Peter J. Schüffler
MMNavAgent: Multi-Magnification WSI Navigation Agent for Clinically Consistent Whole-Slide Analysis
Mar 02, 2026Abstract:Recent AI navigation approaches aim to improve Whole-Slide Image (WSI) diagnosis by modeling spatial exploration and selecting diagnostically relevant regions, yet most operate at a single fixed magnification or rely on predefined magnification traversal. In clinical practice, pathologists examine slides across multiple magnifications and selectively inspect only necessary scales, dynamically integrating global and cellular evidence in a sequential manner. This mismatch prevents existing methods from modeling cross-magnification interactions and adaptive magnification selection inherent to real diagnostic workflows. To these, we propose a clinically consistent Multi-Magnification WSI Navigation Agent (MMNavAgent) that explicitly models multi magnification interaction and adaptive magnification selection. Specifically, we introduce a Cross-Magnification navigation Tool (CMT) that aggregates contextual information from adjacent magnifications to enhance discriminative representations along the navigation path. We further introduce a Magnification Selection Tool (MST) that leverages memory-driven reasoning within the agent framework to enable interactive and adaptive magnification selection, mimicking the sequential decision process of pathologists. Extensive experiments on a public dataset demonstrate improved diagnostic performance, with 1.45% gain of AUC and 2.93% gain of BACC over a non-agent baseline. Code will be public upon acceptance.
Efficient Special Stain Classification
Feb 10, 2026Abstract:Stains are essential in histopathology to visualize specific tissue characteristics, with Haematoxylin and Eosin (H&E) serving as the clinical standard. However, pathologists frequently utilize a variety of special stains for the diagnosis of specific morphologies. Maintaining accurate metadata for these slides is critical for quality control in clinical archives and for the integrity of computational pathology datasets. In this work, we compare two approaches for automated classification of stains using whole slide images, covering the 14 most commonly used special stains in our institute alongside standard and frozen-section H&E. We evaluate a Multi-Instance Learning (MIL) pipeline and a proposed lightweight thumbnail-based approach. On internal test data, MIL achieved the highest performance (macro F1: 0.941 for 16 classes; 0.969 for 14 merged classes), while the thumbnail approach remained competitive (0.897 and 0.953, respectively). On external TCGA data, the thumbnail model generalized best (weighted F1: 0.843 vs. 0.807 for MIL). The thumbnail approach also increased throughput by two orders of magnitude (5.635 vs. 0.018 slides/s for MIL with all patches). We conclude that thumbnail-based classification provides a scalable and robust solution for routine visual quality control in digital pathology workflows.
Deep Learning-Based Fixation Type Prediction for Quality Assurance in Digital Pathology
Feb 09, 2026Abstract:Accurate annotation of fixation type is a critical step in slide preparation for pathology laboratories. However, this manual process is prone to errors, impacting downstream analyses and diagnostic accuracy. Existing methods for verifying formalin-fixed, paraffin-embedded (FFPE), and frozen section (FS) fixation types typically require full-resolution whole-slide images (WSIs), limiting scalability for high-throughput quality control. We propose a deep-learning model to predict fixation types using low-resolution, pre-scan thumbnail images. The model was trained on WSIs from the TUM Institute of Pathology (n=1,200, Leica GT450DX) and evaluated on a class-balanced subset of The Cancer Genome Atlas dataset (TCGA, n=8,800, Leica AT2), as well as on class-balanced datasets from Augsburg (n=695 [392 FFPE, 303 FS], Philips UFS) and Regensburg (n=202, 3DHISTECH P1000). Our model achieves an AUROC of 0.88 on TCGA, outperforming comparable pre-scan methods by 4.8%. It also achieves AUROCs of 0.72 on Regensburg and Augsburg slides, underscoring challenges related to scanner-induced domain shifts. Furthermore, the model processes each slide in 21 ms, $400\times$ faster than existing high-magnification, full-resolution methods, enabling rapid, high-throughput processing. This approach provides an efficient solution for detecting labelling errors without relying on high-magnification scans, offering a valuable tool for quality control in high-throughput pathology workflows. Future work will improve and evaluate the model's generalisation to additional scanner types. Our findings suggest that this method can increase accuracy and efficiency in digital pathology workflows and may be extended to other low-resolution slide annotations.
From Linear Probing to Joint-Weighted Token Hierarchy: A Foundation Model Bridging Global and Cellular Representations in Biomarker Detection
Nov 07, 2025Abstract:AI-based biomarkers can infer molecular features directly from hematoxylin & eosin (H&E) slides, yet most pathology foundation models (PFMs) rely on global patch-level embeddings and overlook cell-level morphology. We present a PFM model, JWTH (Joint-Weighted Token Hierarchy), which integrates large-scale self-supervised pretraining with cell-centric post-tuning and attention pooling to fuse local and global tokens. Across four tasks involving four biomarkers and eight cohorts, JWTH achieves up to 8.3% higher balanced accuracy and 1.2% average improvement over prior PFMs, advancing interpretable and robust AI-based biomarker detection in digital pathology.
Deep Learning Methods for Lung Cancer Segmentation in Whole-slide Histopathology Images -- the ACDC@LungHP Challenge 2019
Aug 21, 2020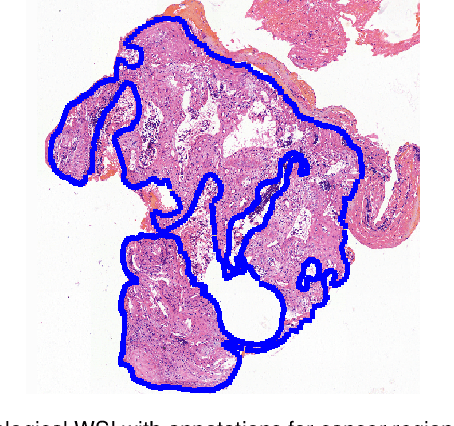
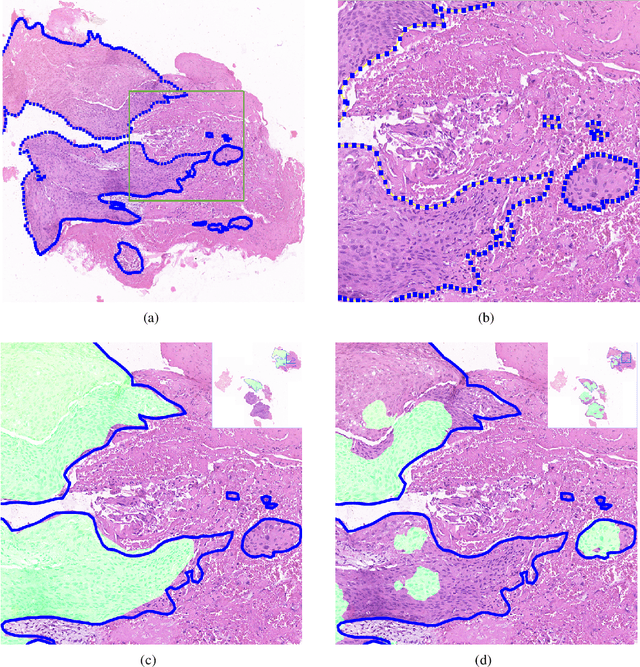
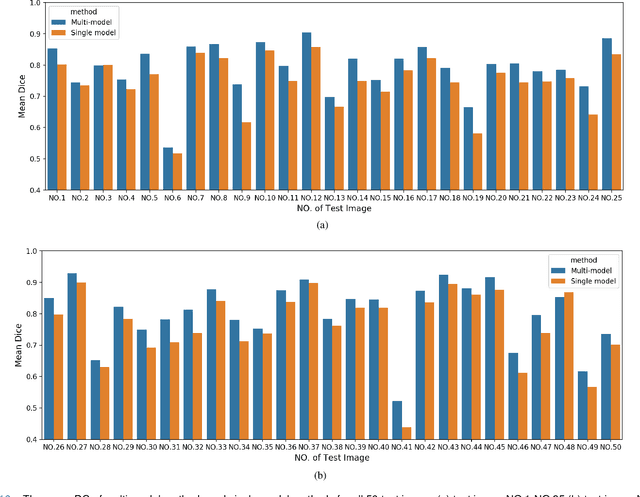
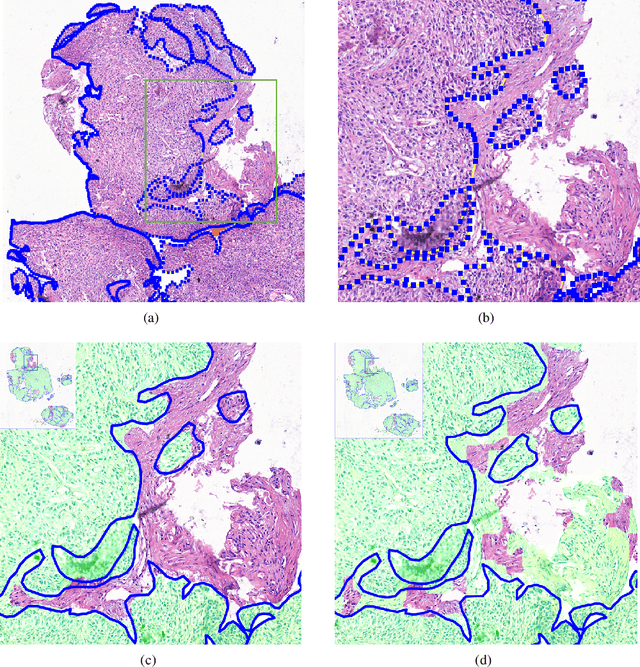

Abstract:Accurate segmentation of lung cancer in pathology slides is a critical step in improving patient care. We proposed the ACDC@LungHP (Automatic Cancer Detection and Classification in Whole-slide Lung Histopathology) challenge for evaluating different computer-aided diagnosis (CADs) methods on the automatic diagnosis of lung cancer. The ACDC@LungHP 2019 focused on segmentation (pixel-wise detection) of cancer tissue in whole slide imaging (WSI), using an annotated dataset of 150 training images and 50 test images from 200 patients. This paper reviews this challenge and summarizes the top 10 submitted methods for lung cancer segmentation. All methods were evaluated using the false positive rate, false negative rate, and DICE coefficient (DC). The DC ranged from 0.7354$\pm$0.1149 to 0.8372$\pm$0.0858. The DC of the best method was close to the inter-observer agreement (0.8398$\pm$0.0890). All methods were based on deep learning and categorized into two groups: multi-model method and single model method. In general, multi-model methods were significantly better ($\textit{p}$<$0.01$) than single model methods, with mean DC of 0.7966 and 0.7544, respectively. Deep learning based methods could potentially help pathologists find suspicious regions for further analysis of lung cancer in WSI.
Mitochondria-based Renal Cell Carcinoma Subtyping: Learning from Deep vs. Flat Feature Representations
Aug 02, 2016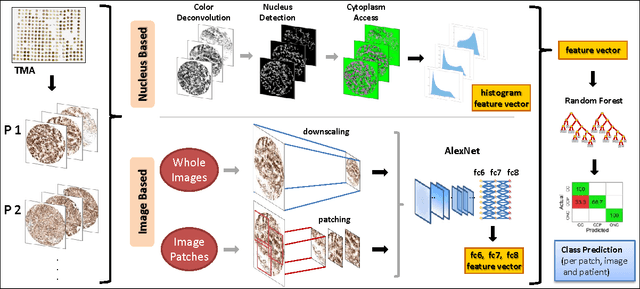
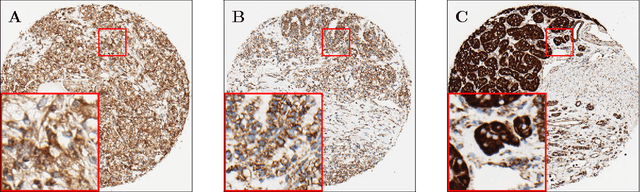
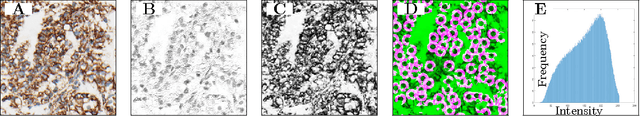

Abstract:Accurate subtyping of renal cell carcinoma (RCC) is of crucial importance for understanding disease progression and for making informed treatment decisions. New discoveries of significant alterations to mitochondria between subtypes make immunohistochemical (IHC) staining based image classification an imperative. Until now, accurate quantification and subtyping was made impossible by huge IHC variations, the absence of cell membrane staining for cytoplasm segmentation as well as the complete lack of systems for robust and reproducible image based classification. In this paper we present a comprehensive classification framework to overcome these challenges for tissue microarrays (TMA) of RCCs. We compare and evaluate models based on domain specific hand-crafted "flat"-features versus "deep" feature representations from various layers of a pre-trained convolutional neural network (CNN). The best model reaches a cross-validation accuracy of 89%, which demonstrates for the first time, that robust mitochondria-based subtyping of renal cancer is feasible
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge