Qingyu Zhao
Dept. of Psychiatry and Behavioral Sciences, Stanford University, Stanford, CA, USA
Generating Realistic 3D Brain MRIs Using a Conditional Diffusion Probabilistic Model
Dec 15, 2022Abstract:Training deep learning models on brain MRI is often plagued by small sample size, which can lead to biased training or overfitting. One potential solution is to synthetically generate realistic MRIs via generative models such as Generative Adversarial Network (GAN). However, existing GANs for synthesizing realistic brain MRIs largely rely on image-to-image conditioned transformations requiring extensive, well-curated pairs of MRI samples for training. On the other hand, unconditioned GAN models (i.e., those generating MRI from random noise) are unstable during training and tend to produce blurred images during inference. Here, we propose an efficient strategy that generates high fidelity 3D brain MRI via Diffusion Probabilistic Model (DPM). To this end, we train a conditional DPM with attention to generate an MRI sub-volume (a set of slices at arbitrary locations) conditioned on another subset of slices from the same MRI. By computing attention weights from slice indices and using a mask to encode the target and conditional slices, the model is able to learn the long-range dependency across distant slices with limited computational resources. After training, the model can progressively synthesize a new 3D brain MRI by generating the first subset of slices from random noise and conditionally generating subsequent slices. Based on 1262 t1-weighted MRIs from three neuroimaging studies, our experiments demonstrate that the proposed method can generate high quality 3D MRIs that share the same distribution as real MRIs and are more realistic than the ones produced by GAN-based models.
Joint Graph Convolution for Analyzing Brain Structural and Functional Connectome
Oct 27, 2022Abstract:The white-matter (micro-)structural architecture of the brain promotes synchrony among neuronal populations, giving rise to richly patterned functional connections. A fundamental problem for systems neuroscience is determining the best way to relate structural and functional networks quantified by diffusion tensor imaging and resting-state functional MRI. As one of the state-of-the-art approaches for network analysis, graph convolutional networks (GCN) have been separately used to analyze functional and structural networks, but have not been applied to explore inter-network relationships. In this work, we propose to couple the two networks of an individual by adding inter-network edges between corresponding brain regions, so that the joint structure-function graph can be directly analyzed by a single GCN. The weights of inter-network edges are learnable, reflecting non-uniform structure-function coupling strength across the brain. We apply our Joint-GCN to predict age and sex of 662 participants from the public dataset of the National Consortium on Alcohol and Neurodevelopment in Adolescence (NCANDA) based on their functional and micro-structural white-matter networks. Our results support that the proposed Joint-GCN outperforms existing multi-modal graph learning approaches for analyzing structural and functional networks.
Identifying Auxiliary or Adversarial Tasks Using Necessary Condition Analysis for Adversarial Multi-task Video Understanding
Aug 22, 2022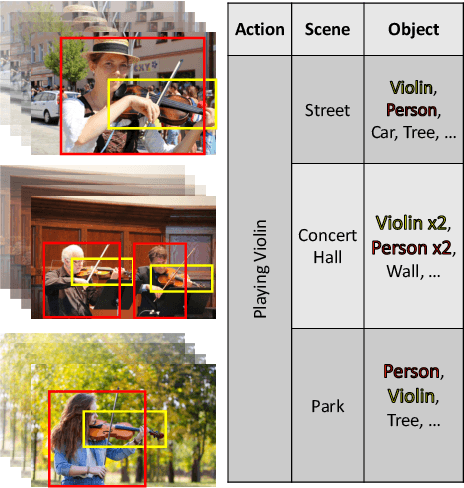
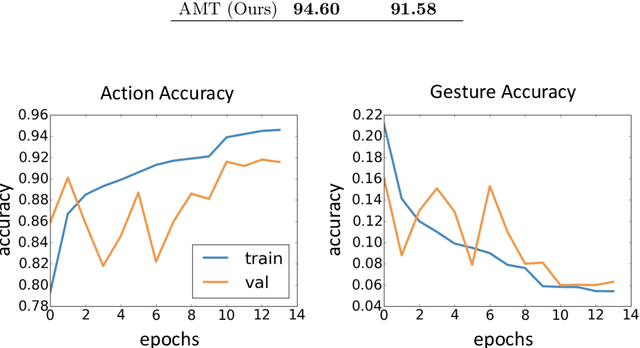
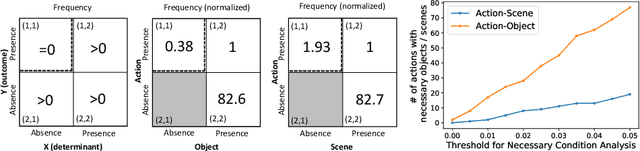
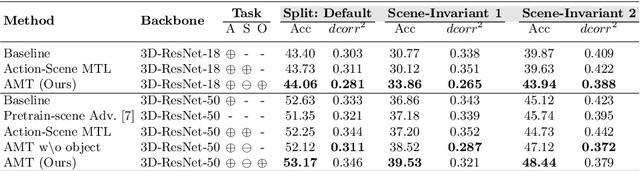
Abstract:There has been an increasing interest in multi-task learning for video understanding in recent years. In this work, we propose a generalized notion of multi-task learning by incorporating both auxiliary tasks that the model should perform well on and adversarial tasks that the model should not perform well on. We employ Necessary Condition Analysis (NCA) as a data-driven approach for deciding what category these tasks should fall in. Our novel proposed framework, Adversarial Multi-Task Neural Networks (AMT), penalizes adversarial tasks, determined by NCA to be scene recognition in the Holistic Video Understanding (HVU) dataset, to improve action recognition. This upends the common assumption that the model should always be encouraged to do well on all tasks in multi-task learning. Simultaneously, AMT still retains all the benefits of multi-task learning as a generalization of existing methods and uses object recognition as an auxiliary task to aid action recognition. We introduce two challenging Scene-Invariant test splits of HVU, where the model is evaluated on action-scene co-occurrences not encountered in training. We show that our approach improves accuracy by ~3% and encourages the model to attend to action features instead of correlation-biasing scene features.
Multiple Instance Neuroimage Transformer
Aug 19, 2022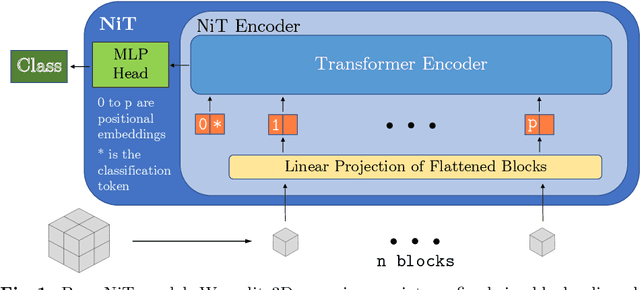
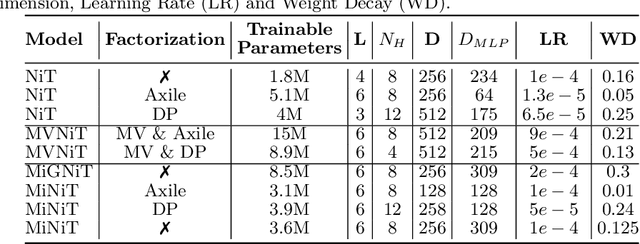
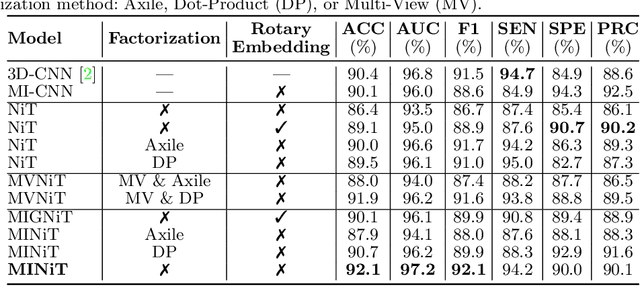
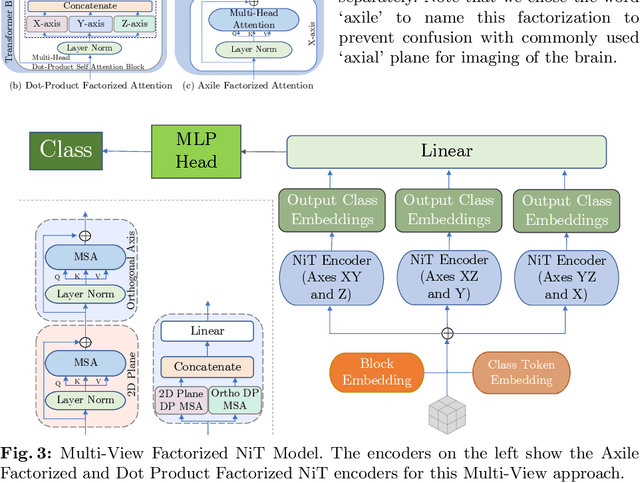
Abstract:For the first time, we propose using a multiple instance learning based convolution-free transformer model, called Multiple Instance Neuroimage Transformer (MINiT), for the classification of T1weighted (T1w) MRIs. We first present several variants of transformer models adopted for neuroimages. These models extract non-overlapping 3D blocks from the input volume and perform multi-headed self-attention on a sequence of their linear projections. MINiT, on the other hand, treats each of the non-overlapping 3D blocks of the input MRI as its own instance, splitting it further into non-overlapping 3D patches, on which multi-headed self-attention is computed. As a proof-of-concept, we evaluate the efficacy of our model by training it to identify sex from T1w-MRIs of two public datasets: Adolescent Brain Cognitive Development (ABCD) and the National Consortium on Alcohol and Neurodevelopment in Adolescence (NCANDA). The learned attention maps highlight voxels contributing to identifying sex differences in brain morphometry. The code is available at https://github.com/singlaayush/MINIT.
Bridging the Gap between Deep Learning and Hypothesis-Driven Analysis via Permutation Testing
Jul 28, 2022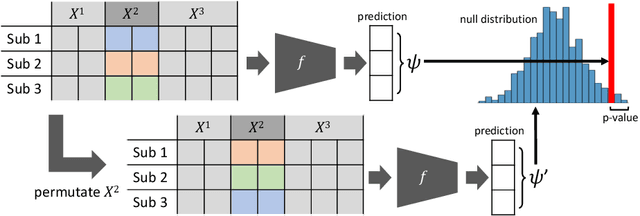
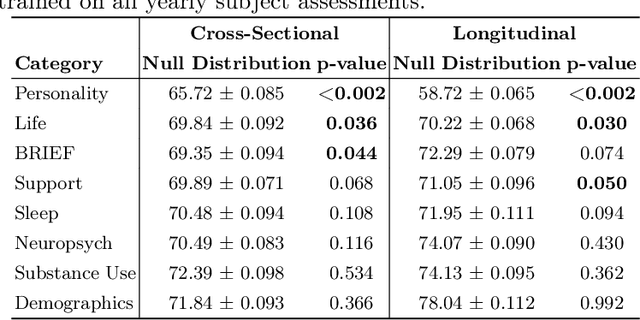

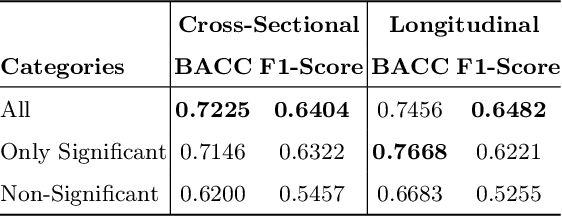
Abstract:A fundamental approach in neuroscience research is to test hypotheses based on neuropsychological and behavioral measures, i.e., whether certain factors (e.g., related to life events) are associated with an outcome (e.g., depression). In recent years, deep learning has become a potential alternative approach for conducting such analyses by predicting an outcome from a collection of factors and identifying the most "informative" ones driving the prediction. However, this approach has had limited impact as its findings are not linked to statistical significance of factors supporting hypotheses. In this article, we proposed a flexible and scalable approach based on the concept of permutation testing that integrates hypothesis testing into the data-driven deep learning analysis. We apply our approach to the yearly self-reported assessments of 621 adolescent participants of the National Consortium of Alcohol and Neurodevelopment in Adolescence (NCANDA) to predict negative valence, a symptom of major depressive disorder according to the NIMH Research Domain Criteria (RDoC). Our method successfully identifies categories of risk factors that further explain the symptom.
A Penalty Approach for Normalizing Feature Distributions to Build Confounder-Free Models
Jul 11, 2022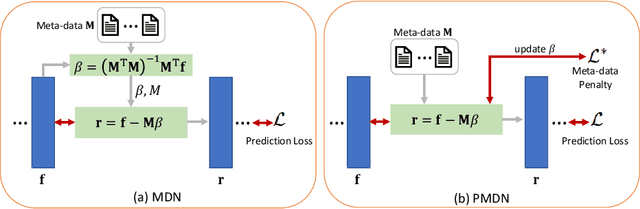

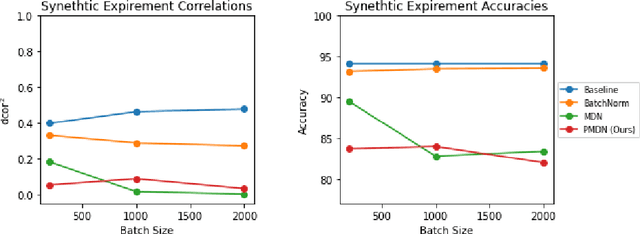
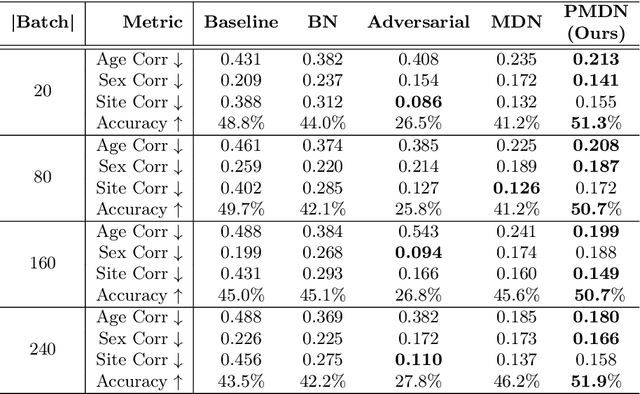
Abstract:Translating machine learning algorithms into clinical applications requires addressing challenges related to interpretability, such as accounting for the effect of confounding variables (or metadata). Confounding variables affect the relationship between input training data and target outputs. When we train a model on such data, confounding variables will bias the distribution of the learned features. A recent promising solution, MetaData Normalization (MDN), estimates the linear relationship between the metadata and each feature based on a non-trainable closed-form solution. However, this estimation is confined by the sample size of a mini-batch and thereby may cause the approach to be unstable during training. In this paper, we extend the MDN method by applying a Penalty approach (referred to as PDMN). We cast the problem into a bi-level nested optimization problem. We then approximate this optimization problem using a penalty method so that the linear parameters within the MDN layer are trainable and learned on all samples. This enables PMDN to be plugged into any architectures, even those unfit to run batch-level operations, such as transformers and recurrent models. We show improvement in model accuracy and greater independence from confounders using PMDN over MDN in a synthetic experiment and a multi-label, multi-site dataset of magnetic resonance images (MRIs).
Longitudinal Correlation Analysis for Decoding Multi-Modal Brain Development
Jul 10, 2021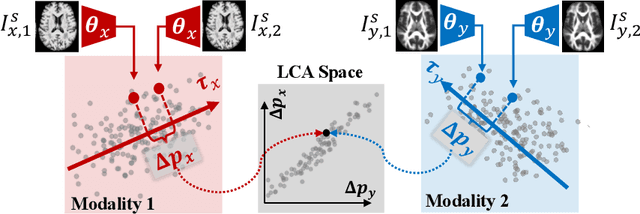
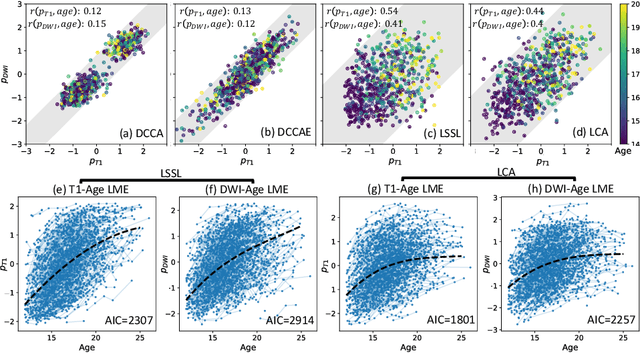
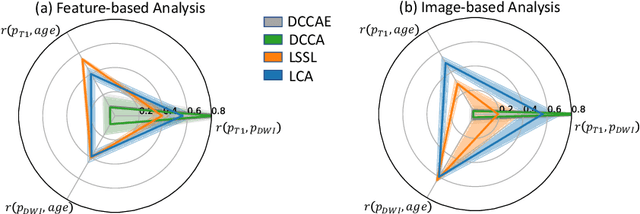
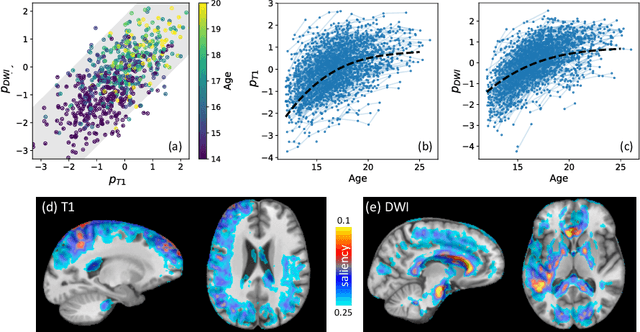
Abstract:Starting from childhood, the human brain restructures and rewires throughout life. Characterizing such complex brain development requires effective analysis of longitudinal and multi-modal neuroimaging data. Here, we propose such an analysis approach named Longitudinal Correlation Analysis (LCA). LCA couples the data of two modalities by first reducing the input from each modality to a latent representation based on autoencoders. A self-supervised strategy then relates the two latent spaces by jointly disentangling two directions, one in each space, such that the longitudinal changes in latent representations along those directions are maximally correlated between modalities. We applied LCA to analyze the longitudinal T1-weighted and diffusion-weighted MRIs of 679 youths from the National Consortium on Alcohol and Neurodevelopment in Adolescence. Unlike existing approaches that focus on either cross-sectional or single-modal modeling, LCA successfully unraveled coupled macrostructural and microstructural brain development from morphological and diffusivity features extracted from the data. A retesting of LCA on raw 3D image volumes of those subjects successfully replicated the findings from the feature-based analysis. Lastly, the developmental effects revealed by LCA were inline with the current understanding of maturational patterns of the adolescent brain.
Metadata Normalization
May 05, 2021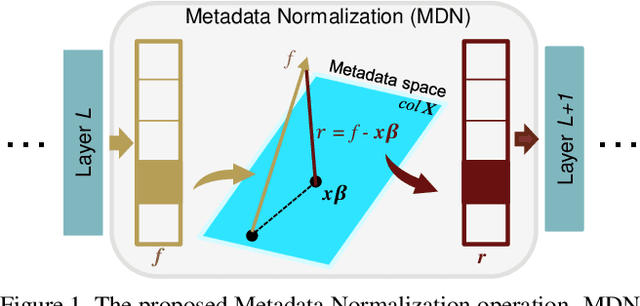
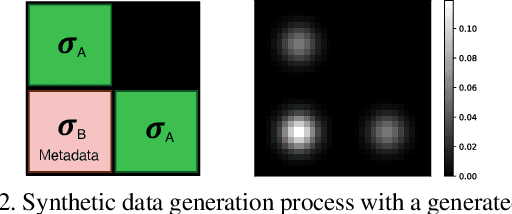
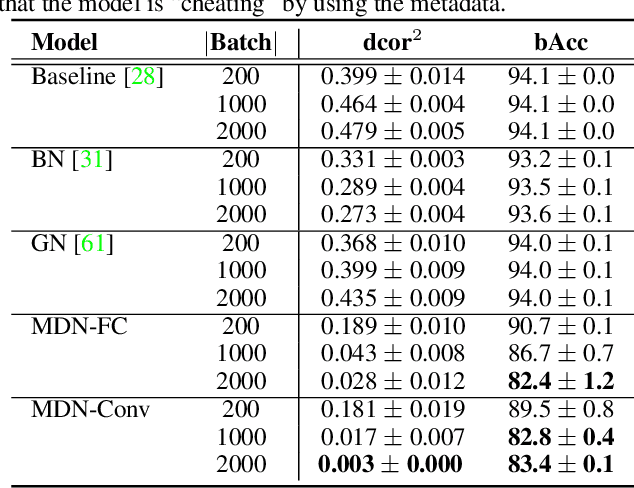

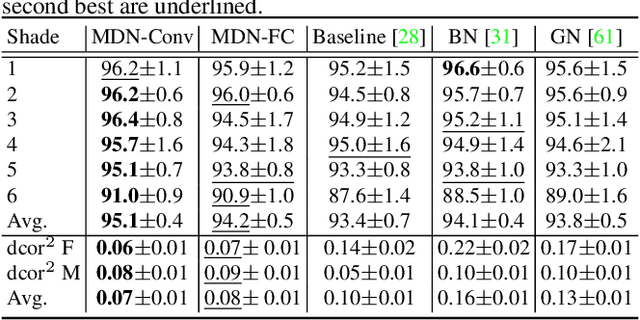
Abstract:Batch Normalization (BN) and its variants have delivered tremendous success in combating the covariate shift induced by the training step of deep learning methods. While these techniques normalize feature distributions by standardizing with batch statistics, they do not correct the influence on features from extraneous variables or multiple distributions. Such extra variables, referred to as metadata here, may create bias or confounding effects (e.g., race when classifying gender from face images). We introduce the Metadata Normalization (MDN) layer, a new batch-level operation which can be used end-to-end within the training framework, to correct the influence of metadata on feature distributions. MDN adopts a regression analysis technique traditionally used for preprocessing to remove (regress out) the metadata effects on model features during training. We utilize a metric based on distance correlation to quantify the distribution bias from the metadata and demonstrate that our method successfully removes metadata effects on four diverse settings: one synthetic, one 2D image, one video, and one 3D medical image dataset.
Self-Supervised Longitudinal Neighbourhood Embedding
Mar 09, 2021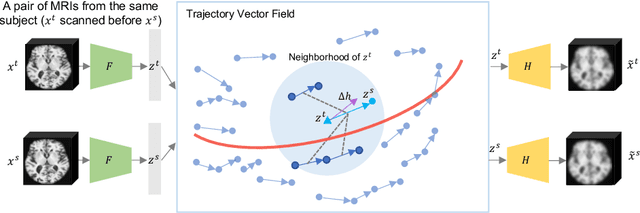
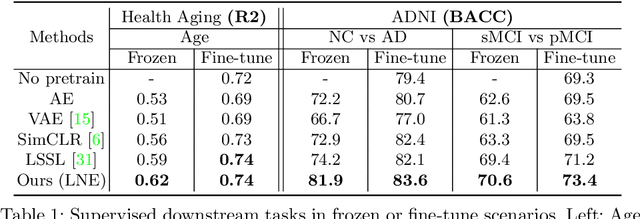
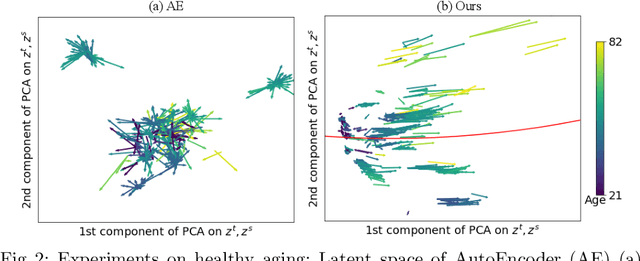
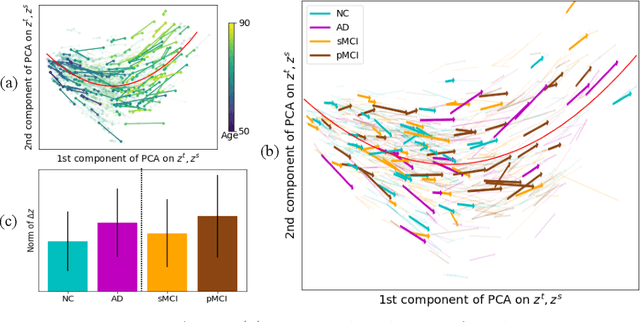
Abstract:Longitudinal MRIs are often used to capture the gradual deterioration of brain structure and function caused by aging or neurological diseases. Analyzing this data via machine learning generally requires a large number of ground-truth labels, which are often missing or expensive to obtain. Reducing the need for labels, we propose a self-supervised strategy for representation learning named Longitudinal Neighborhood Embedding (LNE). Motivated by concepts in contrastive learning, LNE explicitly models the similarity between trajectory vectors across different subjects. We do so by building a graph in each training iteration defining neighborhoods in the latent space so that the progression direction of a subject follows the direction of its neighbors. This results in a smooth trajectory field that captures the global morphological change of the brain while maintaining the local continuity. We apply LNE to longitudinal T1w MRIs of two neuroimaging studies: a dataset composed of 274 healthy subjects, and Alzheimer's Disease Neuroimaging Initiative (ADNI, N=632). The visualization of the smooth trajectory vector field and superior performance on downstream tasks demonstrate the strength of the proposed method over existing self-supervised methods in extracting information associated with normal aging and in revealing the impact of neurodegenerative disorders. The code is available at \url{https://github.com/ouyangjiahong/longitudinal-neighbourhood-embedding.git}.
Representation Disentanglement for Multi-modal MR Analysis
Feb 23, 2021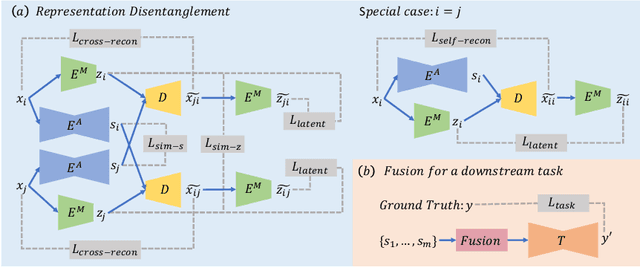
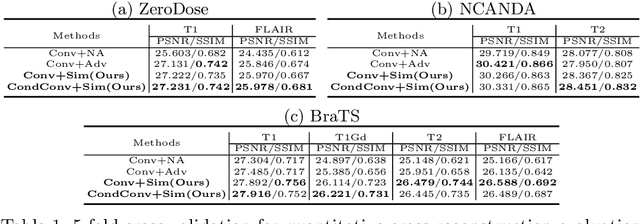
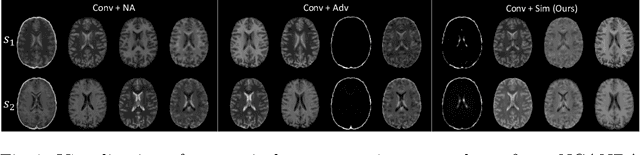
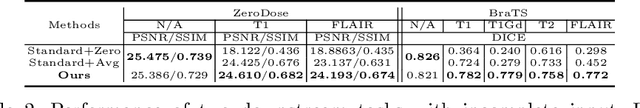
Abstract:Multi-modal MR images are widely used in neuroimaging applications to provide complementary information about the brain structures. Recent works have suggested that multi-modal deep learning analysis can benefit from explicitly disentangling anatomical (shape) and modality (appearance) representations from the images. In this work, we challenge existing strategies by showing that they do not naturally lead to representation disentanglement both in theory and in practice. To address this issue, we propose a margin loss that regularizes the similarity relationships of the representations across subjects and modalities. To enable a robust training, we further introduce a modified conditional convolution to design a single model for encoding images of all modalities. Lastly, we propose a fusion function to combine the disentangled anatomical representations as a set of modality-invariant features for downstream tasks. We evaluate the proposed method on three multi-modal neuroimaging datasets. Experiments show that our proposed method can achieve superior disentangled representations compared to existing disentanglement strategies. Results also indicate that the fused anatomical representation has great potential in the downstream task of zero-dose PET reconstruction and brain tumor segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge