Taebum Lee
Generalizing AI-driven Assessment of Immunohistochemistry across Immunostains and Cancer Types: A Universal Immunohistochemistry Analyzer
Jul 30, 2024Abstract:Despite advancements in methodologies, immunohistochemistry (IHC) remains the most utilized ancillary test for histopathologic and companion diagnostics in targeted therapies. However, objective IHC assessment poses challenges. Artificial intelligence (AI) has emerged as a potential solution, yet its development requires extensive training for each cancer and IHC type, limiting versatility. We developed a Universal IHC (UIHC) analyzer, an AI model for interpreting IHC images regardless of tumor or IHC types, using training datasets from various cancers stained for PD-L1 and/or HER2. This multi-cohort trained model outperforms conventional single-cohort models in interpreting unseen IHCs (Kappa score 0.578 vs. up to 0.509) and consistently shows superior performance across different positive staining cutoff values. Qualitative analysis reveals that UIHC effectively clusters patches based on expression levels. The UIHC model also quantitatively assesses c-MET expression with MET mutations, representing a significant advancement in AI application in the era of personalized medicine and accumulating novel biomarkers.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
MF-Hovernet: An Extension of Hovernet for Colon Nuclei Identification and Counting Challenge
Mar 04, 2022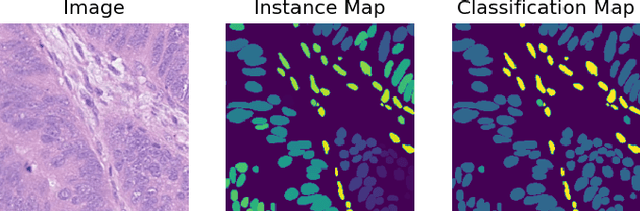
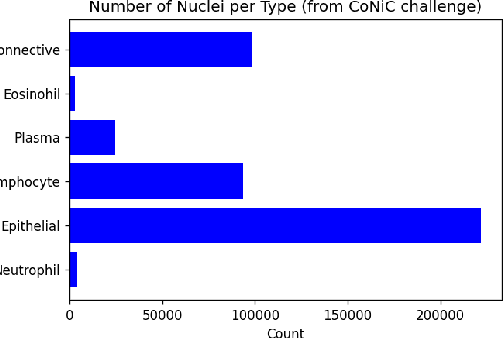
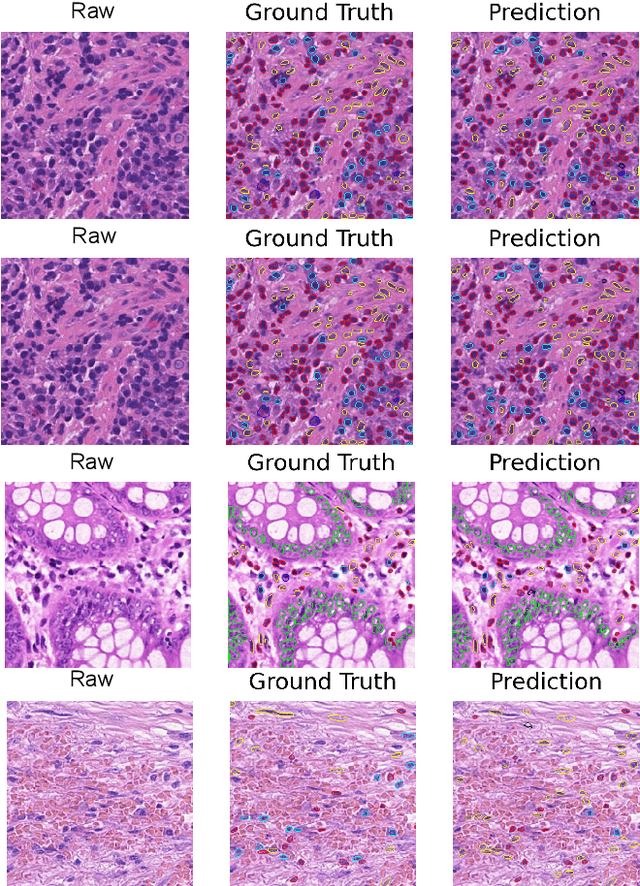
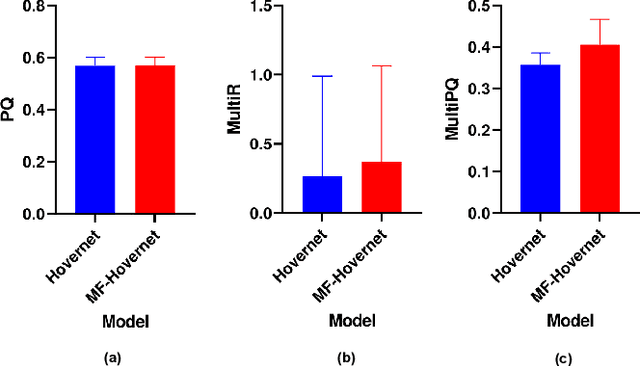
Abstract:Nuclei Identification and Counting is the most important morphological feature of cancers, especially in the colon. Many deep learning-based methods have been proposed to deal with this problem. In this work, we construct an extension of Hovernet for nuclei identification and counting to address the problem named MF-Hovernet. Our proposed model is the combination of multiple filer block to Hovernet architecture. The current result shows the efficiency of multiple filter block to improve the performance of the original Hovernet model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge