Yingtai Li
DiffVP: Differential Visual Semantic Prompting for LLM-Based CT Report Generation
Mar 18, 2026Abstract:While large language models (LLMs) have advanced CT report generation, existing methods typically encode 3D volumes holistically, failing to distinguish informative cues from redundant anatomical background. Inspired by radiological cognitive subtraction, we propose Differential Visual Prompting (DiffVP), which conditions report generation on explicit, high-level semantic scan-to-reference differences rather than solely on absolute visual features. DiffVP employs a hierarchical difference extractor to capture complementary global and local semantic discrepancies into a shared latent space, along with a difference-to-prompt generator that transforms these signals into learnable visual prefix tokens for LLM conditioning. These difference prompts serve as structured conditioning signals that implicitly suppress invariant anatomy while amplifying diagnostically relevant visual evidence, thereby facilitating accurate report generation without explicit lesion localization. On two large-scale benchmarks, DiffVP consistently outperforms prior methods, improving the average BLEU-1-4 by +10.98 and +4.36, respectively, and further boosts clinical efficacy on RadGenome-ChestCT (F1 score 0.421). All codes will be released at https://github.com/ArielTYH/DiffVP/.
QCAgent: An agentic framework for quality-controllable pathology report generation from whole slide image
Mar 02, 2026Abstract:Recent methods for pathology report generation from whole-slide image (WSI) are capable of producing slide-level diagnostic descriptions but fail to ground fine-grained statements in localized visual evidence. Furthermore, they lack control over which diagnostic details to include and how to verify them. Inspired by emerging agentic analysis paradigms and the diagnostic workflow of pathologists,who selectively examine multiple fields of view, we propose QCAgent, an agentic framework for quality-controllable WSI report generation. The core innovations of this framework are as follows: (i) it incorporates a customized critique mechanism guided by a user-defined checklist specifying required diagnostic details and constraints; (ii) it re-identifies informative regions in the WSI based on the critique feedback and text-patch semantic retrieval, a process that iteratively enriches and reconciles the report. Experiments demonstrate that by making report requirements explicitly prompt-defined, constraint-aware, and verifiable through evidence-grounded refinement, QCAgent enables controllable generation of clinically meaningful and high-coverage pathology reports from WSI.
More performant and scalable: Rethinking contrastive vision-language pre-training of radiology in the LLM era
Sep 16, 2025Abstract:The emergence of Large Language Models (LLMs) presents unprecedented opportunities to revolutionize medical contrastive vision-language pre-training. In this paper, we show how LLMs can facilitate large-scale supervised pre-training, thereby advancing vision-language alignment. We begin by demonstrate that modern LLMs can automatically extract diagnostic labels from radiology reports with remarkable precision (>96\% AUC in our experiments) without complex prompt engineering, enabling the creation of large-scale "silver-standard" datasets at a minimal cost (~\$3 for 50k CT image-report pairs). Further, we find that vision encoder trained on this "silver-standard" dataset achieves performance comparable to those trained on labels extracted by specialized BERT-based models, thereby democratizing the access to large-scale supervised pre-training. Building on this foundation, we proceed to reveal that supervised pre-training fundamentally improves contrastive vision-language alignment. Our approach achieves state-of-the-art performance using only a 3D ResNet-18 with vanilla CLIP training, including 83.8\% AUC for zero-shot diagnosis on CT-RATE, 77.3\% AUC on RAD-ChestCT, and substantial improvements in cross-modal retrieval (MAP@50=53.7\% for image-image, Recall@100=52.2\% for report-image). These results demonstrate the potential of utilizing LLMs to facilitate {\bf more performant and scalable} medical AI systems. Our code is avaiable at https://github.com/SadVoxel/More-performant-and-scalable.
Dyna3DGR: 4D Cardiac Motion Tracking with Dynamic 3D Gaussian Representation
Jul 22, 2025Abstract:Accurate analysis of cardiac motion is crucial for evaluating cardiac function. While dynamic cardiac magnetic resonance imaging (CMR) can capture detailed tissue motion throughout the cardiac cycle, the fine-grained 4D cardiac motion tracking remains challenging due to the homogeneous nature of myocardial tissue and the lack of distinctive features. Existing approaches can be broadly categorized into image based and representation-based, each with its limitations. Image-based methods, including both raditional and deep learning-based registration approaches, either struggle with topological consistency or rely heavily on extensive training data. Representation-based methods, while promising, often suffer from loss of image-level details. To address these limitations, we propose Dynamic 3D Gaussian Representation (Dyna3DGR), a novel framework that combines explicit 3D Gaussian representation with implicit neural motion field modeling. Our method simultaneously optimizes cardiac structure and motion in a self-supervised manner, eliminating the need for extensive training data or point-to-point correspondences. Through differentiable volumetric rendering, Dyna3DGR efficiently bridges continuous motion representation with image-space alignment while preserving both topological and temporal consistency. Comprehensive evaluations on the ACDC dataset demonstrate that our approach surpasses state-of-the-art deep learning-based diffeomorphic registration methods in tracking accuracy. The code will be available in https://github.com/windrise/Dyna3DGR.
A General Knowledge Injection Framework for ICD Coding
May 24, 2025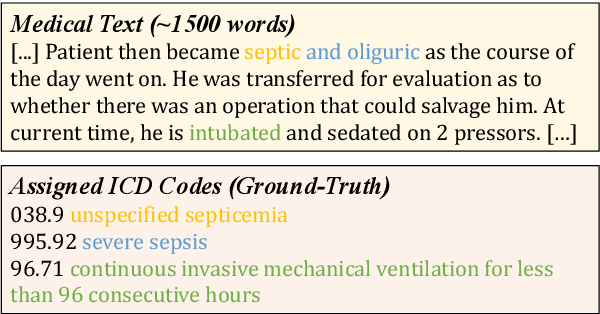
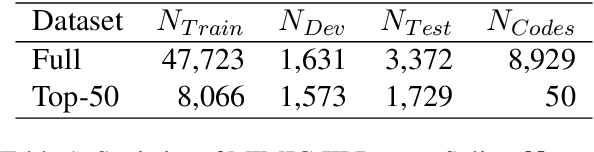
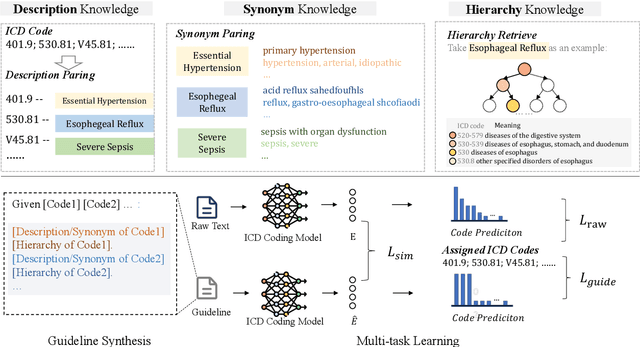
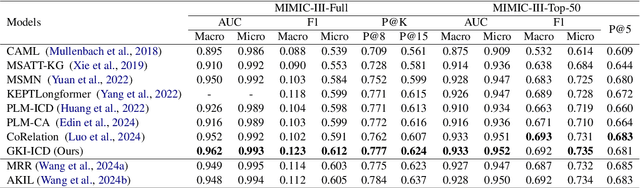
Abstract:ICD Coding aims to assign a wide range of medical codes to a medical text document, which is a popular and challenging task in the healthcare domain. To alleviate the problems of long-tail distribution and the lack of annotations of code-specific evidence, many previous works have proposed incorporating code knowledge to improve coding performance. However, existing methods often focus on a single type of knowledge and design specialized modules that are complex and incompatible with each other, thereby limiting their scalability and effectiveness. To address this issue, we propose GKI-ICD, a novel, general knowledge injection framework that integrates three key types of knowledge, namely ICD Description, ICD Synonym, and ICD Hierarchy, without specialized design of additional modules. The comprehensive utilization of the above knowledge, which exhibits both differences and complementarity, can effectively enhance the ICD coding performance. Extensive experiments on existing popular ICD coding benchmarks demonstrate the effectiveness of GKI-ICD, which achieves the state-of-the-art performance on most evaluation metrics. Code is available at https://github.com/xuzhang0112/GKI-ICD.
AA-CLIP: Enhancing Zero-shot Anomaly Detection via Anomaly-Aware CLIP
Mar 09, 2025Abstract:Anomaly detection (AD) identifies outliers for applications like defect and lesion detection. While CLIP shows promise for zero-shot AD tasks due to its strong generalization capabilities, its inherent Anomaly-Unawareness leads to limited discrimination between normal and abnormal features. To address this problem, we propose Anomaly-Aware CLIP (AA-CLIP), which enhances CLIP's anomaly discrimination ability in both text and visual spaces while preserving its generalization capability. AA-CLIP is achieved through a straightforward yet effective two-stage approach: it first creates anomaly-aware text anchors to differentiate normal and abnormal semantics clearly, then aligns patch-level visual features with these anchors for precise anomaly localization. This two-stage strategy, with the help of residual adapters, gradually adapts CLIP in a controlled manner, achieving effective AD while maintaining CLIP's class knowledge. Extensive experiments validate AA-CLIP as a resource-efficient solution for zero-shot AD tasks, achieving state-of-the-art results in industrial and medical applications. The code is available at https://github.com/Mwxinnn/AA-CLIP.
* 8 pages, 7 figures
3DGR-CAR: Coronary artery reconstruction from ultra-sparse 2D X-ray views with a 3D Gaussians representation
Oct 01, 2024Abstract:Reconstructing 3D coronary arteries is important for coronary artery disease diagnosis, treatment planning and operation navigation. Traditional reconstruction techniques often require many projections, while reconstruction from sparse-view X-ray projections is a potential way of reducing radiation dose. However, the extreme sparsity of coronary arteries in a 3D volume and ultra-limited number of projections pose significant challenges for efficient and accurate 3D reconstruction. To this end, we propose 3DGR-CAR, a 3D Gaussian Representation for Coronary Artery Reconstruction from ultra-sparse X-ray projections. We leverage 3D Gaussian representation to avoid the inefficiency caused by the extreme sparsity of coronary artery data and propose a Gaussian center predictor to overcome the noisy Gaussian initialization from ultra-sparse view projections. The proposed scheme enables fast and accurate 3D coronary artery reconstruction with only 2 views. Experimental results on two datasets indicate that the proposed approach significantly outperforms other methods in terms of voxel accuracy and visual quality of coronary arteries. The code will be available in https://github.com/windrise/3DGR-CAR.
MambaMIM: Pre-training Mamba with State Space Token-interpolation
Aug 15, 2024Abstract:Generative self-supervised learning demonstrates outstanding representation learning capabilities in both Convolutional Neural Networks (CNNs) and Vision Transformers (ViTs). However, there are currently no generative pre-training methods related to selective state space models (Mamba) that can handle long-range dependencies effectively. To address this challenge, we introduce a generative self-supervised learning method for Mamba (MambaMIM) based on Selective Structure State Space Sequence Token-interpolation (S6T), a general-purpose pre-training method for arbitrary Mamba architectures. Our method, MambaMIM, incorporates a bottom-up 3D hybrid masking strategy in the encoder to maintain masking consistency across different architectures. Additionally, S6T is employed to learn causal relationships between the masked sequence in the state space. MambaMIM can be used on any single or hybrid Mamba architectures to enhance the Mamba long-range representation capability. Extensive downstream experiments reveal the feasibility and advancement of using Mamba for pre-training medical image tasks. The code is available at: https://github.com/FengheTan9/MambaMIM
Sparse-view CT Reconstruction with 3D Gaussian Volumetric Representation
Dec 25, 2023



Abstract:Sparse-view CT is a promising strategy for reducing the radiation dose of traditional CT scans, but reconstructing high-quality images from incomplete and noisy data is challenging. Recently, 3D Gaussian has been applied to model complex natural scenes, demonstrating fast convergence and better rendering of novel views compared to implicit neural representations (INRs). Taking inspiration from the successful application of 3D Gaussians in natural scene modeling and novel view synthesis, we investigate their potential for sparse-view CT reconstruction. We leverage prior information from the filtered-backprojection reconstructed image to initialize the Gaussians; and update their parameters via comparing difference in the projection space. Performance is further enhanced by adaptive density control. Compared to INRs, 3D Gaussians benefit more from prior information to explicitly bypass learning in void spaces and allocate the capacity efficiently, accelerating convergence. 3D Gaussians also efficiently learn high-frequency details. Trained in a self-supervised manner, 3D Gaussians avoid the need for large-scale paired data. Our experiments on the AAPM-Mayo dataset demonstrate that 3D Gaussians can provide superior performance compared to INR-based methods. This work is in progress, and the code will be publicly available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge