Nasir Rajpoot
*: shared first/last authors
NuClick: A Deep Learning Framework for Interactive Segmentation of Microscopy Images
May 29, 2020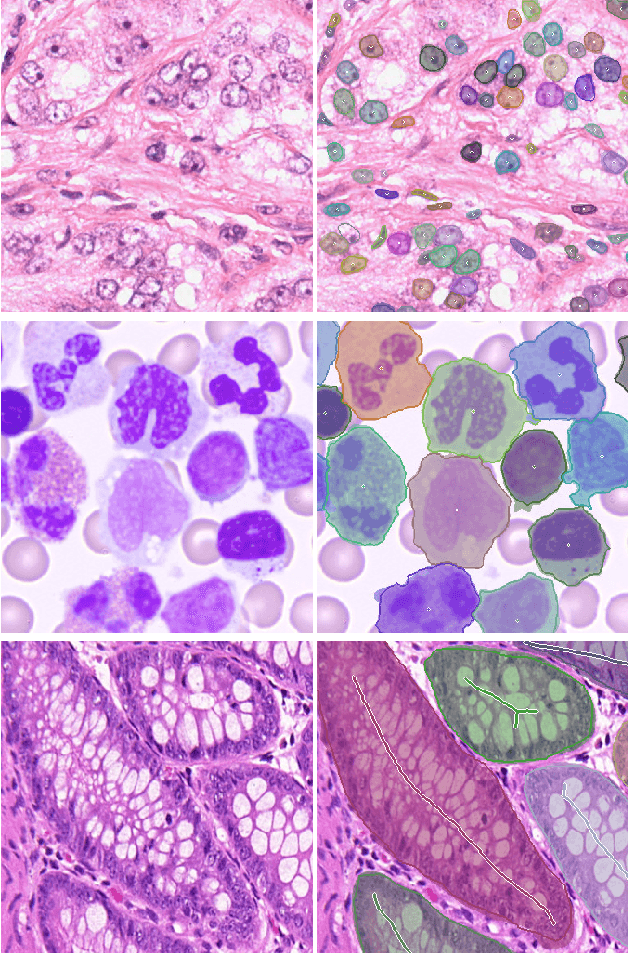
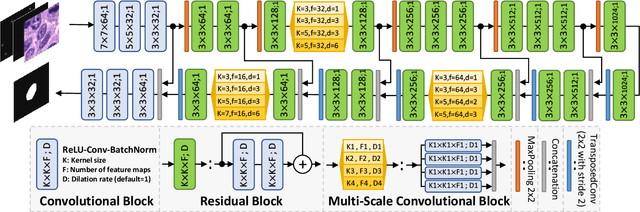
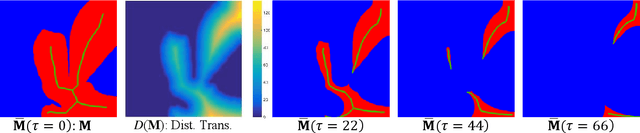
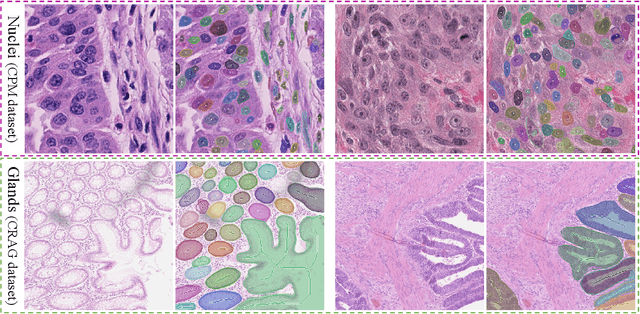
Abstract:Object Segmentation is an important step in the work-flow of computational pathology. Deep learning based models as the best forming models require huge amount of labeled data for precise and reliable prediction. However, collecting labeled data is expensive, because it necessarily involves expert knowledge. This is perhaps best illustrated by medical tasks where measurements call for expensive machinery and labels are the fruit of a time-consuming analysis that draws from multiple human experts. As nuclei, cells and glands are fundamental objects for downstream analysis in histology, in this paper we propose a simple CNN-based approach to speed up collecting segmentation annotation for these objects by utilizing minimum input from an annotator. We show for nuclei and cells as small objects, one click inside objects is enough to have precise annotation. For glands as large objects, providing a squiggle to show the extend of gland can guide the model to outline the exact boundaries. This supervisory signals are fed to network as an auxiliary channels along with RGB channels. With detailed experiments, we show that our approach is generalizable, robust against variations in the user input and that it can be used to obtain annotations for completely different domains. Practically, a model trained on the masks generated by NuClick could achieve first rank in LYON19 challenge. Furthermore, as the output of our framework, we release two data-sets: 1) a dataset of lymphocyte annotations within IHC images and 2) a dataset of WBCs annotated in blood sample images.
Multi-Task Learning in Histo-pathology for Widely Generalizable Model
May 09, 2020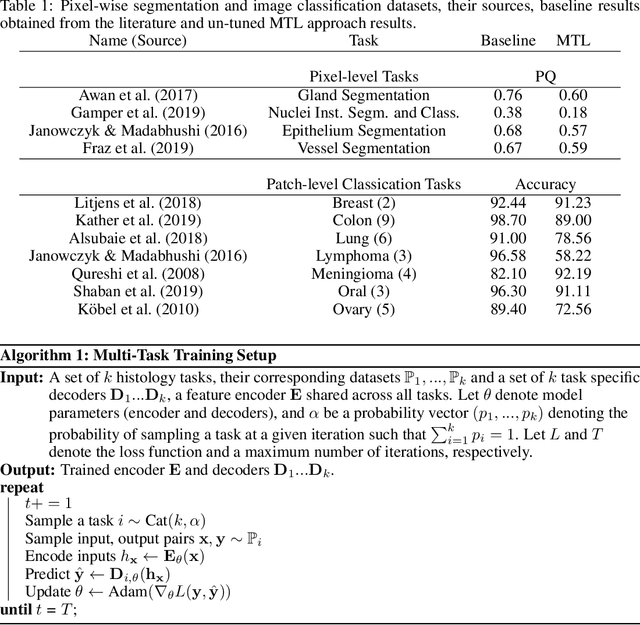
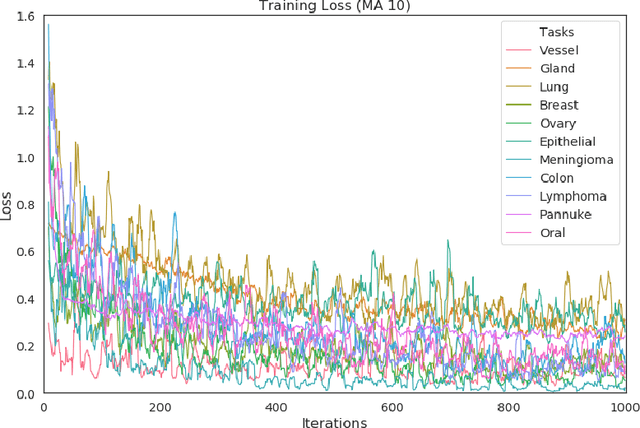
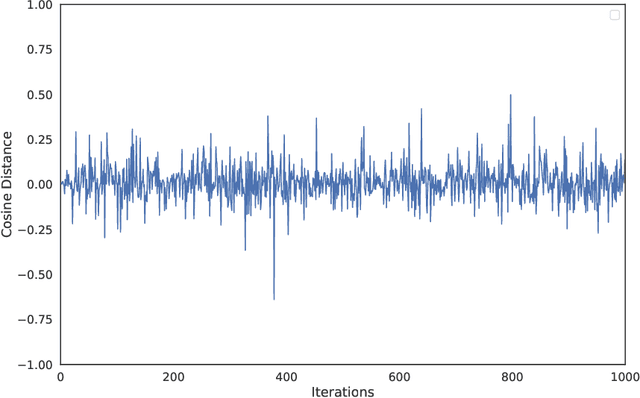
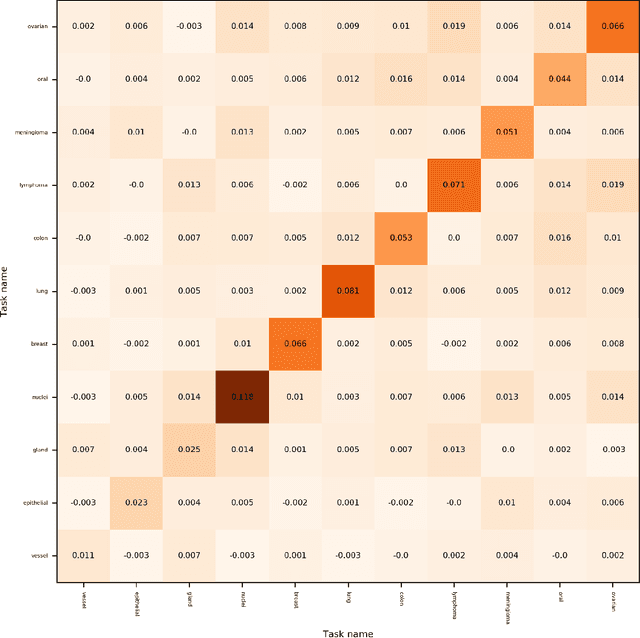
Abstract:In this work we show preliminary results of deep multi-task learning in the area of computational pathology. We combine 11 tasks ranging from patch-wise oral cancer classification, one of the most prevalent cancers in the developing world, to multi-tissue nuclei instance segmentation and classification.
PanNuke Dataset Extension, Insights and Baselines
Apr 22, 2020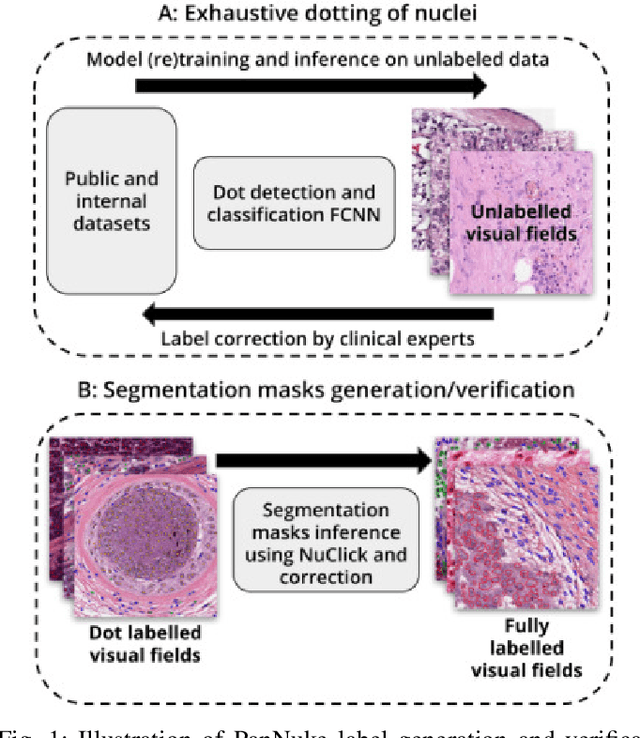
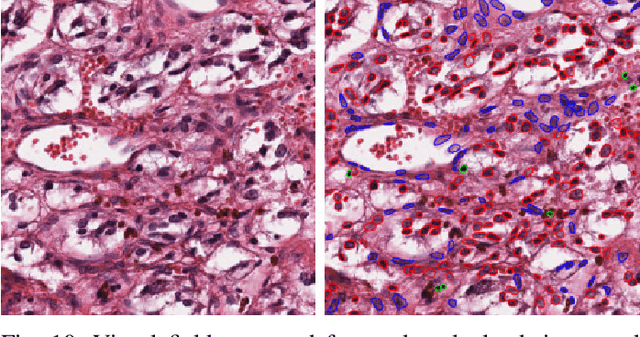
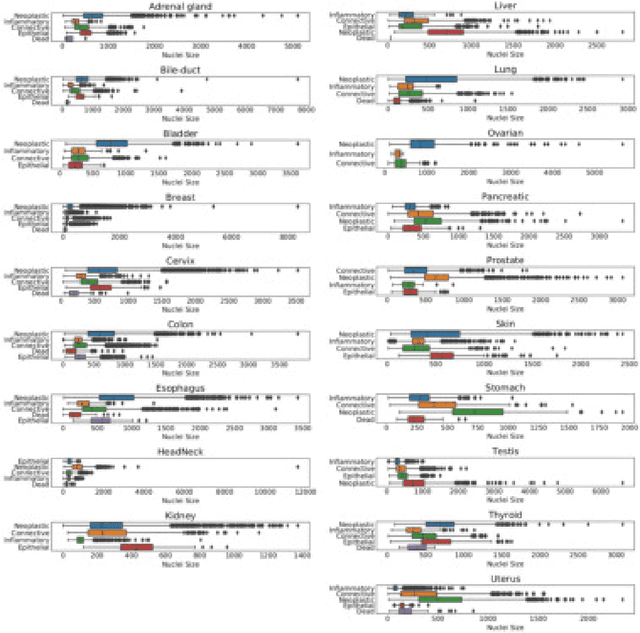
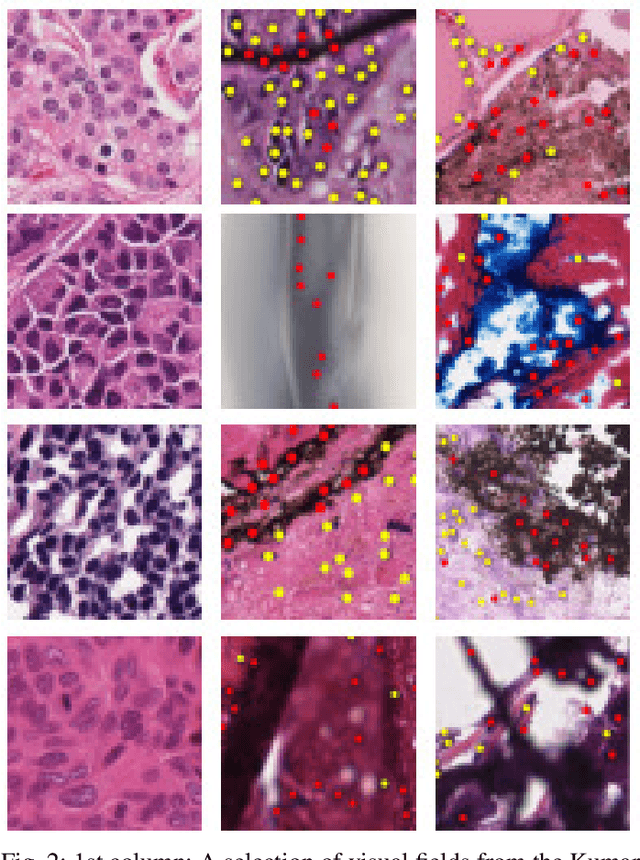
Abstract:The emerging area of computational pathology (CPath) is ripe ground for the application of deep learning (DL) methods to healthcare due to the sheer volume of raw pixel data in whole-slide images (WSIs) of cancerous tissue slides. However, it is imperative for the DL algorithms relying on nuclei-level details to be able to cope with data from `the clinical wild', which tends to be quite challenging. We study, and extend recently released PanNuke dataset consisting of ~200,000 nuclei categorized into 5 clinically important classes for the challenging tasks of segmenting and classifying nuclei in WSIs. Previous pan-cancer datasets consisted of only up to 9 different tissues and up to 21,000 unlabeled nuclei and just over 24,000 labeled nuclei with segmentation masks. PanNuke consists of 19 different tissue types that have been semi-automatically annotated and quality controlled by clinical pathologists, leading to a dataset with statistics similar to the clinical wild and with minimal selection bias. We study the performance of segmentation and classification models when applied to the proposed dataset and demonstrate the application of models trained on PanNuke to whole-slide images. We provide comprehensive statistics about the dataset and outline recommendations and research directions to address the limitations of existing DL tools when applied to real-world CPath applications.
Dense Steerable Filter CNNs for Exploiting Rotational Symmetry in Histology Images
Apr 06, 2020
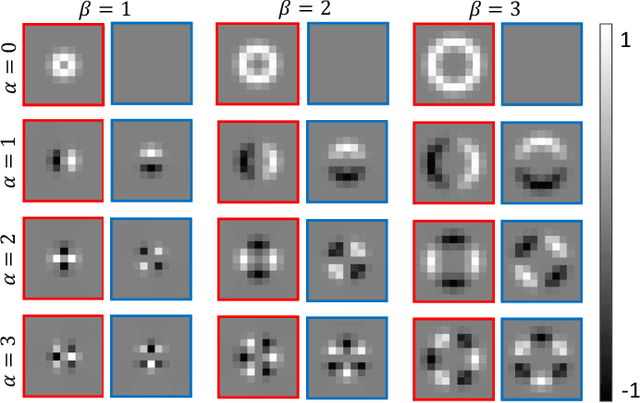

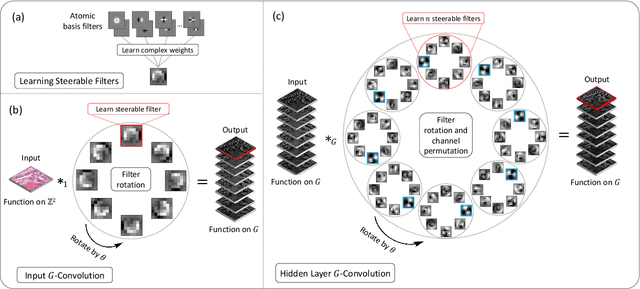
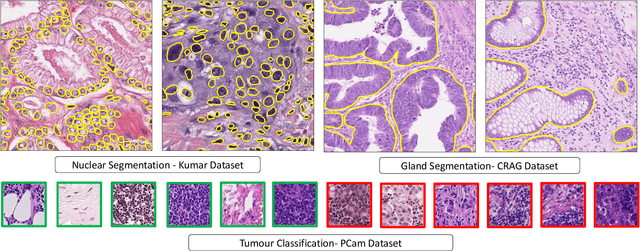
Abstract:Histology images are inherently symmetric under rotation, where each orientation is equally as likely to appear. However, this rotational symmetry is not widely utilised as prior knowledge in modern Convolutional Neural Networks (CNNs), resulting in data hungry models that learn independent features at each orientation. Allowing CNNs to be rotation-equivariant removes the necessity to learn this set of transformations from the data and instead frees up model capacity, allowing more discriminative features to be learned. This reduction in the number of required parameters also reduces the risk of overfitting. In this paper, we propose Dense Steerable Filter CNNs (DSF-CNNs) that use group convolutions with multiple rotated copies of each filter in a densely connected framework. Each filter is defined as a linear combination of steerable basis filters, enabling exact rotation and decreasing the number of trainable parameters compared to standard filters. We also provide the first in-depth comparison of different rotation-equivariant CNNs for histology image analysis and demonstrate the advantage of encoding rotational symmetry into modern architectures. We show that DSF-CNNs achieve state-of-the-art performance, with significantly fewer parameters, when applied to three different tasks in the area of computational pathology: breast tumour classification, colon gland segmentation and multi-tissue nuclear segmentation.
Meta-SVDD: Probabilistic Meta-Learning for One-Class Classification in Cancer Histology Images
Mar 06, 2020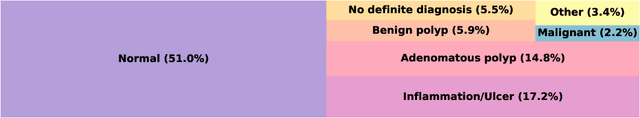
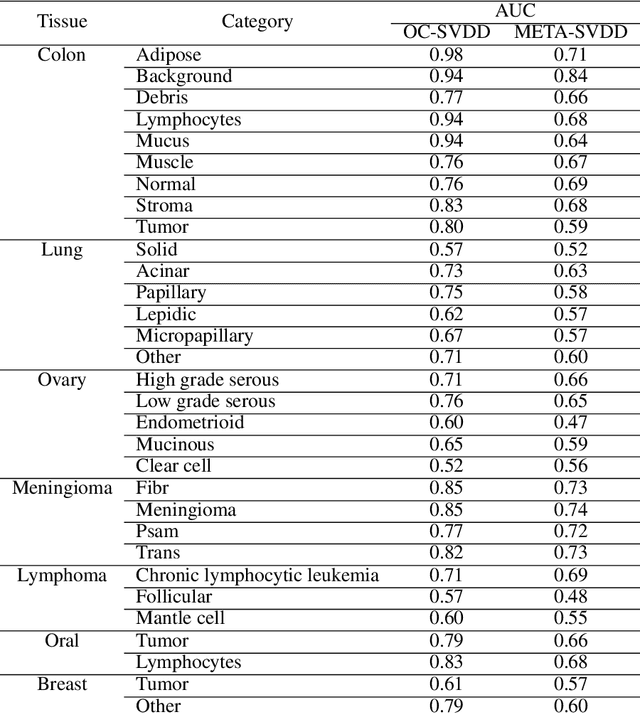
Abstract:To train a robust deep learning model, one usually needs a balanced set of categories in the training data. The data acquired in a medical domain, however, frequently contains an abundance of healthy patients, versus a small variety of positive, abnormal cases. Moreover, the annotation of a positive sample requires time consuming input from medical domain experts. This scenario would suggest a promise for one-class classification type approaches. In this work we propose a general one-class classification model for histology, that is meta-trained on multiple histology datasets simultaneously, and can be applied to new tasks without expensive re-training. This model could be easily used by pathology domain experts, and potentially be used for screening purposes.
NuClick: From Clicks in the Nuclei to Nuclear Boundaries
Sep 07, 2019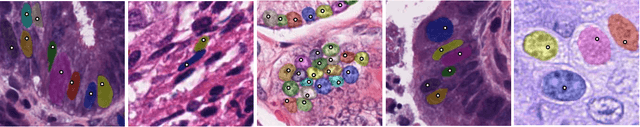
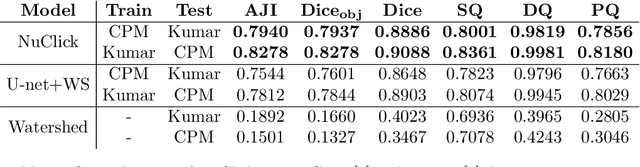
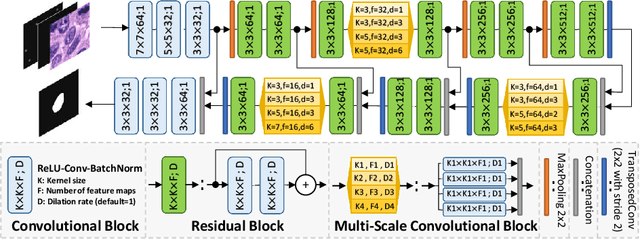
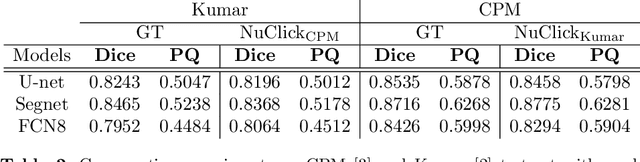
Abstract:Best performing nuclear segmentation methods are based on deep learning algorithms that require a large amount of annotated data. However, collecting annotations for nuclear segmentation is a very labor-intensive and time-consuming task. Thereby, providing a tool that can facilitate and speed up this procedure is very demanding. Here we propose a simple yet efficient framework based on convolutional neural networks, named NuClick, which can precisely segment nuclei boundaries by accepting a single point position (or click) inside each nucleus. Based on the clicked positions, inclusion and exclusion maps are generated which comprise 2D Gaussian distributions centered on those positions. These maps serve as guiding signals for the network as they are concatenated to the input image. The inclusion map focuses on the desired nucleus while the exclusion map indicates neighboring nuclei and improve the results of segmentation in scenes with nuclei clutter. The NuClick not only facilitates collecting more annotation from unseen data but also leads to superior segmentation output for deep models. It is also worth mentioning that an instance segmentation model trained on NuClick generated labels was able to rank first in LYON19 challenge.
CGC-Net: Cell Graph Convolutional Network for Grading of Colorectal Cancer Histology Images
Sep 03, 2019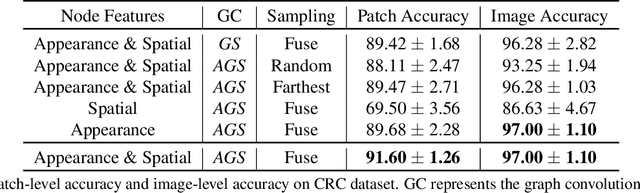
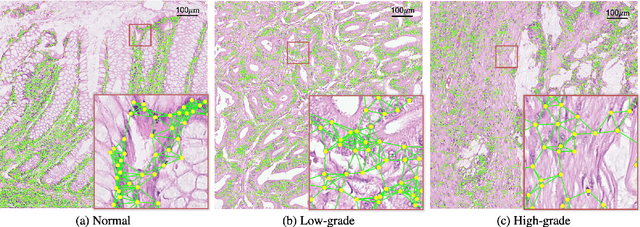
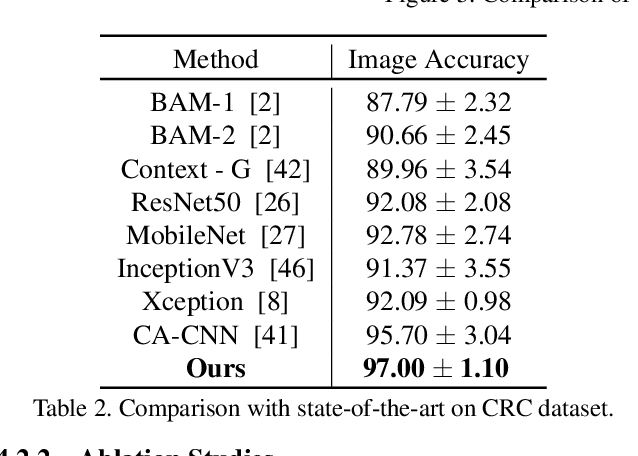
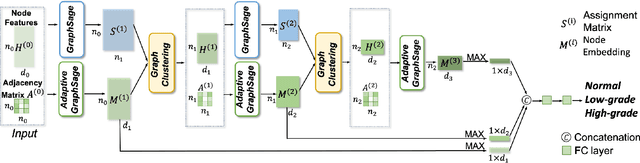
Abstract:Colorectal cancer (CRC) grading is typically carried out by assessing the degree of gland formation within histology images. To do this, it is important to consider the overall tissue micro-environment by assessing the cell-level information along with the morphology of the gland. However, current automated methods for CRC grading typically utilise small image patches and therefore fail to incorporate the entire tissue micro-architecture for grading purposes. To overcome the challenges of CRC grading, we present a novel cell-graph convolutional neural network (CGC-Net) that converts each large histology image into a graph, where each node is represented by a nucleus within the original image and cellular interactions are denoted as edges between these nodes according to node similarity. The CGC-Net utilises nuclear appearance features in addition to the spatial location of nodes to further boost the performance of the algorithm. To enable nodes to fuse multi-scale information, we introduce Adaptive GraphSage, which is a graph convolution technique that combines multi-level features in a data-driven way. Furthermore, to deal with redundancy in the graph, we propose a sampling technique that removes nodes in areas of dense nuclear activity. We show that modeling the image as a graph enables us to effectively consider a much larger image (around 16$\times$ larger) than traditional patch-based approaches and model the complex structure of the tissue micro-environment. We construct cell graphs with an average of over 3,000 nodes on a large CRC histology image dataset and report state-of-the-art results as compared to recent patch-based as well as contextual patch-based techniques, demonstrating the effectiveness of our method.
Nuclear Instance Segmentation using a Proposal-Free Spatially Aware Deep Learning Framework
Aug 27, 2019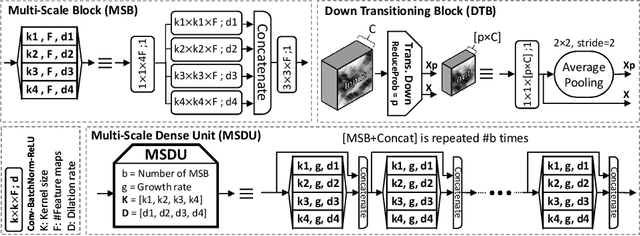
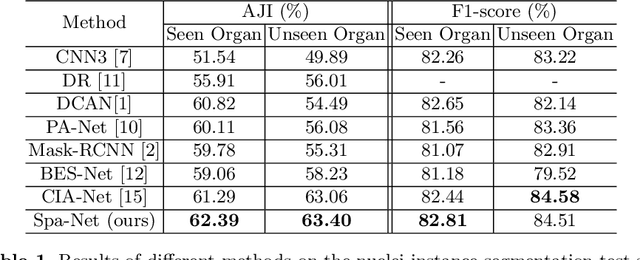
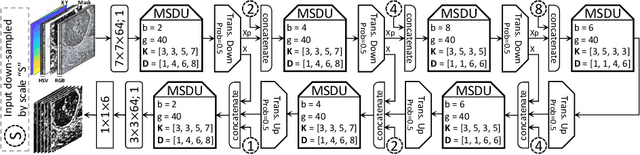
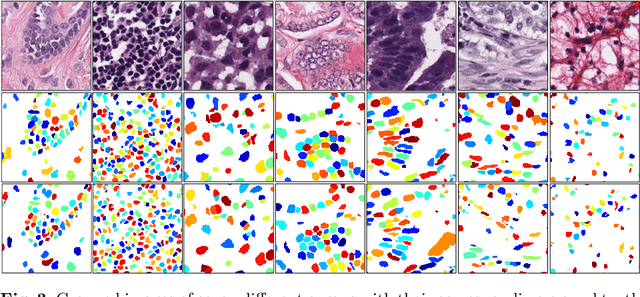
Abstract:Nuclear segmentation in histology images is a challenging task due to significant variations in the shape and appearance of nuclei. One of the main hurdles in nuclear instance segmentation is overlapping nuclei where a smart algorithm is needed to separate each nucleus. In this paper, we introduce a proposal-free deep learning based framework to address these challenges. To this end, we propose a spatially-aware network (SpaNet) to capture spatial information in a multi-scale manner. A dual-head variation of the SpaNet is first utilized to predict the pixel-wise segmentation and centroid detection maps of nuclei. Based on these outputs, a single-head SpaNet predicts the positional information related to each nucleus instance. Spectral clustering method is applied on the output of the last SpaNet, which utilizes the nuclear mask and the Gaussian-like detection map for determining the connected components and associated cluster identifiers, respectively. The output of the clustering method is the final nuclear instance segmentation mask. We applied our method on a publicly available multi-organ data set and achieved state-of-the-art performance for nuclear segmentation.
XY Network for Nuclear Segmentation in Multi-Tissue Histology Images
Dec 16, 2018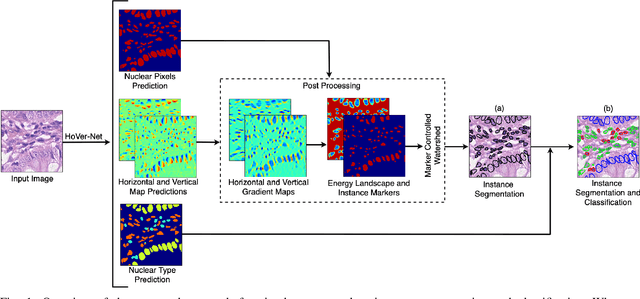
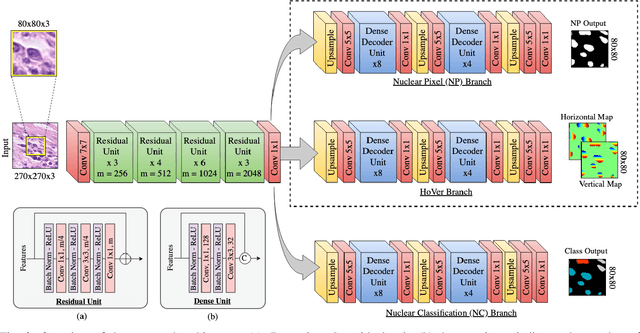
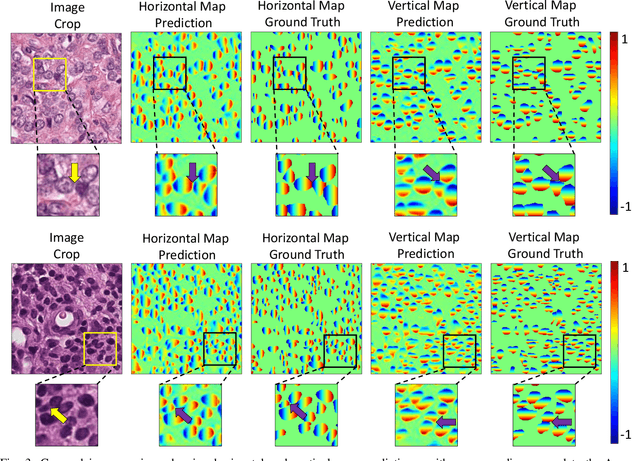
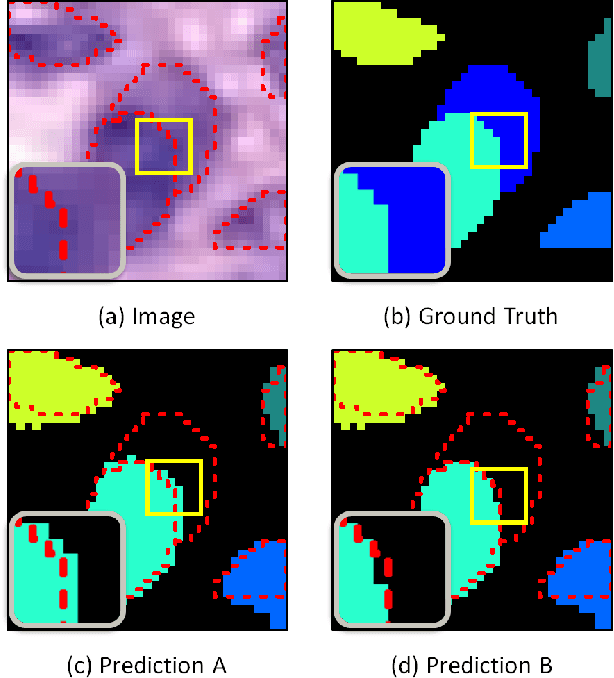
Abstract:Nuclear segmentation within Haematoxylin & Eosin stained histology images is a fundamental prerequisite in the digital pathology work-flow, due to the ability for nuclear features to act as key diagnostic markers. The development of automated methods for nuclear segmentation enables the quantitative analysis of tens of thousands of nuclei within a whole-slide pathology image, opening up possibilities of further analysis of large-scale nuclear morphometry. However, automated nuclear segmentation is faced with a major challenge in that there are several different types of nuclei, some of them exhibiting large intra-class variability such as the tumour cells. Additionally, some of the nuclei are often clustered together. To address these challenges, we present a novel convolutional neural network for automated nuclear segmentation that leverages the instance-rich information encoded within the vertical and horizontal distances of nuclear pixels to their centres of mass. These distances are then utilised to separate clustered nuclei, resulting in an accurate segmentation, particularly in areas with overlapping instances. We demonstrate state-of-the-art performance compared to other methods on four independent multi-tissue histology image datasets. Furthermore, we propose an interpretable and reliable evaluation framework that effectively quantifies nuclear segmentation performance and overcomes the limitations of existing performance measures.
Methods for Segmentation and Classification of Digital Microscopy Tissue Images
Oct 31, 2018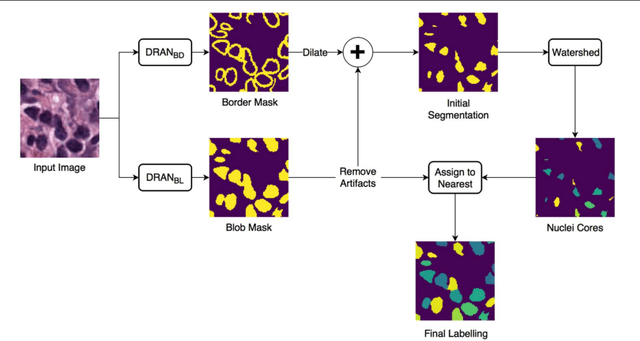
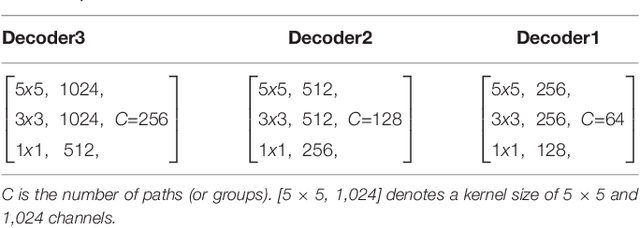

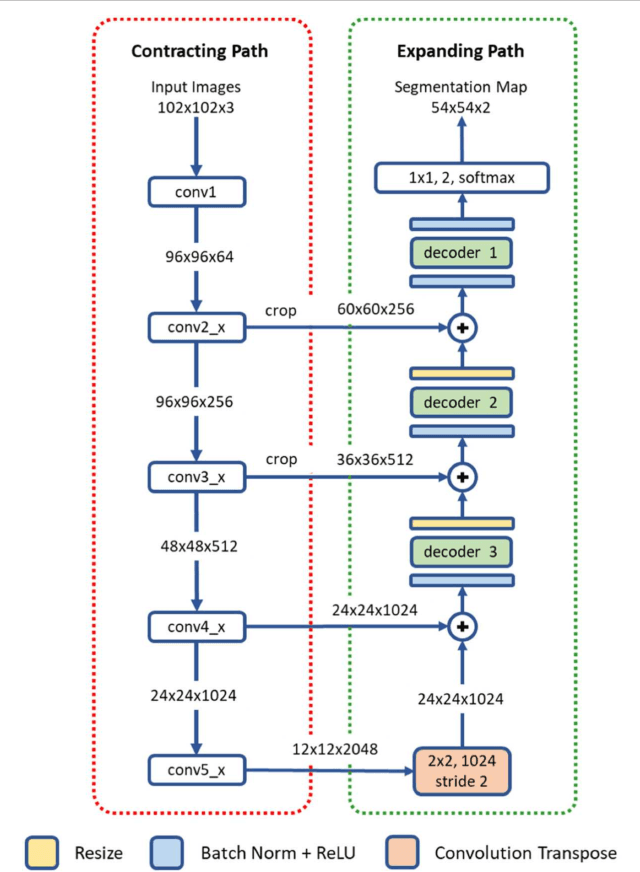
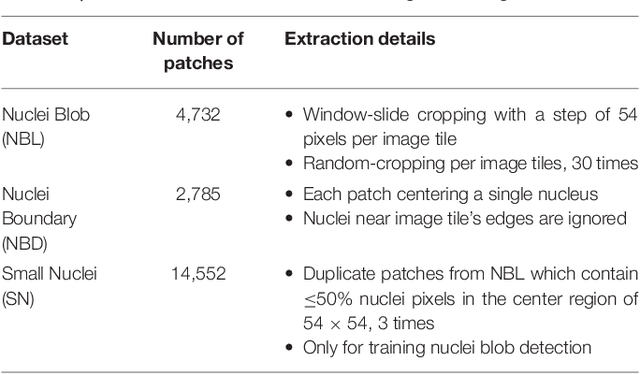
Abstract:High-resolution microscopy images of tissue specimens provide detailed information about the morphology of normal and diseased tissue. Image analysis of tissue morphology can help cancer researchers develop a better understanding of cancer biology. Segmentation of nuclei and classification of tissue images are two common tasks in tissue image analysis. Development of accurate and efficient algorithms for these tasks is a challenging problem because of the complexity of tissue morphology and tumor heterogeneity. In this paper we present two computer algorithms; one designed for segmentation of nuclei and the other for classification of whole slide tissue images. The segmentation algorithm implements a multiscale deep residual aggregation network to accurately segment nuclear material and then separate clumped nuclei into individual nuclei. The classification algorithm initially carries out patch-level classification via a deep learning method, then patch-level statistical and morphological features are used as input to a random forest regression model for whole slide image classification. The segmentation and classification algorithms were evaluated in the MICCAI 2017 Digital Pathology challenge. The segmentation algorithm achieved an accuracy score of 0.78. The classification algorithm achieved an accuracy score of 0.81.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge