Yunzhi Xu
Nanbeige4.1-3B: A Small General Model that Reasons, Aligns, and Acts
Feb 13, 2026Abstract:We present Nanbeige4.1-3B, a unified generalist language model that simultaneously achieves strong agentic behavior, code generation, and general reasoning with only 3B parameters. To the best of our knowledge, it is the first open-source small language model (SLM) to achieve such versatility in a single model. To improve reasoning and preference alignment, we combine point-wise and pair-wise reward modeling, ensuring high-quality, human-aligned responses. For code generation, we design complexity-aware rewards in Reinforcement Learning, optimizing both correctness and efficiency. In deep search, we perform complex data synthesis and incorporate turn-level supervision during training. This enables stable long-horizon tool interactions, allowing Nanbeige4.1-3B to reliably execute up to 600 tool-call turns for complex problem-solving. Extensive experimental results show that Nanbeige4.1-3B significantly outperforms prior models of similar scale, such as Nanbeige4-3B-2511 and Qwen3-4B, even achieving superior performance compared to much larger models, such as Qwen3-30B-A3B. Our results demonstrate that small models can achieve both broad competence and strong specialization simultaneously, redefining the potential of 3B parameter models.
MiniMax-M1: Scaling Test-Time Compute Efficiently with Lightning Attention
Jun 16, 2025



Abstract:We introduce MiniMax-M1, the world's first open-weight, large-scale hybrid-attention reasoning model. MiniMax-M1 is powered by a hybrid Mixture-of-Experts (MoE) architecture combined with a lightning attention mechanism. The model is developed based on our previous MiniMax-Text-01 model, which contains a total of 456 billion parameters with 45.9 billion parameters activated per token. The M1 model natively supports a context length of 1 million tokens, 8x the context size of DeepSeek R1. Furthermore, the lightning attention mechanism in MiniMax-M1 enables efficient scaling of test-time compute. These properties make M1 particularly suitable for complex tasks that require processing long inputs and thinking extensively. MiniMax-M1 is trained using large-scale reinforcement learning (RL) on diverse problems including sandbox-based, real-world software engineering environments. In addition to M1's inherent efficiency advantage for RL training, we propose CISPO, a novel RL algorithm to further enhance RL efficiency. CISPO clips importance sampling weights rather than token updates, outperforming other competitive RL variants. Combining hybrid-attention and CISPO enables MiniMax-M1's full RL training on 512 H800 GPUs to complete in only three weeks, with a rental cost of just $534,700. We release two versions of MiniMax-M1 models with 40K and 80K thinking budgets respectively, where the 40K model represents an intermediate phase of the 80K training. Experiments on standard benchmarks show that our models are comparable or superior to strong open-weight models such as the original DeepSeek-R1 and Qwen3-235B, with particular strengths in complex software engineering, tool utilization, and long-context tasks. We publicly release MiniMax-M1 at https://github.com/MiniMax-AI/MiniMax-M1.
MiniMax-01: Scaling Foundation Models with Lightning Attention
Jan 14, 2025Abstract:We introduce MiniMax-01 series, including MiniMax-Text-01 and MiniMax-VL-01, which are comparable to top-tier models while offering superior capabilities in processing longer contexts. The core lies in lightning attention and its efficient scaling. To maximize computational capacity, we integrate it with Mixture of Experts (MoE), creating a model with 32 experts and 456 billion total parameters, of which 45.9 billion are activated for each token. We develop an optimized parallel strategy and highly efficient computation-communication overlap techniques for MoE and lightning attention. This approach enables us to conduct efficient training and inference on models with hundreds of billions of parameters across contexts spanning millions of tokens. The context window of MiniMax-Text-01 can reach up to 1 million tokens during training and extrapolate to 4 million tokens during inference at an affordable cost. Our vision-language model, MiniMax-VL-01 is built through continued training with 512 billion vision-language tokens. Experiments on both standard and in-house benchmarks show that our models match the performance of state-of-the-art models like GPT-4o and Claude-3.5-Sonnet while offering 20-32 times longer context window. We publicly release MiniMax-01 at https://github.com/MiniMax-AI.
Fetal Brain Tissue Annotation and Segmentation Challenge Results
Apr 20, 2022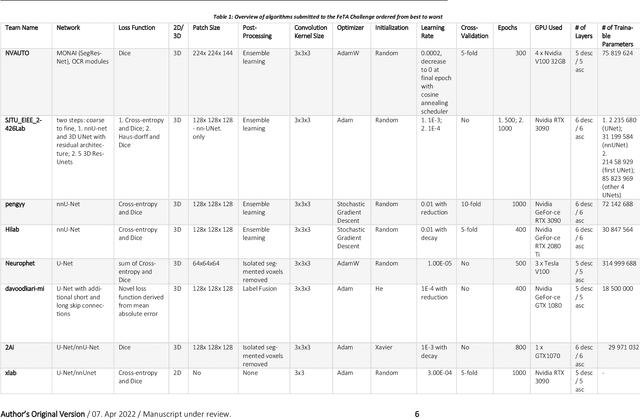
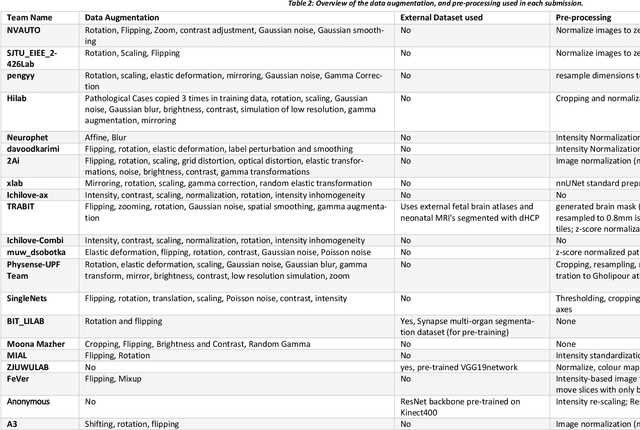
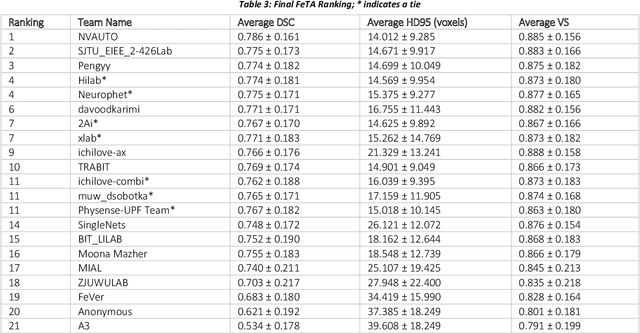
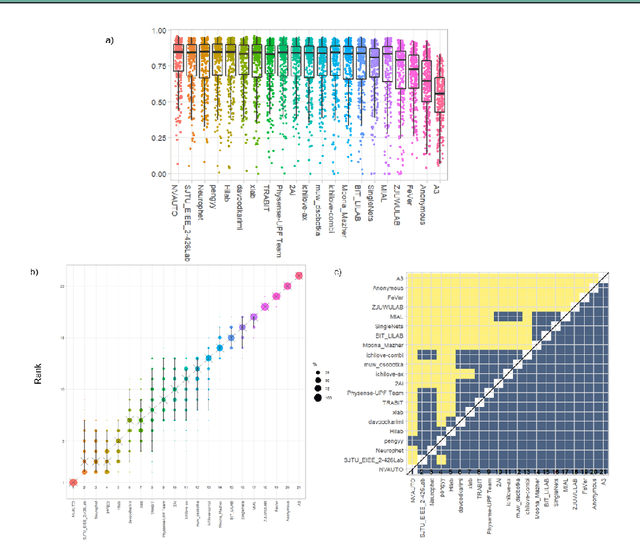
Abstract:In-utero fetal MRI is emerging as an important tool in the diagnosis and analysis of the developing human brain. Automatic segmentation of the developing fetal brain is a vital step in the quantitative analysis of prenatal neurodevelopment both in the research and clinical context. However, manual segmentation of cerebral structures is time-consuming and prone to error and inter-observer variability. Therefore, we organized the Fetal Tissue Annotation (FeTA) Challenge in 2021 in order to encourage the development of automatic segmentation algorithms on an international level. The challenge utilized FeTA Dataset, an open dataset of fetal brain MRI reconstructions segmented into seven different tissues (external cerebrospinal fluid, grey matter, white matter, ventricles, cerebellum, brainstem, deep grey matter). 20 international teams participated in this challenge, submitting a total of 21 algorithms for evaluation. In this paper, we provide a detailed analysis of the results from both a technical and clinical perspective. All participants relied on deep learning methods, mainly U-Nets, with some variability present in the network architecture, optimization, and image pre- and post-processing. The majority of teams used existing medical imaging deep learning frameworks. The main differences between the submissions were the fine tuning done during training, and the specific pre- and post-processing steps performed. The challenge results showed that almost all submissions performed similarly. Four of the top five teams used ensemble learning methods. However, one team's algorithm performed significantly superior to the other submissions, and consisted of an asymmetrical U-Net network architecture. This paper provides a first of its kind benchmark for future automatic multi-tissue segmentation algorithms for the developing human brain in utero.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge