Luoting Zhuang
GLIDE-Reg: Global-to-Local Deformable Registration Using Co-Optimized Foundation and Handcrafted Features
Mar 03, 2026Abstract:Deformable registration is crucial in medical imaging. Several existing applications include lesion tracking, probabilistic atlas generation, and treatment response evaluation. However, current methods often lack robustness and generalizability across two key factors: spatial resolution and differences in anatomical coverage. We jointly optimize a registration field and a learnable dimensionality reduction module so that compressed VFM embeddings remain registration-relevant, and fuse these global semantic cues with MIND local descriptors. GLIDE-Reg achieves average dice similarity coefficients (DSC) across 6 anatomical structures of 0.859, 0.862, and 0.901 in two public cohorts (Lung250M and NLST) and one institution cohort (UCLA5DCT), and outperforms the state-of-the-art DEEDS (0.834, 0.858, 0.900) with relative improvements of 3.0%, 0.5%, and 0.1%. For target registration errors, GLIDE-Reg achieves 1.58 mm on Lung250M landmarks (compared to 1.25 mm on corrField and 1.91 mm on DEEDS) and 1.11 mm on NLST nodule centers (compared to 1.11 mm on DEEDS). The substantiated performance on the nodule centers also demonstrates its robustness across challenging downstream tasks, such as nodule tracking, which is an essential prior step for early-stage lung cancer diagnosis.
Semi-Supervised Domain Adaptation with Latent Diffusion for Pathology Image Classification
Jan 23, 2026Abstract:Deep learning models in computational pathology often fail to generalize across cohorts and institutions due to domain shift. Existing approaches either fail to leverage unlabeled data from the target domain or rely on image-to-image translation, which can distort tissue structures and compromise model accuracy. In this work, we propose a semi-supervised domain adaptation (SSDA) framework that utilizes a latent diffusion model trained on unlabeled data from both the source and target domains to generate morphology-preserving and target-aware synthetic images. By conditioning the diffusion model on foundation model features, cohort identity, and tissue preparation method, we preserve tissue structure in the source domain while introducing target-domain appearance characteristics. The target-aware synthetic images, combined with real, labeled images from the source cohort, are subsequently used to train a downstream classifier, which is then tested on the target cohort. The effectiveness of the proposed SSDA framework is demonstrated on the task of lung adenocarcinoma prognostication. The proposed augmentation yielded substantially better performance on the held-out test set from the target cohort, without degrading source-cohort performance. The approach improved the weighted F1 score on the target-cohort held-out test set from 0.611 to 0.706 and the macro F1 score from 0.641 to 0.716. Our results demonstrate that target-aware diffusion-based synthetic data augmentation provides a promising and effective approach for improving domain generalization in computational pathology.
Vision-Language Model-Based Semantic-Guided Imaging Biomarker for Early Lung Cancer Detection
Apr 30, 2025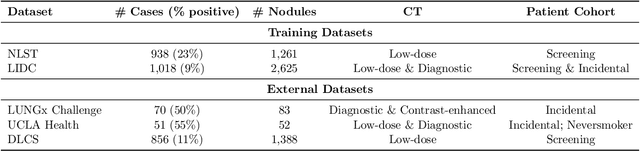
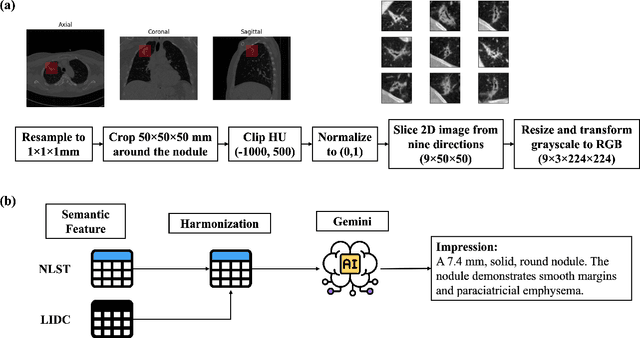
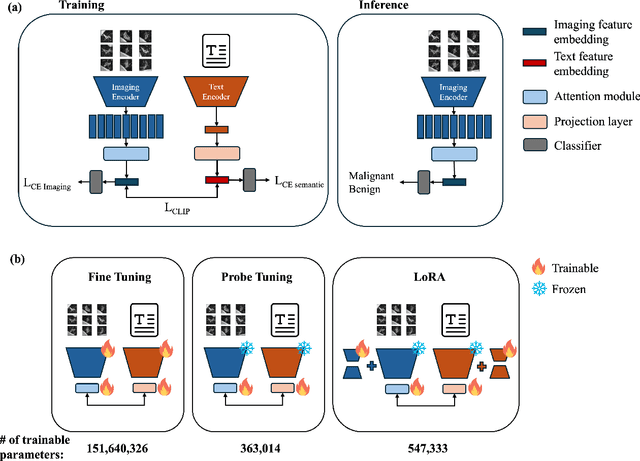
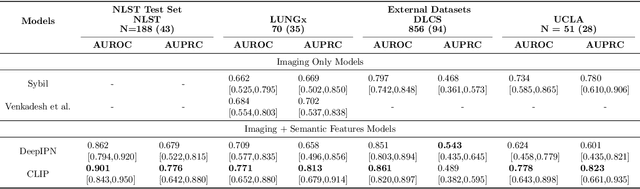
Abstract:Objective: A number of machine learning models have utilized semantic features, deep features, or both to assess lung nodule malignancy. However, their reliance on manual annotation during inference, limited interpretability, and sensitivity to imaging variations hinder their application in real-world clinical settings. Thus, this research aims to integrate semantic features derived from radiologists' assessments of nodules, allowing the model to learn clinically relevant, robust, and explainable features for predicting lung cancer. Methods: We obtained 938 low-dose CT scans from the National Lung Screening Trial with 1,246 nodules and semantic features. The Lung Image Database Consortium dataset contains 1,018 CT scans, with 2,625 lesions annotated for nodule characteristics. Three external datasets were obtained from UCLA Health, the LUNGx Challenge, and the Duke Lung Cancer Screening. We finetuned a pretrained Contrastive Language-Image Pretraining model with a parameter-efficient fine-tuning approach to align imaging and semantic features and predict the one-year lung cancer diagnosis. Results: We evaluated the performance of the one-year diagnosis of lung cancer with AUROC and AUPRC and compared it to three state-of-the-art models. Our model demonstrated an AUROC of 0.90 and AUPRC of 0.78, outperforming baseline state-of-the-art models on external datasets. Using CLIP, we also obtained predictions on semantic features, such as nodule margin (AUROC: 0.81), nodule consistency (0.81), and pleural attachment (0.84), that can be used to explain model predictions. Conclusion: Our approach accurately classifies lung nodules as benign or malignant, providing explainable outputs, aiding clinicians in comprehending the underlying meaning of model predictions. This approach also prevents the model from learning shortcuts and generalizes across clinical settings.
Advancing Precision Oncology Through Modeling of Longitudinal and Multimodal Data
Feb 11, 2025



Abstract:Cancer evolves continuously over time through a complex interplay of genetic, epigenetic, microenvironmental, and phenotypic changes. This dynamic behavior drives uncontrolled cell growth, metastasis, immune evasion, and therapy resistance, posing challenges for effective monitoring and treatment. However, today's data-driven research in oncology has primarily focused on cross-sectional analysis using data from a single modality, limiting the ability to fully characterize and interpret the disease's dynamic heterogeneity. Advances in multiscale data collection and computational methods now enable the discovery of longitudinal multimodal biomarkers for precision oncology. Longitudinal data reveal patterns of disease progression and treatment response that are not evident from single-timepoint data, enabling timely abnormality detection and dynamic treatment adaptation. Multimodal data integration offers complementary information from diverse sources for more precise risk assessment and targeting of cancer therapy. In this review, we survey methods of longitudinal and multimodal modeling, highlighting their synergy in providing multifaceted insights for personalized care tailored to the unique characteristics of a patient's cancer. We summarize the current challenges and future directions of longitudinal multimodal analysis in advancing precision oncology.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge