIDE-Reg achieves average dice similarity coefficients (DSC) across 6 anatomical structures of 0.859, 0.862, and 0.901 in two public cohorts (Lung250M and NLST) and one institution cohort (UCLA5DCT), and outperforms the state-of-the-art DEEDS (0.834, 0.858, 0.900) with relative improvements of 3.0%, 0.5%, and 0.1%. For target registration errors, GLIDE-Reg achieves 1.58 mm on Lung250M landmarks (compared to 1.25 mm on corrField and 1.91 mm on DEEDS) and 1.11 mm on NLST nodule centers (compared to 1.11 mm on DEEDS). The substantiated performance on the nodule centers also demonstrates its robustness across challenging downstream tasks, such as nodule tracking, which is an essential prior step for early-stage lung cancer diagnosis.
Semi-Supervised Domain Adaptation with Latent Diffusion for Pathology Image Classification
Jan 23, 2026Abstract:Deep learning models in computational pathology often fail to generalize across cohorts and institutions due to domain shift. Existing approaches either fail to leverage unlabeled data from the target domain or rely on image-to-image translation, which can distort tissue structures and compromise model accuracy. In this work, we propose a semi-supervised domain adaptation (SSDA) framework that utilizes a latent diffusion model trained on unlabeled data from both the source and target domains to generate morphology-preserving and target-aware synthetic images. By conditioning the diffusion model on foundation model features, cohort identity, and tissue preparation method, we preserve tissue structure in the source domain while introducing target-domain appearance characteristics. The target-aware synthetic images, combined with real, labeled images from the source cohort, are subsequently used to train a downstream classifier, which is then tested on the target cohort. The effectiveness of the proposed SSDA framework is demonstrated on the task of lung adenocarcinoma prognostication. The proposed augmentation yielded substantially better performance on the held-out test set from the target cohort, without degrading source-cohort performance. The approach improved the weighted F1 score on the target-cohort held-out test set from 0.611 to 0.706 and the macro F1 score from 0.641 to 0.716. Our results demonstrate that target-aware diffusion-based synthetic data augmentation provides a promising and effective approach for improving domain generalization in computational pathology.
Vision-Language Model-Based Semantic-Guided Imaging Biomarker for Early Lung Cancer Detection
Apr 30, 2025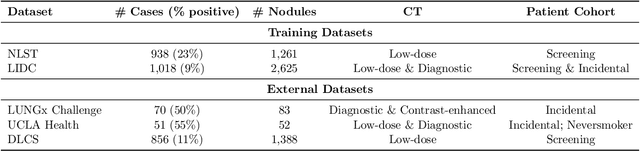
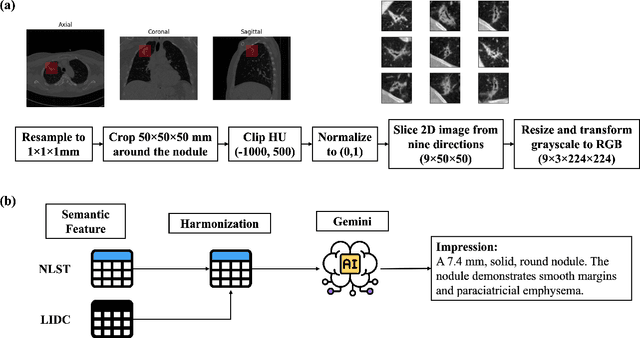
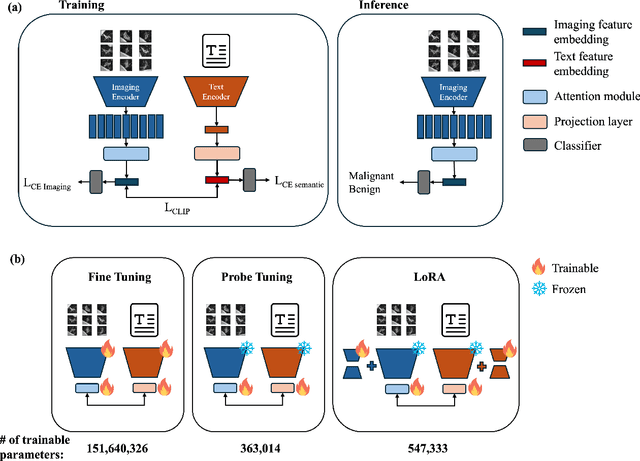
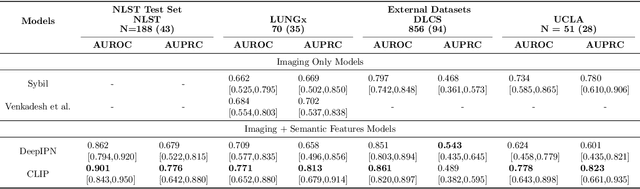
Abstract:Objective: A number of machine learning models have utilized semantic features, deep features, or both to assess lung nodule malignancy. However, their reliance on manual annotation during inference, limited interpretability, and sensitivity to imaging variations hinder their application in real-world clinical settings. Thus, this research aims to integrate semantic features derived from radiologists' assessments of nodules, allowing the model to learn clinically relevant, robust, and explainable features for predicting lung cancer. Methods: We obtained 938 low-dose CT scans from the National Lung Screening Trial with 1,246 nodules and semantic features. The Lung Image Database Consortium dataset contains 1,018 CT scans, with 2,625 lesions annotated for nodule characteristics. Three external datasets were obtained from UCLA Health, the LUNGx Challenge, and the Duke Lung Cancer Screening. We finetuned a pretrained Contrastive Language-Image Pretraining model with a parameter-efficient fine-tuning approach to align imaging and semantic features and predict the one-year lung cancer diagnosis. Results: We evaluated the performance of the one-year diagnosis of lung cancer with AUROC and AUPRC and compared it to three state-of-the-art models. Our model demonstrated an AUROC of 0.90 and AUPRC of 0.78, outperforming baseline state-of-the-art models on external datasets. Using CLIP, we also obtained predictions on semantic features, such as nodule margin (AUROC: 0.81), nodule consistency (0.81), and pleural attachment (0.84), that can be used to explain model predictions. Conclusion: Our approach accurately classifies lung nodules as benign or malignant, providing explainable outputs, aiding clinicians in comprehending the underlying meaning of model predictions. This approach also prevents the model from learning shortcuts and generalizes across clinical settings.
Advancing Precision Oncology Through Modeling of Longitudinal and Multimodal Data
Feb 11, 2025



Abstract:Cancer evolves continuously over time through a complex interplay of genetic, epigenetic, microenvironmental, and phenotypic changes. This dynamic behavior drives uncontrolled cell growth, metastasis, immune evasion, and therapy resistance, posing challenges for effective monitoring and treatment. However, today's data-driven research in oncology has primarily focused on cross-sectional analysis using data from a single modality, limiting the ability to fully characterize and interpret the disease's dynamic heterogeneity. Advances in multiscale data collection and computational methods now enable the discovery of longitudinal multimodal biomarkers for precision oncology. Longitudinal data reveal patterns of disease progression and treatment response that are not evident from single-timepoint data, enabling timely abnormality detection and dynamic treatment adaptation. Multimodal data integration offers complementary information from diverse sources for more precise risk assessment and targeting of cancer therapy. In this review, we survey methods of longitudinal and multimodal modeling, highlighting their synergy in providing multifaceted insights for personalized care tailored to the unique characteristics of a patient's cancer. We summarize the current challenges and future directions of longitudinal multimodal analysis in advancing precision oncology.
Unifying Interpretability and Explainability for Alzheimer's Disease Progression Prediction
Jun 11, 2024Abstract:Reinforcement learning (RL) has recently shown promise in predicting Alzheimer's disease (AD) progression due to its unique ability to model domain knowledge. However, it is not clear which RL algorithms are well-suited for this task. Furthermore, these methods are not inherently explainable, limiting their applicability in real-world clinical scenarios. Our work addresses these two important questions. Using a causal, interpretable model of AD, we first compare the performance of four contemporary RL algorithms in predicting brain cognition over 10 years using only baseline (year 0) data. We then apply SHAP (SHapley Additive exPlanations) to explain the decisions made by each algorithm in the model. Our approach combines interpretability with explainability to provide insights into the key factors influencing AD progression, offering both global and individual, patient-level analysis. Our findings show that only one of the RL methods is able to satisfactorily model disease progression, but the post-hoc explanations indicate that all methods fail to properly capture the importance of amyloid accumulation, one of the pathological hallmarks of Alzheimer's disease. Our work aims to merge predictive accuracy with transparency, assisting clinicians and researchers in enhancing disease progression modeling for informed healthcare decisions. Code is available at https://github.com/rfali/xrlad.
Combining Graph Neural Network and Mamba to Capture Local and Global Tissue Spatial Relationships in Whole Slide Images
Jun 05, 2024Abstract:In computational pathology, extracting spatial features from gigapixel whole slide images (WSIs) is a fundamental task, but due to their large size, WSIs are typically segmented into smaller tiles. A critical aspect of this analysis is aggregating information from these tiles to make predictions at the WSI level. We introduce a model that combines a message-passing graph neural network (GNN) with a state space model (Mamba) to capture both local and global spatial relationships among the tiles in WSIs. The model's effectiveness was demonstrated in predicting progression-free survival among patients with early-stage lung adenocarcinomas (LUAD). We compared the model with other state-of-the-art methods for tile-level information aggregation in WSIs, including tile-level information summary statistics-based aggregation, multiple instance learning (MIL)-based aggregation, GNN-based aggregation, and GNN-transformer-based aggregation. Additional experiments showed the impact of different types of node features and different tile sampling strategies on the model performance. This work can be easily extended to any WSI-based analysis. Code: https://github.com/rina-ding/gat-mamba.
Spatial Matching of 2D Mammography Images and Specimen Radiographs: Towards Improved Characterization of Suspicious Microcalcifications
May 21, 2024Abstract:Accurate characterization of suspicious microcalcifications is critical to determine whether these calcifications are associated with invasive disease. Our overarching objective is to enable the joint characterization of microcalcifications and surrounding breast tissue using mammography images and digital histopathology images. Towards this goal, we investigate a template matching-based approach that utilizes microcalcifications as landmarks to match radiographs taken of biopsy core specimens to groups of calcifications that are visible on mammography. Our approach achieved a high negative predictive value (0.98) but modest precision (0.66) and recall (0.58) in identifying the mammographic region where microcalcifications were taken during a core needle biopsy.
Multimodal Machine Learning in Image-Based and Clinical Biomedicine: Survey and Prospects
Nov 20, 2023Abstract:Machine learning (ML) applications in medical artificial intelligence (AI) systems have shifted from traditional and statistical methods to increasing application of deep learning models. This survey navigates the current landscape of multimodal ML, focusing on its profound impact on medical image analysis and clinical decision support systems. Emphasizing challenges and innovations in addressing multimodal representation, fusion, translation, alignment, and co-learning, the paper explores the transformative potential of multimodal models for clinical predictions. It also questions practical implementation of such models, bringing attention to the dynamics between decision support systems and healthcare providers. Despite advancements, challenges such as data biases and the scarcity of "big data" in many biomedical domains persist. We conclude with a discussion on effective innovation and collaborative efforts to further the miss
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge