Yufan He
Disruptive Autoencoders: Leveraging Low-level features for 3D Medical Image Pre-training
Jul 31, 2023Abstract:Harnessing the power of pre-training on large-scale datasets like ImageNet forms a fundamental building block for the progress of representation learning-driven solutions in computer vision. Medical images are inherently different from natural images as they are acquired in the form of many modalities (CT, MR, PET, Ultrasound etc.) and contain granulated information like tissue, lesion, organs etc. These characteristics of medical images require special attention towards learning features representative of local context. In this work, we focus on designing an effective pre-training framework for 3D radiology images. First, we propose a new masking strategy called local masking where the masking is performed across channel embeddings instead of tokens to improve the learning of local feature representations. We combine this with classical low-level perturbations like adding noise and downsampling to further enable low-level representation learning. To this end, we introduce Disruptive Autoencoders, a pre-training framework that attempts to reconstruct the original image from disruptions created by a combination of local masking and low-level perturbations. Additionally, we also devise a cross-modal contrastive loss (CMCL) to accommodate the pre-training of multiple modalities in a single framework. We curate a large-scale dataset to enable pre-training of 3D medical radiology images (MRI and CT). The proposed pre-training framework is tested across multiple downstream tasks and achieves state-of-the-art performance. Notably, our proposed method tops the public test leaderboard of BTCV multi-organ segmentation challenge.
Spatially-varying Regularization with Conditional Transformer for Unsupervised Image Registration
Mar 10, 2023Abstract:In the past, optimization-based registration models have used spatially-varying regularization to account for deformation variations in different image regions. However, deep learning-based registration models have mostly relied on spatially-invariant regularization. Here, we introduce an end-to-end framework that uses neural networks to learn a spatially-varying deformation regularizer directly from data. The hyperparameter of the proposed regularizer is conditioned into the network, enabling easy tuning of the regularization strength. The proposed method is built upon a Transformer-based model, but it can be readily adapted to any network architecture. We thoroughly evaluated the proposed approach using publicly available datasets and observed a significant performance improvement while maintaining smooth deformation. The source code of this work will be made available after publication.
Deformable Cross-Attention Transformer for Medical Image Registration
Mar 10, 2023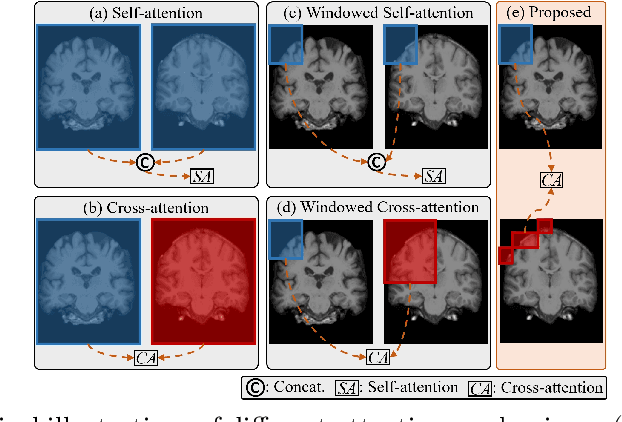
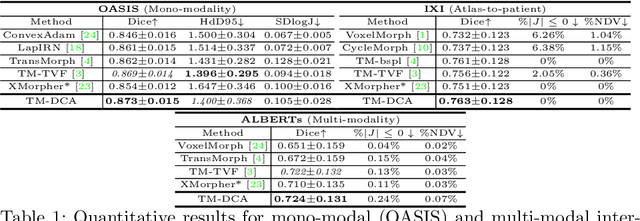
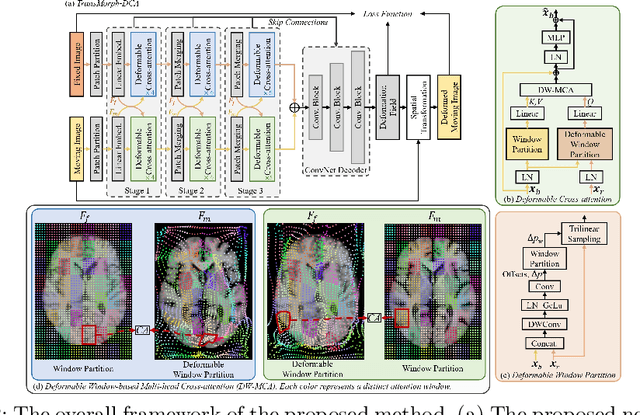
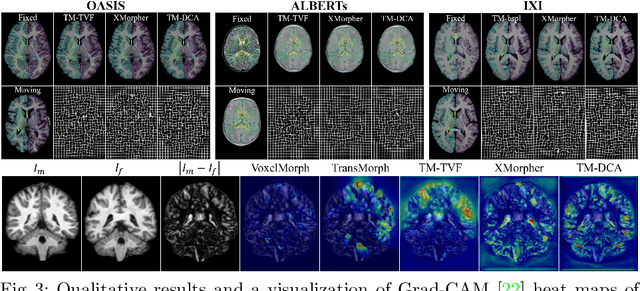
Abstract:Transformers have recently shown promise for medical image applications, leading to an increasing interest in developing such models for medical image registration. Recent advancements in designing registration Transformers have focused on using cross-attention (CA) to enable a more precise understanding of spatial correspondences between moving and fixed images. Here, we propose a novel CA mechanism that computes windowed attention using deformable windows. In contrast to existing CA mechanisms that require intensive computational complexity by either computing CA globally or locally with a fixed and expanded search window, the proposed deformable CA can selectively sample a diverse set of features over a large search window while maintaining low computational complexity. The proposed model was extensively evaluated on multi-modal, mono-modal, and atlas-to-patient registration tasks, demonstrating promising performance against state-of-the-art methods and indicating its effectiveness for medical image registration. The source code for this work will be available after publication.
MONAI: An open-source framework for deep learning in healthcare
Nov 04, 2022



Abstract:Artificial Intelligence (AI) is having a tremendous impact across most areas of science. Applications of AI in healthcare have the potential to improve our ability to detect, diagnose, prognose, and intervene on human disease. For AI models to be used clinically, they need to be made safe, reproducible and robust, and the underlying software framework must be aware of the particularities (e.g. geometry, physiology, physics) of medical data being processed. This work introduces MONAI, a freely available, community-supported, and consortium-led PyTorch-based framework for deep learning in healthcare. MONAI extends PyTorch to support medical data, with a particular focus on imaging, and provide purpose-specific AI model architectures, transformations and utilities that streamline the development and deployment of medical AI models. MONAI follows best practices for software-development, providing an easy-to-use, robust, well-documented, and well-tested software framework. MONAI preserves the simple, additive, and compositional approach of its underlying PyTorch libraries. MONAI is being used by and receiving contributions from research, clinical and industrial teams from around the world, who are pursuing applications spanning nearly every aspect of healthcare.
Automated head and neck tumor segmentation from 3D PET/CT
Sep 22, 2022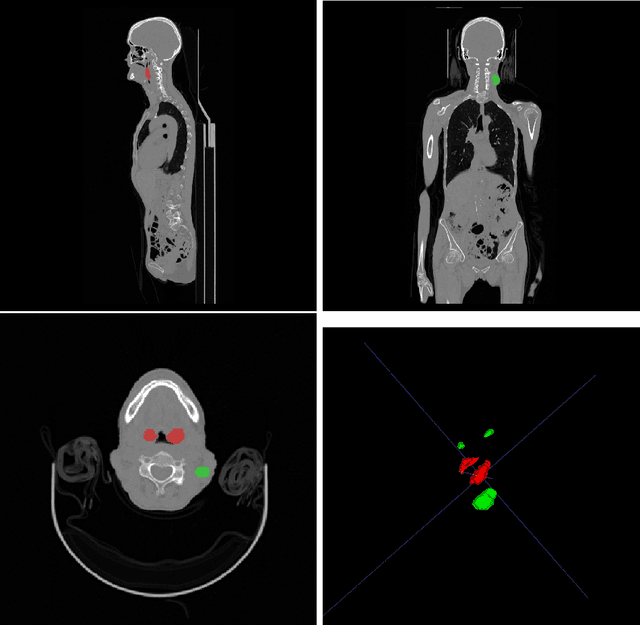
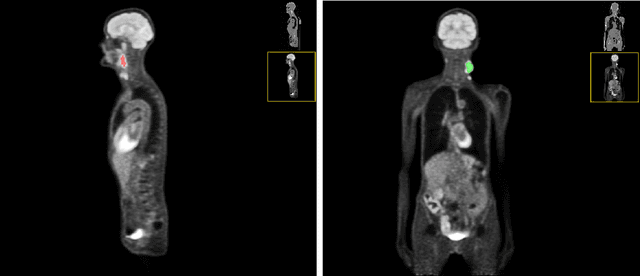
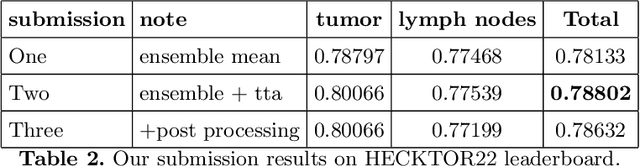
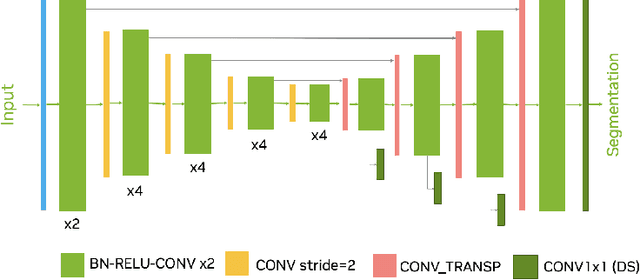
Abstract:Head and neck tumor segmentation challenge (HECKTOR) 2022 offers a platform for researchers to compare their solutions to segmentation of tumors and lymph nodes from 3D CT and PET images. In this work, we describe our solution to HECKTOR 2022 segmentation task. We re-sample all images to a common resolution, crop around head and neck region, and train SegResNet semantic segmentation network from MONAI. We use 5-fold cross validation to select best model checkpoints. The final submission is an ensemble of 15 models from 3 runs. Our solution (team name NVAUTO) achieves the 1st place on the HECKTOR22 challenge leaderboard with an aggregated dice score of 0.78802.
Automated segmentation of intracranial hemorrhages from 3D CT
Sep 21, 2022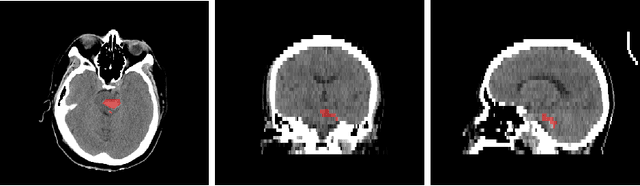
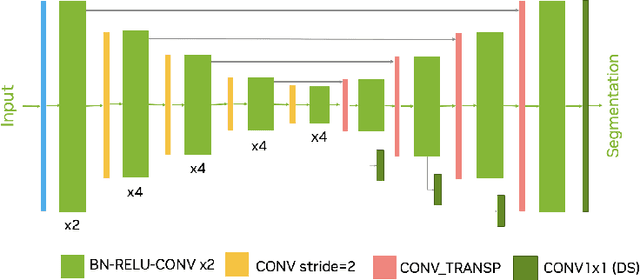
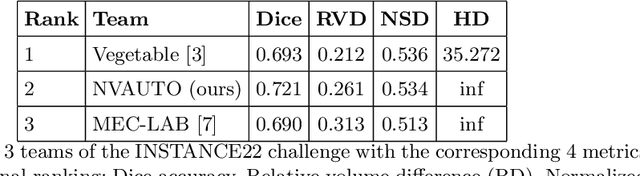
Abstract:Intracranial hemorrhage segmentation challenge (INSTANCE 2022) offers a platform for researchers to compare their solutions to segmentation of hemorrhage stroke regions from 3D CTs. In this work, we describe our solution to INSTANCE 2022. We use a 2D segmentation network, SegResNet from MONAI, operating slice-wise without resampling. The final submission is an ensemble of 18 models. Our solution (team name NVAUTO) achieves the top place in terms of Dice metric (0.721), and overall rank 2. It is implemented with Auto3DSeg.
Automated ischemic stroke lesion segmentation from 3D MRI
Sep 21, 2022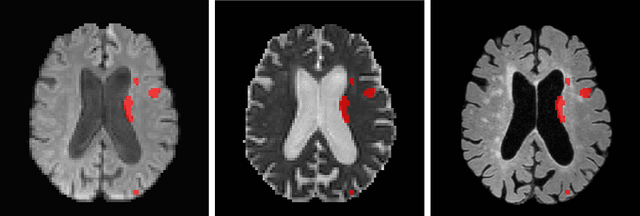
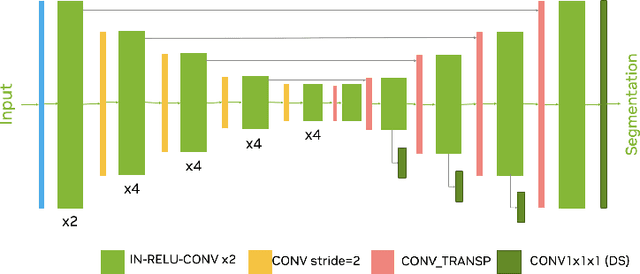
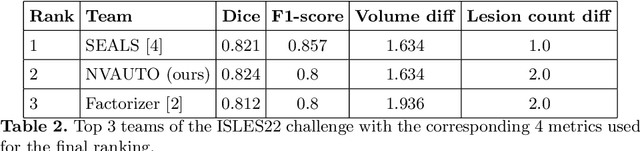
Abstract:Ischemic Stroke Lesion Segmentation challenge (ISLES 2022) offers a platform for researchers to compare their solutions to 3D segmentation of ischemic stroke regions from 3D MRIs. In this work, we describe our solution to ISLES 2022 segmentation task. We re-sample all images to a common resolution, use two input MRI modalities (DWI and ADC) and train SegResNet semantic segmentation network from MONAI. The final submission is an ensemble of 15 models (from 3 runs of 5-fold cross validation). Our solution (team name NVAUTO) achieves the top place in terms of Dice metric (0.824), and overall rank 2 (based on the combined metric ranking).
HyperSegNAS: Bridging One-Shot Neural Architecture Search with 3D Medical Image Segmentation using HyperNet
Dec 20, 2021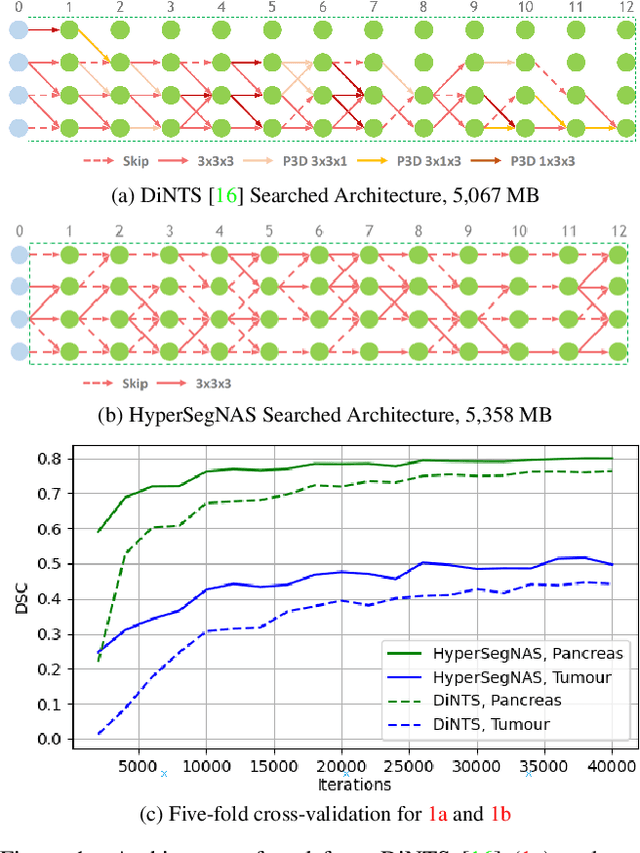
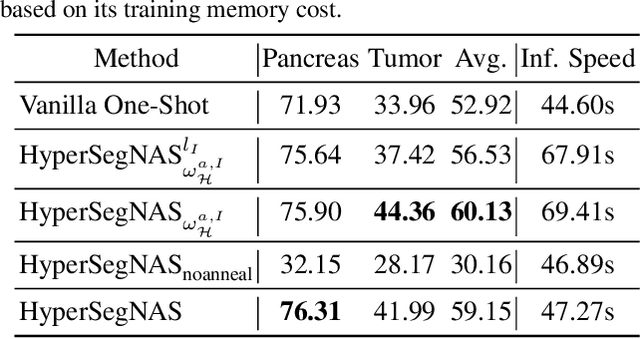
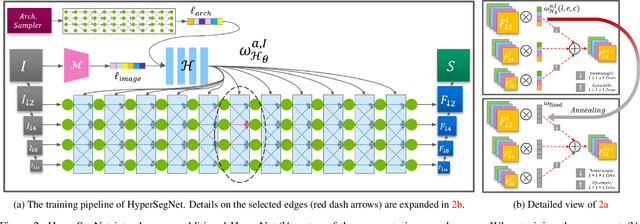
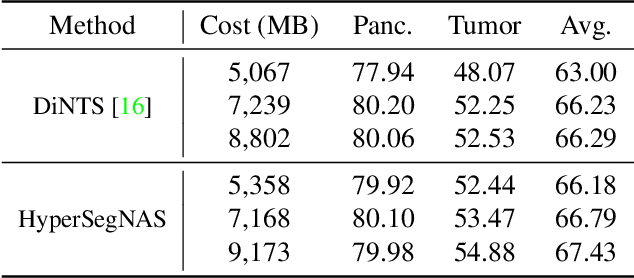
Abstract:Semantic segmentation of 3D medical images is a challenging task due to the high variability of the shape and pattern of objects (such as organs or tumors). Given the recent success of deep learning in medical image segmentation, Neural Architecture Search (NAS) has been introduced to find high-performance 3D segmentation network architectures. However, because of the massive computational requirements of 3D data and the discrete optimization nature of architecture search, previous NAS methods require a long search time or necessary continuous relaxation, and commonly lead to sub-optimal network architectures. While one-shot NAS can potentially address these disadvantages, its application in the segmentation domain has not been well studied in the expansive multi-scale multi-path search space. To enable one-shot NAS for medical image segmentation, our method, named HyperSegNAS, introduces a HyperNet to assist super-net training by incorporating architecture topology information. Such a HyperNet can be removed once the super-net is trained and introduces no overhead during architecture search. We show that HyperSegNAS yields better performing and more intuitive architectures compared to the previous state-of-the-art (SOTA) segmentation networks; furthermore, it can quickly and accurately find good architecture candidates under different computing constraints. Our method is evaluated on public datasets from the Medical Segmentation Decathlon (MSD) challenge, and achieves SOTA performances.
TransMorph: Transformer for unsupervised medical image registration
Nov 23, 2021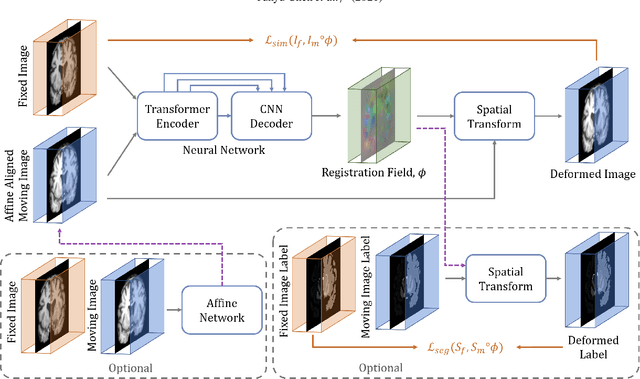

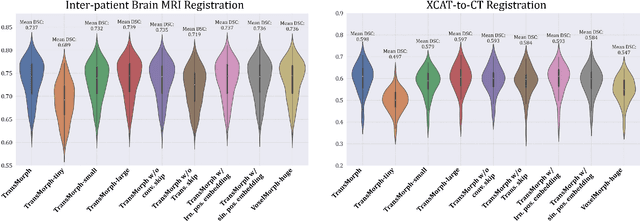
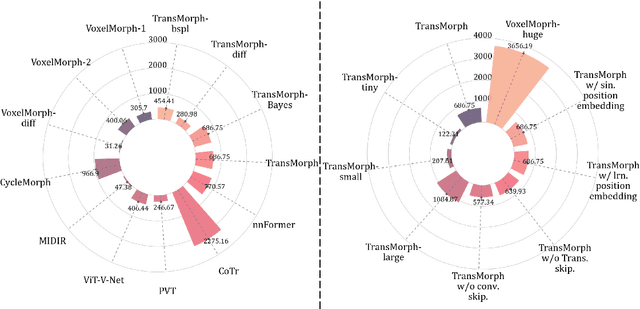
Abstract:In the last decade, convolutional neural networks (ConvNets) have dominated the field of medical image analysis. However, it is found that the performances of ConvNets may still be limited by their inability to model long-range spatial relations between voxels in an image. Numerous vision Transformers have been proposed recently to address the shortcomings of ConvNets, demonstrating state-of-the-art performances in many medical imaging applications. Transformers may be a strong candidate for image registration because their self-attention mechanism enables a more precise comprehension of the spatial correspondence between moving and fixed images. In this paper, we present TransMorph, a hybrid Transformer-ConvNet model for volumetric medical image registration. We also introduce three variants of TransMorph, with two diffeomorphic variants ensuring the topology-preserving deformations and a Bayesian variant producing a well-calibrated registration uncertainty estimate. The proposed models are extensively validated against a variety of existing registration methods and Transformer architectures using volumetric medical images from two applications: inter-patient brain MRI registration and phantom-to-CT registration. Qualitative and quantitative results demonstrate that TransMorph and its variants lead to a substantial performance improvement over the baseline methods, demonstrating the effectiveness of Transformers for medical image registration.
ViT-V-Net: Vision Transformer for Unsupervised Volumetric Medical Image Registration
Apr 13, 2021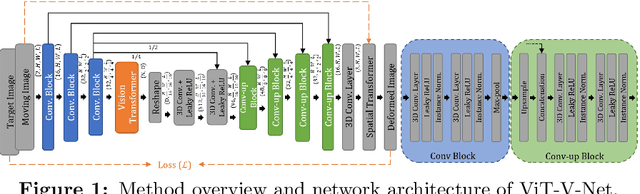
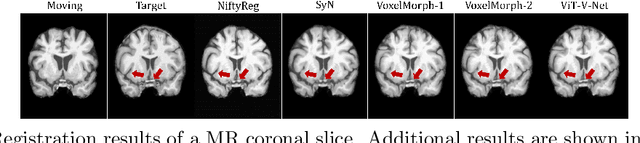
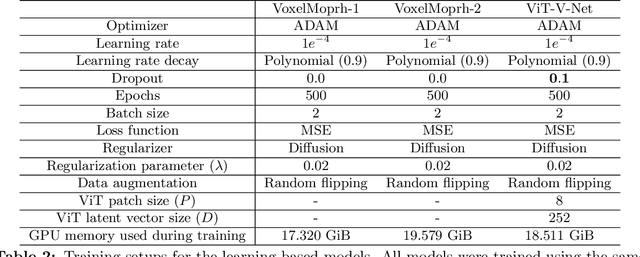
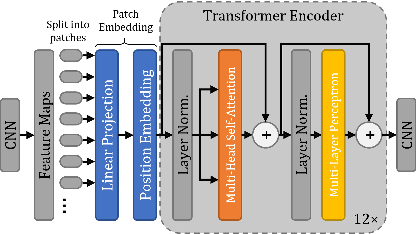
Abstract:In the last decade, convolutional neural networks (ConvNets) have dominated and achieved state-of-the-art performances in a variety of medical imaging applications. However, the performances of ConvNets are still limited by lacking the understanding of long-range spatial relations in an image. The recently proposed Vision Transformer (ViT) for image classification uses a purely self-attention-based model that learns long-range spatial relations to focus on the relevant parts of an image. Nevertheless, ViT emphasizes the low-resolution features because of the consecutive downsamplings, result in a lack of detailed localization information, making it unsuitable for image registration. Recently, several ViT-based image segmentation methods have been combined with ConvNets to improve the recovery of detailed localization information. Inspired by them, we present ViT-V-Net, which bridges ViT and ConvNet to provide volumetric medical image registration. The experimental results presented here demonstrate that the proposed architecture achieves superior performance to several top-performing registration methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge