Pallavi Tiwari
Department of Radiology, University of Wisconsin School of Medicine & Public Health, Madison, WI, USA, Department of Biomedical Engineering, University of Wisconsin Madison, Madison, WI, USA
Cyst-X: AI-Powered Pancreatic Cancer Risk Prediction from Multicenter MRI in Centralized and Federated Learning
Jul 29, 2025



Abstract:Pancreatic cancer is projected to become the second-deadliest malignancy in Western countries by 2030, highlighting the urgent need for better early detection. Intraductal papillary mucinous neoplasms (IPMNs), key precursors to pancreatic cancer, are challenging to assess with current guidelines, often leading to unnecessary surgeries or missed malignancies. We present Cyst-X, an AI framework that predicts IPMN malignancy using multicenter MRI data, leveraging MRI's superior soft tissue contrast over CT. Trained on 723 T1- and 738 T2-weighted scans from 764 patients across seven institutions, our models (AUC=0.82) significantly outperform both Kyoto guidelines (AUC=0.75) and expert radiologists. The AI-derived imaging features align with known clinical markers and offer biologically meaningful insights. We also demonstrate strong performance in a federated learning setting, enabling collaborative training without sharing patient data. To promote privacy-preserving AI development and improve IPMN risk stratification, the Cyst-X dataset is released as the first large-scale, multi-center pancreatic cysts MRI dataset.
3D Nephrographic Image Synthesis in CT Urography with the Diffusion Model and Swin Transformer
Feb 26, 2025Abstract:Purpose: This study aims to develop and validate a method for synthesizing 3D nephrographic phase images in CT urography (CTU) examinations using a diffusion model integrated with a Swin Transformer-based deep learning approach. Materials and Methods: This retrospective study was approved by the local Institutional Review Board. A dataset comprising 327 patients who underwent three-phase CTU (mean $\pm$ SD age, 63 $\pm$ 15 years; 174 males, 153 females) was curated for deep learning model development. The three phases for each patient were aligned with an affine registration algorithm. A custom deep learning model coined dsSNICT (diffusion model with a Swin transformer for synthetic nephrographic phase images in CT) was developed and implemented to synthesize the nephrographic images. Performance was assessed using Peak Signal-to-Noise Ratio (PSNR), Structural Similarity Index (SSIM), Mean Absolute Error (MAE), and Fr\'{e}chet Video Distance (FVD). Qualitative evaluation by two fellowship-trained abdominal radiologists was performed. Results: The synthetic nephrographic images generated by our proposed approach achieved high PSNR (26.3 $\pm$ 4.4 dB), SSIM (0.84 $\pm$ 0.069), MAE (12.74 $\pm$ 5.22 HU), and FVD (1323). Two radiologists provided average scores of 3.5 for real images and 3.4 for synthetic images (P-value = 0.5) on a Likert scale of 1-5, indicating that our synthetic images closely resemble real images. Conclusion: The proposed approach effectively synthesizes high-quality 3D nephrographic phase images. This model can be used to reduce radiation dose in CTU by 33.3\% without compromising image quality, which thereby enhances the safety and diagnostic utility of CT urography.
IPMN Risk Assessment under Federated Learning Paradigm
Nov 08, 2024



Abstract:Accurate classification of Intraductal Papillary Mucinous Neoplasms (IPMN) is essential for identifying high-risk cases that require timely intervention. In this study, we develop a federated learning framework for multi-center IPMN classification utilizing a comprehensive pancreas MRI dataset. This dataset includes 653 T1-weighted and 656 T2-weighted MRI images, accompanied by corresponding IPMN risk scores from 7 leading medical institutions, making it the largest and most diverse dataset for IPMN classification to date. We assess the performance of DenseNet-121 in both centralized and federated settings for training on distributed data. Our results demonstrate that the federated learning approach achieves high classification accuracy comparable to centralized learning while ensuring data privacy across institutions. This work marks a significant advancement in collaborative IPMN classification, facilitating secure and high-accuracy model training across multiple centers.
Adaptive Aggregation Weights for Federated Segmentation of Pancreas MRI
Oct 29, 2024

Abstract:Federated learning (FL) enables collaborative model training across institutions without sharing sensitive data, making it an attractive solution for medical imaging tasks. However, traditional FL methods, such as Federated Averaging (FedAvg), face difficulties in generalizing across domains due to variations in imaging protocols and patient demographics across institutions. This challenge is particularly evident in pancreas MRI segmentation, where anatomical variability and imaging artifacts significantly impact performance. In this paper, we conduct a comprehensive evaluation of FL algorithms for pancreas MRI segmentation and introduce a novel approach that incorporates adaptive aggregation weights. By dynamically adjusting the contribution of each client during model aggregation, our method accounts for domain-specific differences and improves generalization across heterogeneous datasets. Experimental results demonstrate that our approach enhances segmentation accuracy and reduces the impact of domain shift compared to conventional FL methods while maintaining privacy-preserving capabilities. Significant performance improvements are observed across multiple hospitals (centers).
Large-Scale Multi-Center CT and MRI Segmentation of Pancreas with Deep Learning
May 20, 2024



Abstract:Automated volumetric segmentation of the pancreas on cross-sectional imaging is needed for diagnosis and follow-up of pancreatic diseases. While CT-based pancreatic segmentation is more established, MRI-based segmentation methods are understudied, largely due to a lack of publicly available datasets, benchmarking research efforts, and domain-specific deep learning methods. In this retrospective study, we collected a large dataset (767 scans from 499 participants) of T1-weighted (T1W) and T2-weighted (T2W) abdominal MRI series from five centers between March 2004 and November 2022. We also collected CT scans of 1,350 patients from publicly available sources for benchmarking purposes. We developed a new pancreas segmentation method, called PanSegNet, combining the strengths of nnUNet and a Transformer network with a new linear attention module enabling volumetric computation. We tested PanSegNet's accuracy in cross-modality (a total of 2,117 scans) and cross-center settings with Dice and Hausdorff distance (HD95) evaluation metrics. We used Cohen's kappa statistics for intra and inter-rater agreement evaluation and paired t-tests for volume and Dice comparisons, respectively. For segmentation accuracy, we achieved Dice coefficients of 88.3% (std: 7.2%, at case level) with CT, 85.0% (std: 7.9%) with T1W MRI, and 86.3% (std: 6.4%) with T2W MRI. There was a high correlation for pancreas volume prediction with R^2 of 0.91, 0.84, and 0.85 for CT, T1W, and T2W, respectively. We found moderate inter-observer (0.624 and 0.638 for T1W and T2W MRI, respectively) and high intra-observer agreement scores. All MRI data is made available at https://osf.io/kysnj/. Our source code is available at https://github.com/NUBagciLab/PaNSegNet.
ResNCT: A Deep Learning Model for the Synthesis of Nephrographic Phase Images in CT Urography
May 07, 2024



Abstract:Purpose: To develop and evaluate a transformer-based deep learning model for the synthesis of nephrographic phase images in CT urography (CTU) examinations from the unenhanced and urographic phases. Materials and Methods: This retrospective study was approved by the local Institutional Review Board. A dataset of 119 patients (mean $\pm$ SD age, 65 $\pm$ 12 years; 75/44 males/females) with three-phase CT urography studies was curated for deep learning model development. The three phases for each patient were aligned with an affine registration algorithm. A custom model, coined Residual transformer model for Nephrographic phase CT image synthesis (ResNCT), was developed and implemented with paired inputs of non-contrast and urographic sets of images trained to produce the nephrographic phase images, that were compared with the corresponding ground truth nephrographic phase images. The synthesized images were evaluated with multiple performance metrics, including peak signal to noise ratio (PSNR), structural similarity index (SSIM), normalized cross correlation coefficient (NCC), mean absolute error (MAE), and root mean squared error (RMSE). Results: The ResNCT model successfully generated synthetic nephrographic images from non-contrast and urographic image inputs. With respect to ground truth nephrographic phase images, the images synthesized by the model achieved high PSNR (27.8 $\pm$ 2.7 dB), SSIM (0.88 $\pm$ 0.05), and NCC (0.98 $\pm$ 0.02), and low MAE (0.02 $\pm$ 0.005) and RMSE (0.042 $\pm$ 0.016). Conclusion: The ResNCT model synthesized nephrographic phase CT images with high similarity to ground truth images. The ResNCT model provides a means of eliminating the acquisition of the nephrographic phase with a resultant 33% reduction in radiation dose for CTU examinations.
Federated Learning Enables Big Data for Rare Cancer Boundary Detection
Apr 25, 2022Abstract:Although machine learning (ML) has shown promise in numerous domains, there are concerns about generalizability to out-of-sample data. This is currently addressed by centrally sharing ample, and importantly diverse, data from multiple sites. However, such centralization is challenging to scale (or even not feasible) due to various limitations. Federated ML (FL) provides an alternative to train accurate and generalizable ML models, by only sharing numerical model updates. Here we present findings from the largest FL study to-date, involving data from 71 healthcare institutions across 6 continents, to generate an automatic tumor boundary detector for the rare disease of glioblastoma, utilizing the largest dataset of such patients ever used in the literature (25,256 MRI scans from 6,314 patients). We demonstrate a 33% improvement over a publicly trained model to delineate the surgically targetable tumor, and 23% improvement over the tumor's entire extent. We anticipate our study to: 1) enable more studies in healthcare informed by large and diverse data, ensuring meaningful results for rare diseases and underrepresented populations, 2) facilitate further quantitative analyses for glioblastoma via performance optimization of our consensus model for eventual public release, and 3) demonstrate the effectiveness of FL at such scale and task complexity as a paradigm shift for multi-site collaborations, alleviating the need for data sharing.
Radiomic Deformation and Textural Heterogeneity (R-DepTH) Descriptor to characterize Tumor Field Effect: Application to Survival Prediction in Glioblastoma
Mar 12, 2021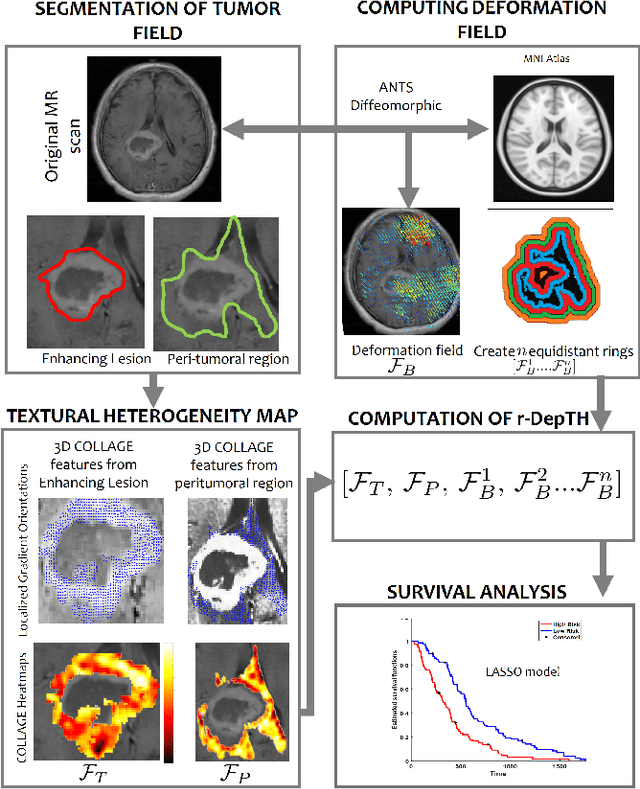
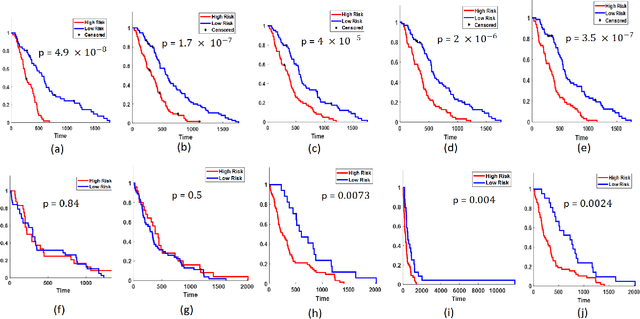
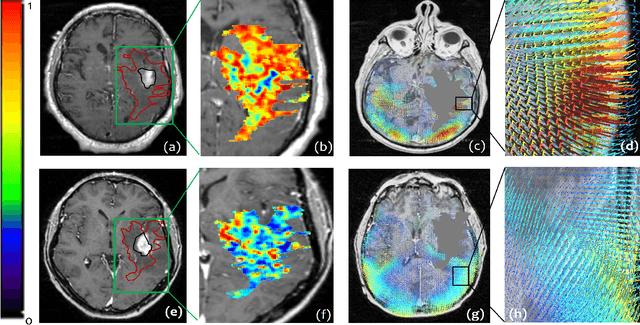
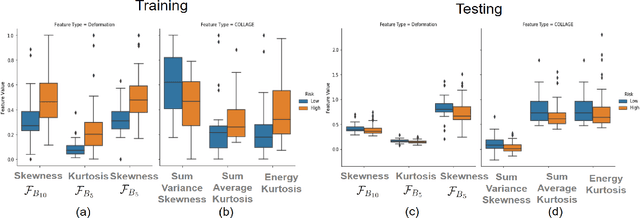
Abstract:The concept of tumor field effect implies that cancer is a systemic disease with its impact way beyond the visible tumor confines. For instance, in Glioblastoma (GBM), an aggressive brain tumor, the increase in intracranial pressure due to tumor burden often leads to brain herniation and poor outcomes. Our work is based on the rationale that highly aggressive tumors tend to grow uncontrollably, leading to pronounced biomechanical tissue deformations in the normal parenchyma, which when combined with local morphological differences in the tumor confines on MRI scans, will comprehensively capture tumor field effect. Specifically, we present an integrated MRI-based descriptor, radiomic-Deformation and Textural Heterogeneity (r-DepTH). This descriptor comprises measurements of the subtle perturbations in tissue deformations throughout the surrounding normal parenchyma due to mass effect. This involves non-rigidly aligning the patients MRI scans to a healthy atlas via diffeomorphic registration. The resulting inverse mapping is used to obtain the deformation field magnitudes in the normal parenchyma. These measurements are then combined with a 3D texture descriptor, Co-occurrence of Local Anisotropic Gradient Orientations (COLLAGE), which captures the morphological heterogeneity within the tumor confines, on MRI scans. R-DepTH, on N = 207 GBM cases (training set (St) = 128, testing set (Sv) = 79), demonstrated improved prognosis of overall survival by categorizing patients into low- (prolonged survival) and high-risk (poor survival) groups (on St, p-value = 0.0000035, and on Sv, p-value = 0.0024). R-DepTH descriptor may serve as a comprehensive MRI-based prognostic marker of disease aggressiveness and survival in solid tumors.
Spatial-And-Context aware (SpACe) "virtual biopsy" radiogenomic maps to target tumor mutational status on structural MRI
Jun 17, 2020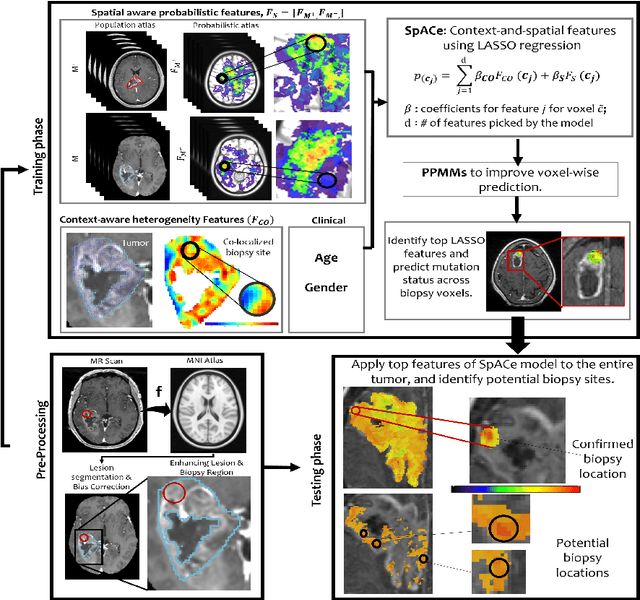
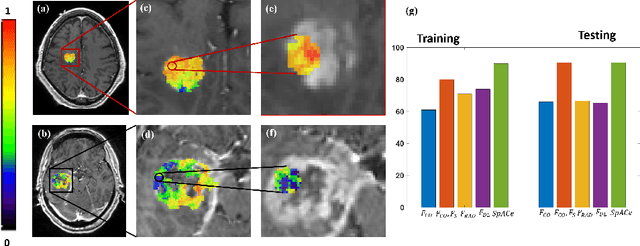
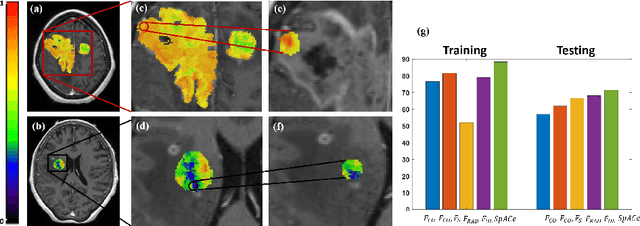
Abstract:With growing emphasis on personalized cancer-therapies,radiogenomics has shown promise in identifying target tumor mutational status on routine imaging (i.e. MRI) scans. These approaches fall into 2 categories: (1) deep-learning/radiomics (context-based), using image features from the entire tumor to identify the gene mutation status, or (2) atlas (spatial)-based to obtain likelihood of gene mutation status based on population statistics. While many genes (i.e. EGFR, MGMT) are spatially variant, a significant challenge in reliable assessment of gene mutation status on imaging has been the lack of available co-localized ground truth for training the models. We present Spatial-And-Context aware (SpACe) "virtual biopsy" maps that incorporate context-features from co-localized biopsy site along with spatial-priors from population atlases, within a Least Absolute Shrinkage and Selection Operator (LASSO) regression model, to obtain a per-voxel probability of the presence of a mutation status (M+ vs M-). We then use probabilistic pair-wise Markov model to improve the voxel-wise prediction probability. We evaluate the efficacy of SpACe maps on MRI scans with co-localized ground truth obtained from corresponding biopsy, to predict the mutation status of 2 driver genes in Glioblastoma: (1) EGFR (n=91), and (2) MGMT (n=81). When compared against deep-learning (DL) and radiomic models, SpACe maps obtained training and testing accuracies of 90% (n=71) and 90.48% (n=21) in identifying EGFR amplification status,compared to 80% and 71.4% via radiomics, and 74.28% and 65.5% via DL. For MGMT status, training and testing accuracies using SpACe were 88.3% (n=61) and 71.5% (n=20), compared to 52.4% and 66.7% using radiomics,and 79.3% and 68.4% using DL. Following validation,SpACe maps could provide surgical navigation to improve localization of sampling sites for targeting of specific driver genes in cancer.
Can tumor location on pre-treatment MRI predict likelihood of pseudo-progression versus tumor recurrence in Glioblastoma? A feasibility study
Jun 16, 2020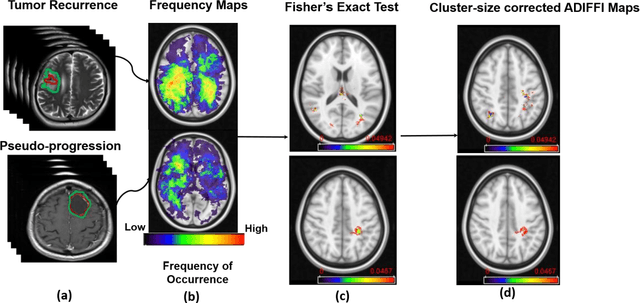

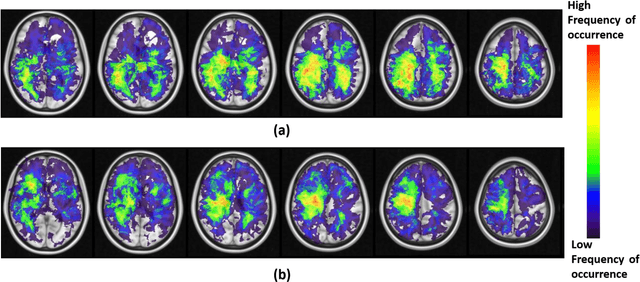
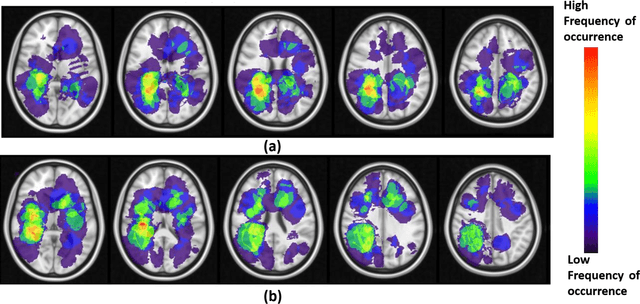
Abstract:A significant challenge in Glioblastoma (GBM) management is identifying pseudo-progression (PsP), a benign radiation-induced effect, from tumor recurrence, on routine imaging following conventional treatment. Previous studies have linked tumor lobar presence and laterality to GBM outcomes, suggesting that disease etiology and progression in GBM may be impacted by tumor location. Hence, in this feasibility study, we seek to investigate the following question: Can tumor location on treatment-na\"ive MRI provide early cues regarding likelihood of a patient developing pseudo-progression versus tumor recurrence? In this study, 74 pre-treatment Glioblastoma MRI scans with PsP (33) and tumor recurrence (41) were analyzed. First, enhancing lesion on Gd-T1w MRI and peri-lesional hyperintensities on T2w/FLAIR were segmented by experts and then registered to a brain atlas. Using patients from the two phenotypes, we construct two atlases by quantifying frequency of occurrence of enhancing lesion and peri-lesion hyperintensities, by averaging voxel intensities across the population. Analysis of differential involvement was then performed to compute voxel-wise significant differences (p-value<0.05) across the atlases. Statistically significant clusters were finally mapped to a structural atlas to provide anatomic localization of their location. Our results demonstrate that patients with tumor recurrence showed prominence of their initial tumor in the parietal lobe, while patients with PsP showed a multi-focal distribution of the initial tumor in the frontal and temporal lobes, insula, and putamen. These preliminary results suggest that lateralization of pre-treatment lesions towards certain anatomical areas of the brain may allow to provide early cues regarding assessing likelihood of occurrence of pseudo-progression from tumor recurrence on MRI scans.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge