Sheng Xu
Representation Disparity-aware Distillation for 3D Object Detection
Aug 20, 2023



Abstract:In this paper, we focus on developing knowledge distillation (KD) for compact 3D detectors. We observe that off-the-shelf KD methods manifest their efficacy only when the teacher model and student counterpart share similar intermediate feature representations. This might explain why they are less effective in building extreme-compact 3D detectors where significant representation disparity arises due primarily to the intrinsic sparsity and irregularity in 3D point clouds. This paper presents a novel representation disparity-aware distillation (RDD) method to address the representation disparity issue and reduce performance gap between compact students and over-parameterized teachers. This is accomplished by building our RDD from an innovative perspective of information bottleneck (IB), which can effectively minimize the disparity of proposal region pairs from student and teacher in features and logits. Extensive experiments are performed to demonstrate the superiority of our RDD over existing KD methods. For example, our RDD increases mAP of CP-Voxel-S to 57.1% on nuScenes dataset, which even surpasses teacher performance while taking up only 42% FLOPs.
DCP-NAS: Discrepant Child-Parent Neural Architecture Search for 1-bit CNNs
Jun 27, 2023Abstract:Neural architecture search (NAS) proves to be among the effective approaches for many tasks by generating an application-adaptive neural architecture, which is still challenged by high computational cost and memory consumption. At the same time, 1-bit convolutional neural networks (CNNs) with binary weights and activations show their potential for resource-limited embedded devices. One natural approach is to use 1-bit CNNs to reduce the computation and memory cost of NAS by taking advantage of the strengths of each in a unified framework, while searching the 1-bit CNNs is more challenging due to the more complicated processes involved. In this paper, we introduce Discrepant Child-Parent Neural Architecture Search (DCP-NAS) to efficiently search 1-bit CNNs, based on a new framework of searching the 1-bit model (Child) under the supervision of a real-valued model (Parent). Particularly, we first utilize a Parent model to calculate a tangent direction, based on which the tangent propagation method is introduced to search the optimized 1-bit Child. We further observe a coupling relationship between the weights and architecture parameters existing in such differentiable frameworks. To address the issue, we propose a decoupled optimization method to search an optimized architecture. Extensive experiments demonstrate that our DCP-NAS achieves much better results than prior arts on both CIFAR-10 and ImageNet datasets. In particular, the backbones achieved by our DCP-NAS achieve strong generalization performance on person re-identification and object detection.
Bi-ViT: Pushing the Limit of Vision Transformer Quantization
May 21, 2023



Abstract:Vision transformers (ViTs) quantization offers a promising prospect to facilitate deploying large pre-trained networks on resource-limited devices. Fully-binarized ViTs (Bi-ViT) that pushes the quantization of ViTs to its limit remain largely unexplored and a very challenging task yet, due to their unacceptable performance. Through extensive empirical analyses, we identify the severe drop in ViT binarization is caused by attention distortion in self-attention, which technically stems from the gradient vanishing and ranking disorder. To address these issues, we first introduce a learnable scaling factor to reactivate the vanished gradients and illustrate its effectiveness through theoretical and experimental analyses. We then propose a ranking-aware distillation method to rectify the disordered ranking in a teacher-student framework. Bi-ViT achieves significant improvements over popular DeiT and Swin backbones in terms of Top-1 accuracy and FLOPs. For example, with DeiT-Tiny and Swin-Tiny, our method significantly outperforms baselines by 22.1% and 21.4% respectively, while 61.5x and 56.1x theoretical acceleration in terms of FLOPs compared with real-valued counterparts on ImageNet.
AF2-Mutation: Adversarial Sequence Mutations against AlphaFold2 on Protein Tertiary Structure Prediction
May 15, 2023Abstract:Deep learning-based approaches, such as AlphaFold2 (AF2), have significantly advanced protein tertiary structure prediction, achieving results comparable to real biological experimental methods. While AF2 has shown limitations in predicting the effects of mutations, its robustness against sequence mutations remains to be determined. Starting with the wild-type (WT) sequence, we investigate adversarial sequences generated via an evolutionary approach, which AF2 predicts to be substantially different from WT. Our experiments on CASP14 reveal that by modifying merely three residues in the protein sequence using a combination of replacement, deletion, and insertion strategies, the alteration in AF2's predictions, as measured by the Local Distance Difference Test (lDDT), reaches 46.61. Moreover, when applied to a specific protein, SPNS2, our proposed algorithm successfully identifies biologically meaningful residues critical to protein structure determination and potentially indicates alternative conformations, thus significantly expediting the experimental process.
Q-DETR: An Efficient Low-Bit Quantized Detection Transformer
Apr 01, 2023



Abstract:The recent detection transformer (DETR) has advanced object detection, but its application on resource-constrained devices requires massive computation and memory resources. Quantization stands out as a solution by representing the network in low-bit parameters and operations. However, there is a significant performance drop when performing low-bit quantized DETR (Q-DETR) with existing quantization methods. We find that the bottlenecks of Q-DETR come from the query information distortion through our empirical analyses. This paper addresses this problem based on a distribution rectification distillation (DRD). We formulate our DRD as a bi-level optimization problem, which can be derived by generalizing the information bottleneck (IB) principle to the learning of Q-DETR. At the inner level, we conduct a distribution alignment for the queries to maximize the self-information entropy. At the upper level, we introduce a new foreground-aware query matching scheme to effectively transfer the teacher information to distillation-desired features to minimize the conditional information entropy. Extensive experimental results show that our method performs much better than prior arts. For example, the 4-bit Q-DETR can theoretically accelerate DETR with ResNet-50 backbone by 6.6x and achieve 39.4% AP, with only 2.6% performance gaps than its real-valued counterpart on the COCO dataset.
Implicit Diffusion Models for Continuous Super-Resolution
Mar 29, 2023



Abstract:Image super-resolution (SR) has attracted increasing attention due to its wide applications. However, current SR methods generally suffer from over-smoothing and artifacts, and most work only with fixed magnifications. This paper introduces an Implicit Diffusion Model (IDM) for high-fidelity continuous image super-resolution. IDM integrates an implicit neural representation and a denoising diffusion model in a unified end-to-end framework, where the implicit neural representation is adopted in the decoding process to learn continuous-resolution representation. Furthermore, we design a scale-controllable conditioning mechanism that consists of a low-resolution (LR) conditioning network and a scaling factor. The scaling factor regulates the resolution and accordingly modulates the proportion of the LR information and generated features in the final output, which enables the model to accommodate the continuous-resolution requirement. Extensive experiments validate the effectiveness of our IDM and demonstrate its superior performance over prior arts.
Fossil Image Identification using Deep Learning Ensembles of Data Augmented Multiviews
Feb 16, 2023Abstract:Identification of fossil species is crucial to evolutionary studies. Recent advances from deep learning have shown promising prospects in fossil image identification. However, the quantity and quality of labeled fossil images are often limited due to fossil preservation, conditioned sampling, and expensive and inconsistent label annotation by domain experts, which pose great challenges to the training of deep learning based image classification models. To address these challenges, we follow the idea of the wisdom of crowds and propose a novel multiview ensemble framework, which collects multiple views of each fossil specimen image reflecting its different characteristics to train multiple base deep learning models and then makes final decisions via soft voting. We further develop OGS method that integrates original, gray, and skeleton views under this framework to demonstrate the effectiveness. Experimental results on the fusulinid fossil dataset over five deep learning based milestone models show that OGS using three base models consistently outperforms the baseline using a single base model, and the ablation study verifies the usefulness of each selected view. Besides, OGS obtains the superior or comparable performance compared to the method under well-known bagging framework. Moreover, as the available training data decreases, the proposed framework achieves more performance gains compared to the baseline. Furthermore, a consistency test with two human experts shows that OGS obtains the highest agreement with both the labels of dataset and the two experts. Notably, this methodology is designed for general fossil identification and it is expected to see applications on other fossil datasets. The results suggest the potential application when the quantity and quality of labeled data are particularly restricted, e.g., to identify rare fossil images.
Resilient Binary Neural Network
Feb 05, 2023



Abstract:Binary neural networks (BNNs) have received ever-increasing popularity for their great capability of reducing storage burden as well as quickening inference time. However, there is a severe performance drop compared with real-valued networks, due to its intrinsic frequent weight oscillation during training. In this paper, we introduce a Resilient Binary Neural Network (ReBNN) to mitigate the frequent oscillation for better BNNs' training. We identify that the weight oscillation mainly stems from the non-parametric scaling factor. To address this issue, we propose to parameterize the scaling factor and introduce a weighted reconstruction loss to build an adaptive training objective. For the first time, we show that the weight oscillation is controlled by the balanced parameter attached to the reconstruction loss, which provides a theoretical foundation to parameterize it in back propagation. Based on this, we learn our ReBNN by calculating the balanced parameter based on its maximum magnitude, which can effectively mitigate the weight oscillation with a resilient training process. Extensive experiments are conducted upon various network models, such as ResNet and Faster-RCNN for computer vision, as well as BERT for natural language processing. The results demonstrate the overwhelming performance of our ReBNN over prior arts. For example, our ReBNN achieves 66.9% Top-1 accuracy with ResNet-18 backbone on the ImageNet dataset, surpassing existing state-of-the-arts by a significant margin. Our code is open-sourced at https://github.com/SteveTsui/ReBNN.
Q-ViT: Accurate and Fully Quantized Low-bit Vision Transformer
Oct 13, 2022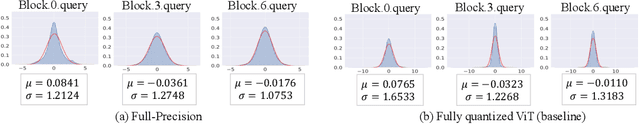
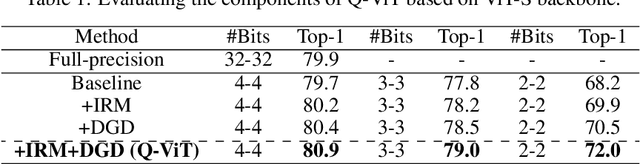
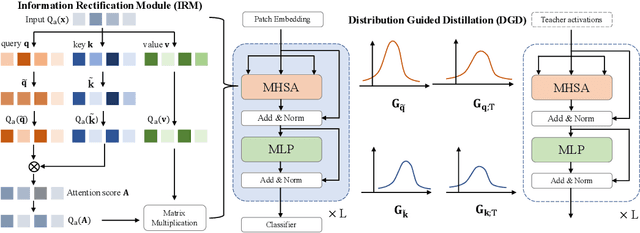
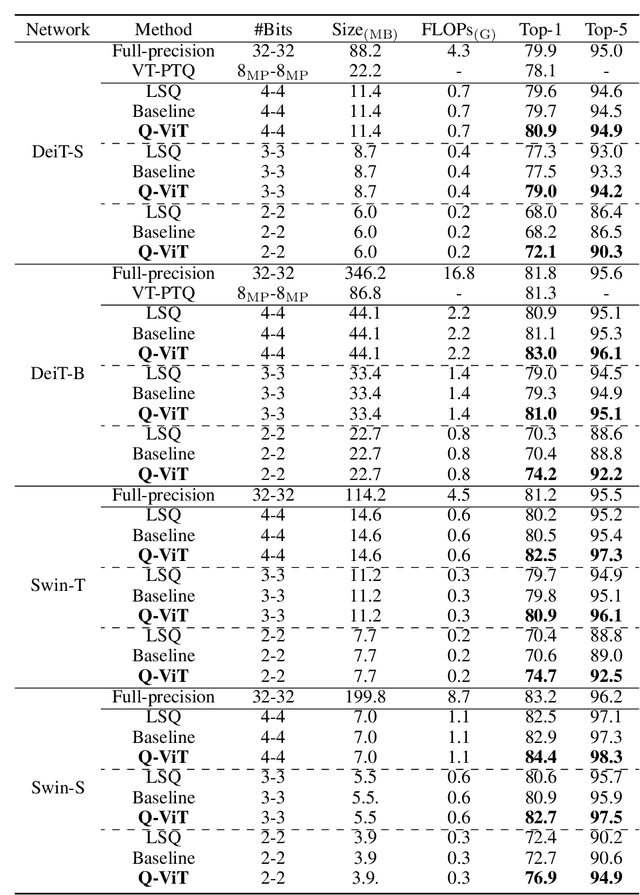
Abstract:The large pre-trained vision transformers (ViTs) have demonstrated remarkable performance on various visual tasks, but suffer from expensive computational and memory cost problems when deployed on resource-constrained devices. Among the powerful compression approaches, quantization extremely reduces the computation and memory consumption by low-bit parameters and bit-wise operations. However, low-bit ViTs remain largely unexplored and usually suffer from a significant performance drop compared with the real-valued counterparts. In this work, through extensive empirical analysis, we first identify the bottleneck for severe performance drop comes from the information distortion of the low-bit quantized self-attention map. We then develop an information rectification module (IRM) and a distribution guided distillation (DGD) scheme for fully quantized vision transformers (Q-ViT) to effectively eliminate such distortion, leading to a fully quantized ViTs. We evaluate our methods on popular DeiT and Swin backbones. Extensive experimental results show that our method achieves a much better performance than the prior arts. For example, our Q-ViT can theoretically accelerates the ViT-S by 6.14x and achieves about 80.9% Top-1 accuracy, even surpassing the full-precision counterpart by 1.0% on ImageNet dataset. Our codes and models are attached on https://github.com/YanjingLi0202/Q-ViT
Distance Map Supervised Landmark Localization for MR-TRUS Registration
Oct 11, 2022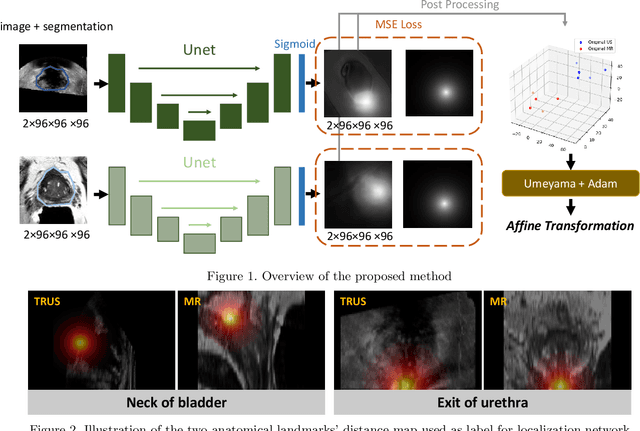
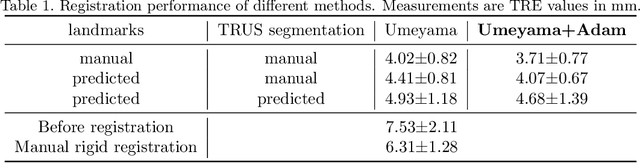

Abstract:In this work, we propose to explicitly use the landmarks of prostate to guide the MR-TRUS image registration. We first train a deep neural network to automatically localize a set of meaningful landmarks, and then directly generate the affine registration matrix from the location of these landmarks. For landmark localization, instead of directly training a network to predict the landmark coordinates, we propose to regress a full-resolution distance map of the landmark, which is demonstrated effective in avoiding statistical bias to unsatisfactory performance and thus improving performance. We then use the predicted landmarks to generate the affine transformation matrix, which outperforms the clinicians' manual rigid registration by a significant margin in terms of TRE.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge