Xundong Wu
Darwin3: A large-scale neuromorphic chip with a Novel ISA and On-Chip Learning
Dec 29, 2023



Abstract:Spiking Neural Networks (SNNs) are gaining increasing attention for their biological plausibility and potential for improved computational efficiency. To match the high spatial-temporal dynamics in SNNs, neuromorphic chips are highly desired to execute SNNs in hardware-based neuron and synapse circuits directly. This paper presents a large-scale neuromorphic chip named Darwin3 with a novel instruction set architecture(ISA), which comprises 10 primary instructions and a few extended instructions. It supports flexible neuron model programming and local learning rule designs. The Darwin3 chip architecture is designed in a mesh of computing nodes with an innovative routing algorithm. We used a compression mechanism to represent synaptic connections, significantly reducing memory usage. The Darwin3 chip supports up to 2.35 million neurons, making it the largest of its kind in neuron scale. The experimental results showed that code density was improved up to 28.3x in Darwin3, and neuron core fan-in and fan-out were improved up to 4096x and 3072x by connection compression compared to the physical memory depth. Our Darwin3 chip also provided memory saving between 6.8X and 200.8X when mapping convolutional spiking neural networks (CSNN) onto the chip, demonstrating state-of-the-art performance in accuracy and latency compared to other neuromorphic chips.
Mitigating Communication Costs in Neural Networks: The Role of Dendritic Nonlinearity
Jun 21, 2023



Abstract:Our comprehension of biological neuronal networks has profoundly influenced the evolution of artificial neural networks (ANNs). However, the neurons employed in ANNs exhibit remarkable deviations from their biological analogs, mainly due to the absence of complex dendritic trees encompassing local nonlinearity. Despite such disparities, previous investigations have demonstrated that point neurons can functionally substitute dendritic neurons in executing computational tasks. In this study, we scrutinized the importance of nonlinear dendrites within neural networks. By employing machine-learning methodologies, we assessed the impact of dendritic structure nonlinearity on neural network performance. Our findings reveal that integrating dendritic structures can substantially enhance model capacity and performance while keeping signal communication costs effectively restrained. This investigation offers pivotal insights that hold considerable implications for the development of future neural network accelerators.
Cross-Channel Intragroup Sparsity Neural Network
Oct 26, 2019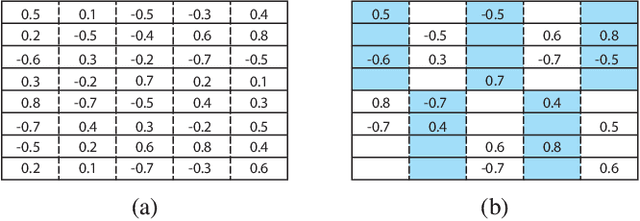
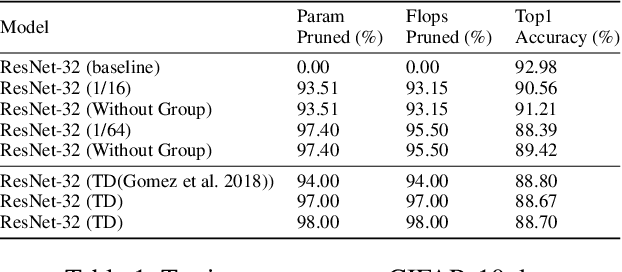
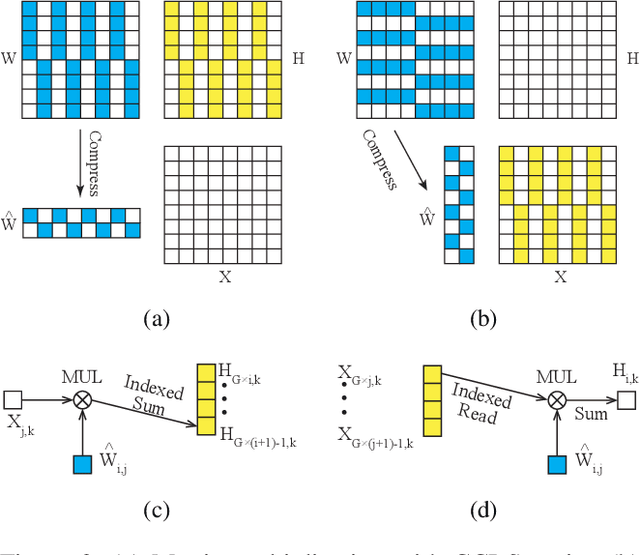
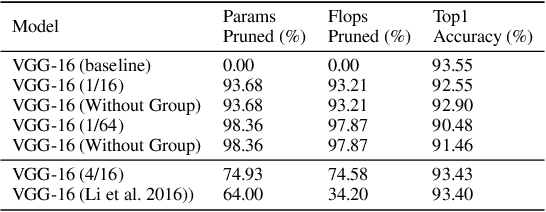
Abstract:Modern deep neural network models generally build upon heavy over-parameterization for their exceptional performance. Network pruning is one often employed approach to obtain less demanding models for their deployment. Fine-grained pruning, while can achieve good model compression ratio, introduces irregularity in the computing data flow, often does not give improved model inference efficiency. Coarse-grained model pruning, while allows good inference speed through removing network weights in whole groups, for example, a whole filter, can lead to significant model performance deterioration. In this study, we introduce the cross-channel intragroup (CCI) sparsity structure that can avoid the inference inefficiency of fine-grained pruning while maintaining outstanding model performance.
Binarized Neural Networks on the ImageNet Classification Task
Nov 19, 2016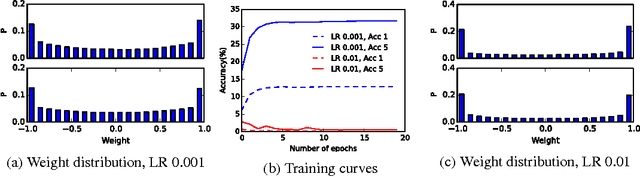
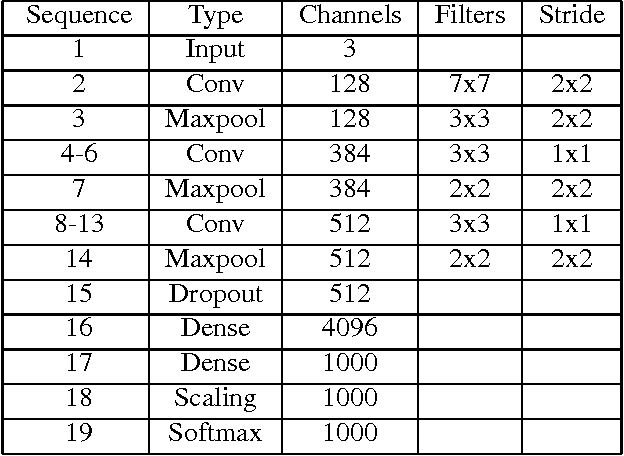
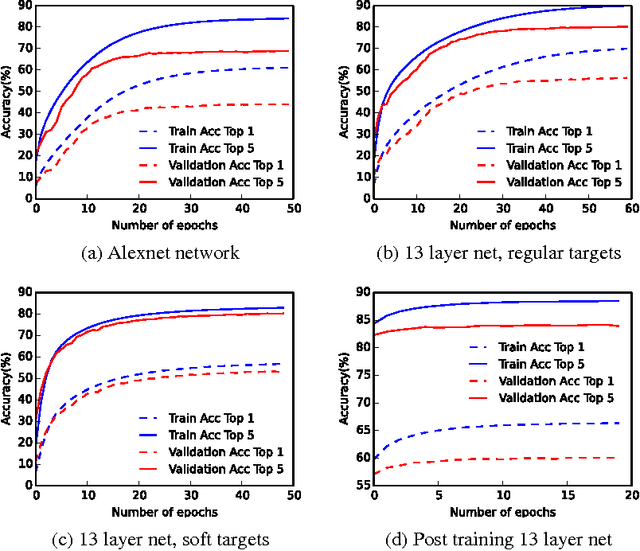
Abstract:We trained Binarized Neural Networks (BNNs) on the high resolution ImageNet ILSVRC-2102 dataset classification task and achieved a good performance. With a moderate size network of 13 layers, we obtained top-5 classification accuracy rate of 84.1 % on validation set through network distillation, much better than previous published results of 73.2% on XNOR network and 69.1% on binarized GoogleNET. We expect networks of better performance can be obtained by following our current strategies. We provide a detailed discussion and preliminary analysis on strategies used in the network training.
An Iterative Convolutional Neural Network Algorithm Improves Electron Microscopy Image Segmentation
Jun 18, 2015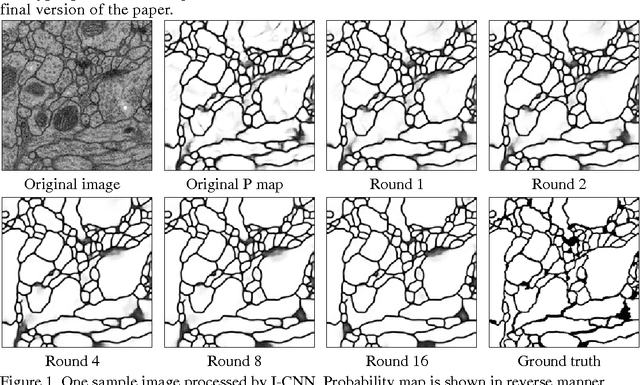
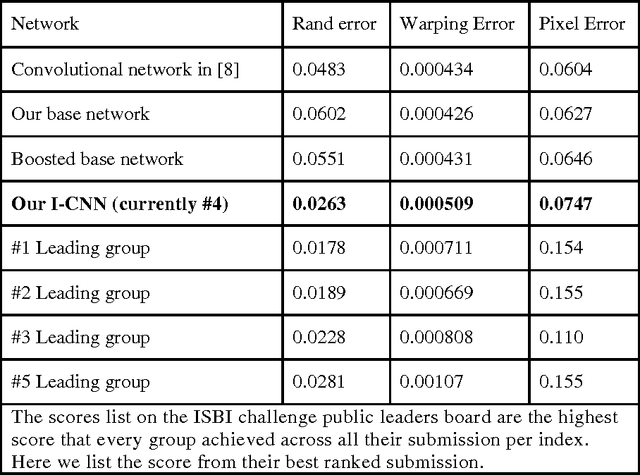
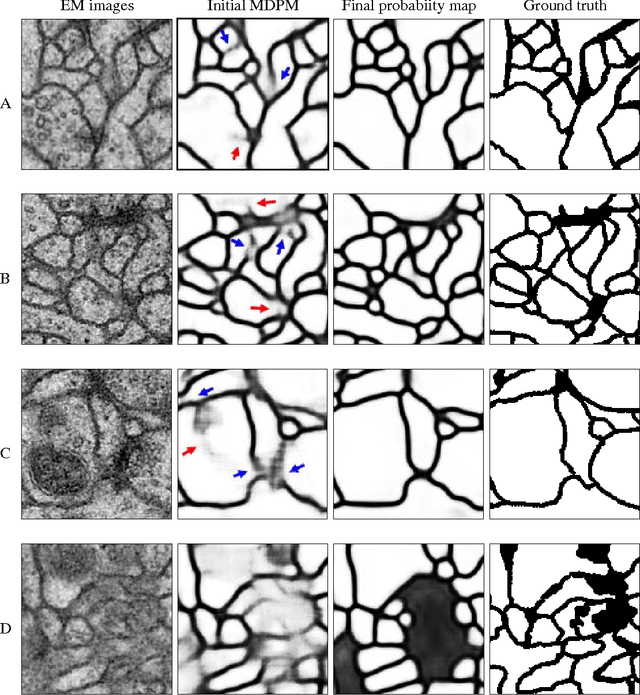
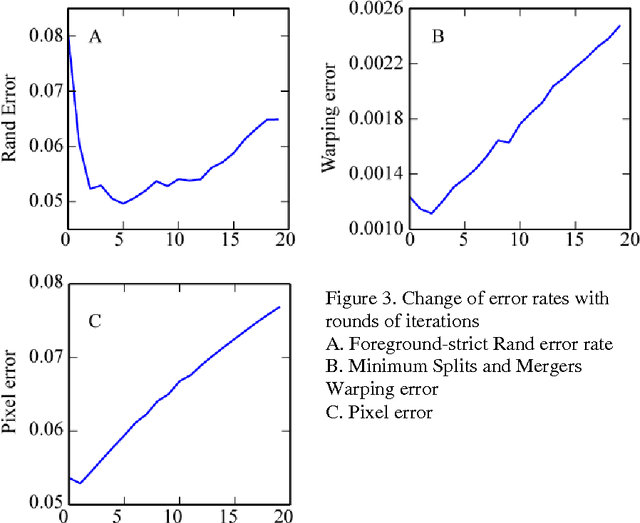
Abstract:To build the connectomics map of the brain, we developed a new algorithm that can automatically refine the Membrane Detection Probability Maps (MDPM) generated to perform automatic segmentation of electron microscopy (EM) images. To achieve this, we executed supervised training of a convolutional neural network to recover the removed center pixel label of patches sampled from a MDPM. MDPM can be generated from other machine learning based algorithms recognizing whether a pixel in an image corresponds to the cell membrane. By iteratively applying this network over MDPM for multiple rounds, we were able to significantly improve membrane segmentation results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge