Zina Ibrahim
Fine-tuning -- a Transfer Learning approach
Nov 06, 2024Abstract:Secondary research use of Electronic Health Records (EHRs) is often hampered by the abundance of missing data in this valuable resource. Missingness in EHRs occurs naturally as a result of the data recording practices during routine clinical care, but handling it is crucial to the precision of medical analysis and the decision-making that follows. The literature contains a variety of imputation methodologies based on deep neural networks. Those aim to overcome the dynamic, heterogeneous and multivariate missingness patterns of EHRs, which cannot be handled by classical and statistical imputation methods. However, all existing deep imputation methods rely on end-to-end pipelines that incorporate both imputation and downstream analyses, e.g. classification. This coupling makes it difficult to assess the quality of imputation and takes away the flexibility of re-using the imputer for a different task. Furthermore, most end-to-end deep architectures tend to use complex networks to perform the downstream task, in addition to the already sophisticated deep imputation network. We, therefore ask if the high performance reported in the literature is due to the imputer or the classifier and further ask if an optimised state-of-the-art imputer is used, a simpler classifier can achieve comparable performance. This paper explores the development of a modular, deep learning-based imputation and classification pipeline, specifically built to leverage the capabilities of state-of-the-art imputation models for downstream classification tasks. Such a modular approach enables a) objective assessment of the quality of the imputer and classifier independently, and b) enables the exploration of the performance of simpler classification architectures using an optimised imputer.
How Deep is your Guess? A Fresh Perspective on Deep Learning for Medical Time-Series Imputation
Jul 11, 2024Abstract:We introduce a novel classification framework for time-series imputation using deep learning, with a particular focus on clinical data. By identifying conceptual gaps in the literature and existing reviews, we devise a taxonomy grounded on the inductive bias of neural imputation frameworks, resulting in a classification of existing deep imputation strategies based on their suitability for specific imputation scenarios and data-specific properties. Our review further examines the existing methodologies employed to benchmark deep imputation models, evaluating their effectiveness in capturing the missingness scenarios found in clinical data and emphasising the importance of reconciling mathematical abstraction with clinical insights. Our classification aims to serve as a guide for researchers to facilitate the selection of appropriate deep learning imputation techniques tailored to their specific clinical data. Our novel perspective also highlights the significance of bridging the gap between computational methodologies and medical insights to achieve clinically sound imputation models.
TSI-Bench: Benchmarking Time Series Imputation
Jun 18, 2024Abstract:Effective imputation is a crucial preprocessing step for time series analysis. Despite the development of numerous deep learning algorithms for time series imputation, the community lacks standardized and comprehensive benchmark platforms to effectively evaluate imputation performance across different settings. Moreover, although many deep learning forecasting algorithms have demonstrated excellent performance, whether their modeling achievements can be transferred to time series imputation tasks remains unexplored. To bridge these gaps, we develop TSI-Bench, the first (to our knowledge) comprehensive benchmark suite for time series imputation utilizing deep learning techniques. The TSI-Bench pipeline standardizes experimental settings to enable fair evaluation of imputation algorithms and identification of meaningful insights into the influence of domain-appropriate missingness ratios and patterns on model performance. Furthermore, TSI-Bench innovatively provides a systematic paradigm to tailor time series forecasting algorithms for imputation purposes. Our extensive study across 34,804 experiments, 28 algorithms, and 8 datasets with diverse missingness scenarios demonstrates TSI-Bench's effectiveness in diverse downstream tasks and potential to unlock future directions in time series imputation research and analysis. The source code and experiment logs are available at https://github.com/WenjieDu/AwesomeImputation.
Unveiling the Secrets: How Masking Strategies Shape Time Series Imputation
May 26, 2024Abstract:In this study, we explore the impact of different masking strategies on time series imputation models. We evaluate the effects of pre-masking versus in-mini-batch masking, normalization timing, and the choice between augmenting and overlaying artificial missingness. Using three diverse datasets, we benchmark eleven imputation models with different missing rates. Our results demonstrate that masking strategies significantly influence imputation accuracy, revealing that more sophisticated and data-driven masking designs are essential for robust model evaluation. We advocate for refined experimental designs and comprehensive disclosureto better simulate real-world patterns, enhancing the practical applicability of imputation models.
Uncertainty-Aware Deep Attention Recurrent Neural Network for Heterogeneous Time Series Imputation
Jan 04, 2024Abstract:Missingness is ubiquitous in multivariate time series and poses an obstacle to reliable downstream analysis. Although recurrent network imputation achieved the SOTA, existing models do not scale to deep architectures that can potentially alleviate issues arising in complex data. Moreover, imputation carries the risk of biased estimations of the ground truth. Yet, confidence in the imputed values is always unmeasured or computed post hoc from model output. We propose DEep Attention Recurrent Imputation (DEARI), which jointly estimates missing values and their associated uncertainty in heterogeneous multivariate time series. By jointly representing feature-wise correlations and temporal dynamics, we adopt a self attention mechanism, along with an effective residual component, to achieve a deep recurrent neural network with good imputation performance and stable convergence. We also leverage self-supervised metric learning to boost performance by optimizing sample similarity. Finally, we transform DEARI into a Bayesian neural network through a novel Bayesian marginalization strategy to produce stochastic DEARI, which outperforms its deterministic equivalent. Experiments show that DEARI surpasses the SOTA in diverse imputation tasks using real-world datasets, namely air quality control, healthcare and traffic.
Knowledge Enhanced Conditional Imputation for Healthcare Time-series
Jan 04, 2024Abstract:This study presents a novel approach to addressing the challenge of missing data in multivariate time series, with a particular focus on the complexities of healthcare data. Our Conditional Self-Attention Imputation (CSAI) model, grounded in a transformer-based framework, introduces a conditional hidden state initialization tailored to the intricacies of medical time series data. This methodology diverges from traditional imputation techniques by specifically targeting the imbalance in missing data distribution, a crucial aspect often overlooked in healthcare datasets. By integrating advanced knowledge embedding and a non-uniform masking strategy, CSAI adeptly adjusts to the distinct patterns of missing data in Electronic Health Records (EHRs).
Exploring Multimodal Large Language Models for Radiology Report Error-checking
Dec 20, 2023



Abstract:This paper proposes one of the first clinical applications of multimodal large language models (LLMs) as an assistant for radiologists to check errors in their reports. We created an evaluation dataset from two real-world radiology datasets (MIMIC-CXR and IU-Xray), with 1,000 subsampled reports each. A subset of original reports was modified to contain synthetic errors by introducing various type of mistakes. The evaluation contained two difficulty levels: SIMPLE for binary error-checking and COMPLEX for identifying error types. LLaVA (Large Language and Visual Assistant) variant models, including our instruction-tuned model, were used for the evaluation. Additionally, a domain expert evaluation was conducted on a small test set. At the SIMPLE level, the LLaVA v1.5 model outperformed other publicly available models. Instruction tuning significantly enhanced performance by 47.4% and 25.4% on MIMIC-CXR and IU-Xray data, respectively. The model also surpassed the domain experts accuracy in the MIMIC-CXR dataset by 1.67%. Notably, among the subsets (N=21) of the test set where a clinician did not achieve the correct conclusion, the LLaVA ensemble mode correctly identified 71.4% of these cases. This study marks a promising step toward utilizing multi-modal LLMs to enhance diagnostic accuracy in radiology. The ensemble model demonstrated comparable performance to clinicians, even capturing errors overlooked by humans. Nevertheless, future work is needed to improve the model ability to identify the types of inconsistency.
Summarisation of Electronic Health Records with Clinical Concept Guidance
Nov 14, 2022



Abstract:Brief Hospital Course (BHC) summaries are succinct summaries of an entire hospital encounter, embedded within discharge summaries, written by senior clinicians responsible for the overall care of a patient. Methods to automatically produce summaries from inpatient documentation would be invaluable in reducing clinician manual burden of summarising documents under high time-pressure to admit and discharge patients. Automatically producing these summaries from the inpatient course, is a complex, multi-document summarisation task, as source notes are written from various perspectives (e.g. nursing, doctor, radiology), during the course of the hospitalisation. We demonstrate a range of methods for BHC summarisation demonstrating the performance of deep learning summarisation models across extractive and abstractive summarisation scenarios. We also test a novel ensemble extractive and abstractive summarisation model that incorporates a medical concept ontology (SNOMED) as a clinical guidance signal and shows superior performance in 2 real-world clinical data sets.
Estimating Redundancy in Clinical Text
May 25, 2021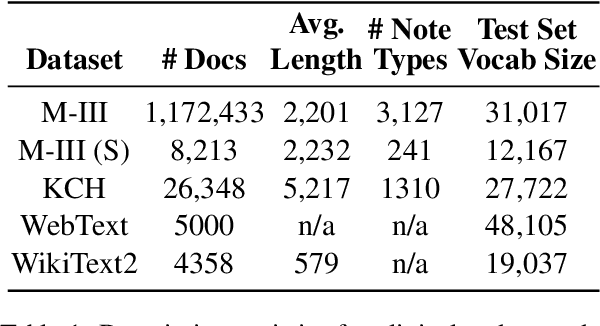
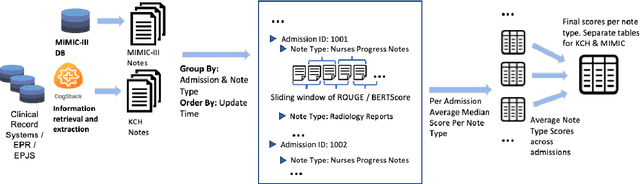
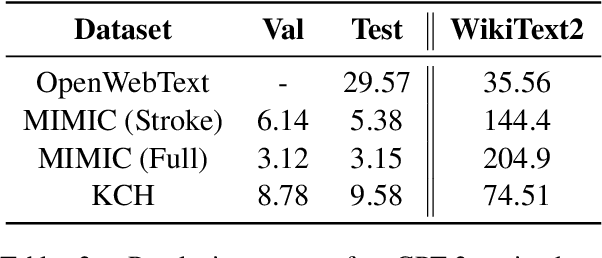
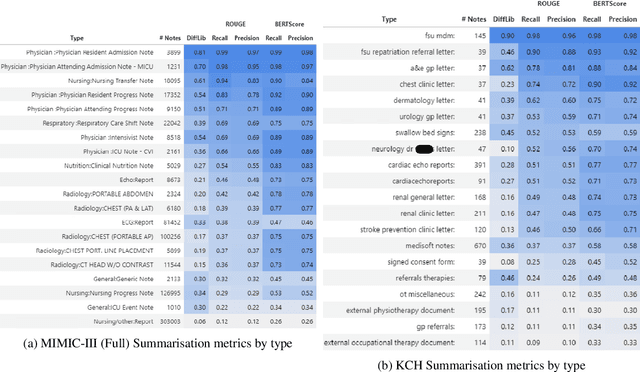
Abstract:The current mode of use of Electronic Health Record (EHR) elicits text redundancy. Clinicians often populate new documents by duplicating existing notes, then updating accordingly. Data duplication can lead to a propagation of errors, inconsistencies and misreporting of care. Therefore, quantifying information redundancy can play an essential role in evaluating innovations that operate on clinical narratives. This work is a quantitative examination of information redundancy in EHR notes. We present and evaluate two strategies to measure redundancy: an information-theoretic approach and a lexicosyntactic and semantic model. We evaluate the measures by training large Transformer-based language models using clinical text from a large openly available US-based ICU dataset and a large multi-site UK based Trust. By comparing the information-theoretic content of the trained models with open-domain language models, the language models trained using clinical text have shown ~1.5x to ~3x less efficient than open-domain corpora. Manual evaluation shows a high correlation with lexicosyntactic and semantic redundancy, with averages ~43 to ~65%.
Multi-domain Clinical Natural Language Processing with MedCAT: the Medical Concept Annotation Toolkit
Oct 02, 2020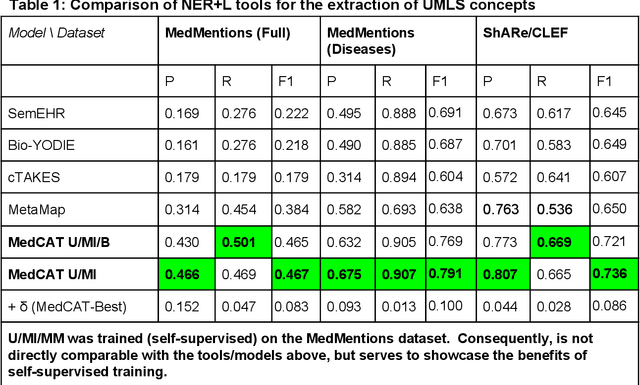
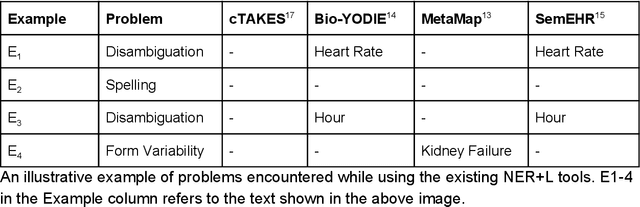
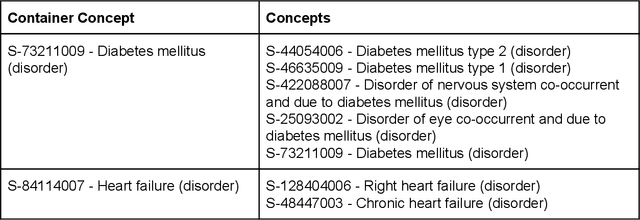
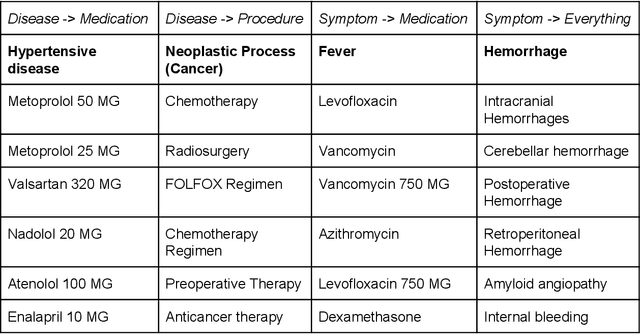
Abstract:Electronic health records (EHR) contain large volumes of unstructured text, requiring the application of Information Extraction (IE) technologies to enable clinical analysis. We present the open source Medical Concept Annotation Toolkit (MedCAT) that provides: a) a novel self-supervised machine learning algorithm for extracting concepts using any concept vocabulary including UMLS/SNOMED-CT; b) a feature-rich annotation interface for customizing and training IE models; and c) integrations to the broader CogStack ecosystem for vendor-agnostic health system deployment. We show improved performance in extracting UMLS concepts from open datasets ( F1 0.467-0.791 vs 0.384-0.691). Further real-world validation demonstrates SNOMED-CT extraction at 3 large London hospitals with self-supervised training over ~8.8B words from ~17M clinical records and further fine-tuning with ~6K clinician annotated examples. We show strong transferability ( F1 >0.94) between hospitals, datasets and concept types indicating cross-domain EHR-agnostic utility for accelerated clinical and research use cases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge