Nan Wu
Program-to-Circuit: Exploiting GNNs for Program Representation and Circuit Translation
Sep 13, 2021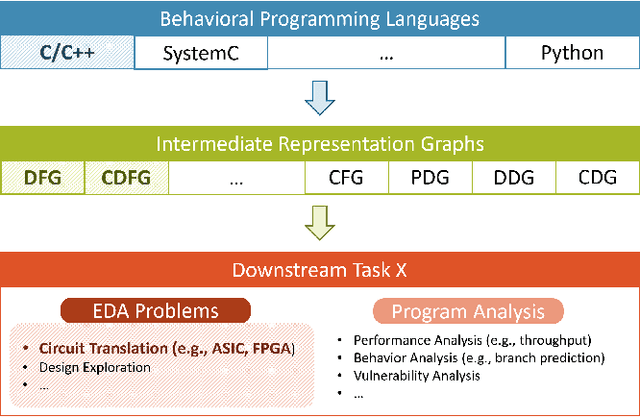
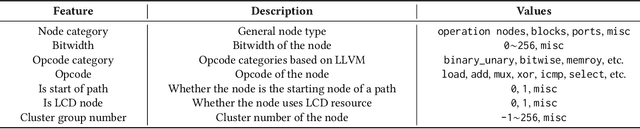

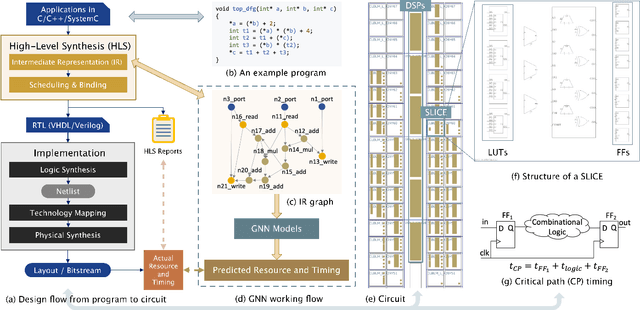

Abstract:Circuit design is complicated and requires extensive domain-specific expertise. One major obstacle stuck on the way to hardware agile development is the considerably time-consuming process of accurate circuit quality evaluation. To significantly expedite the circuit evaluation during the translation from behavioral languages to circuit designs, we formulate it as a Program-to-Circuit problem, aiming to exploit the representation power of graph neural networks (GNNs) by representing C/C++ programs as graphs. The goal of this work is four-fold. First, we build a standard benchmark containing 40k C/C++ programs, each of which is translated to a circuit design with actual hardware quality metrics, aiming to facilitate the development of effective GNNs targeting this high-demand circuit design area. Second, 14 state-of-the-art GNN models are analyzed on the Program-to-Circuit problem. We identify key design challenges of this problem, which should be carefully handled but not yet solved by existing GNNs. The goal is to provide domain-specific knowledge for designing GNNs with suitable inductive biases. Third, we discuss three sets of real-world benchmarks for GNN generalization evaluation, and analyze the performance gap between standard programs and the real-case ones. The goal is to enable transfer learning from limited training data to real-world large-scale circuit design problems. Fourth, the Program-to-Circuit problem is a representative within the Program-to-X framework, a set of program-based analysis problems with various downstream tasks. The in-depth understanding of strength and weaknesses in applying GNNs on Program-to-Circuit could largely benefit the entire family of Program-to-X. Pioneering in this direction, we expect more GNN endeavors to revolutionize this high-demand Program-to-Circuit problem and to enrich the expressiveness of GNNs on programs.
Weakly-supervised High-resolution Segmentation of Mammography Images for Breast Cancer Diagnosis
Jun 15, 2021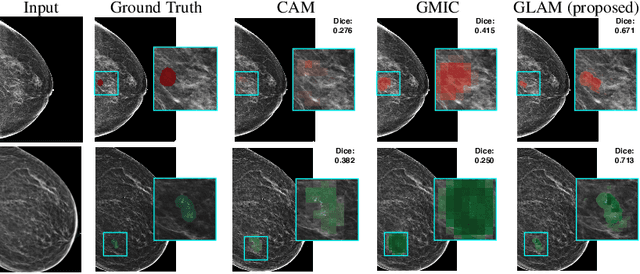
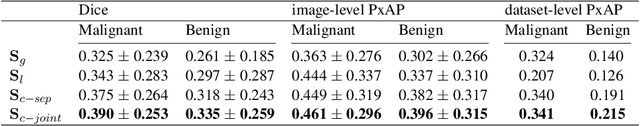
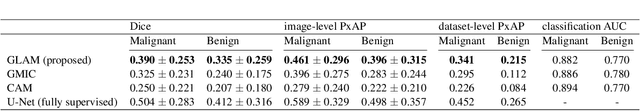
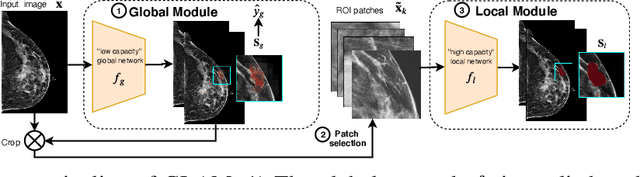

Abstract:In the last few years, deep learning classifiers have shown promising results in image-based medical diagnosis. However, interpreting the outputs of these models remains a challenge. In cancer diagnosis, interpretability can be achieved by localizing the region of the input image responsible for the output, i.e. the location of a lesion. Alternatively, segmentation or detection models can be trained with pixel-wise annotations indicating the locations of malignant lesions. Unfortunately, acquiring such labels is labor-intensive and requires medical expertise. To overcome this difficulty, weakly-supervised localization can be utilized. These methods allow neural network classifiers to output saliency maps highlighting the regions of the input most relevant to the classification task (e.g. malignant lesions in mammograms) using only image-level labels (e.g. whether the patient has cancer or not) during training. When applied to high-resolution images, existing methods produce low-resolution saliency maps. This is problematic in applications in which suspicious lesions are small in relation to the image size. In this work, we introduce a novel neural network architecture to perform weakly-supervised segmentation of high-resolution images. The proposed model selects regions of interest via coarse-level localization, and then performs fine-grained segmentation of those regions. We apply this model to breast cancer diagnosis with screening mammography, and validate it on a large clinically-realistic dataset. Measured by Dice similarity score, our approach outperforms existing methods by a large margin in terms of localization performance of benign and malignant lesions, relatively improving the performance by 39.6% and 20.0%, respectively. Code and the weights of some of the models are available at https://github.com/nyukat/GLAM
IronMan: GNN-assisted Design Space Exploration in High-Level Synthesis via Reinforcement Learning
Feb 16, 2021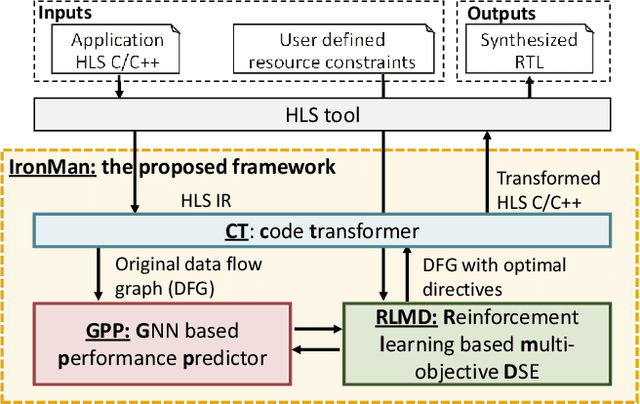
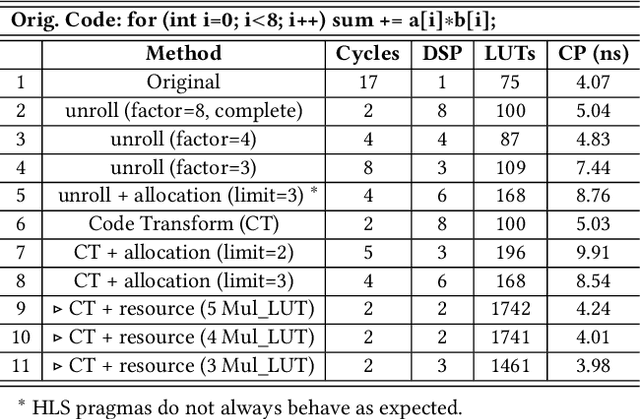
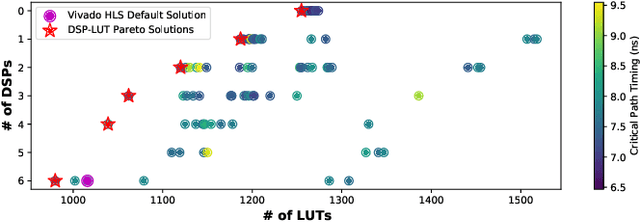
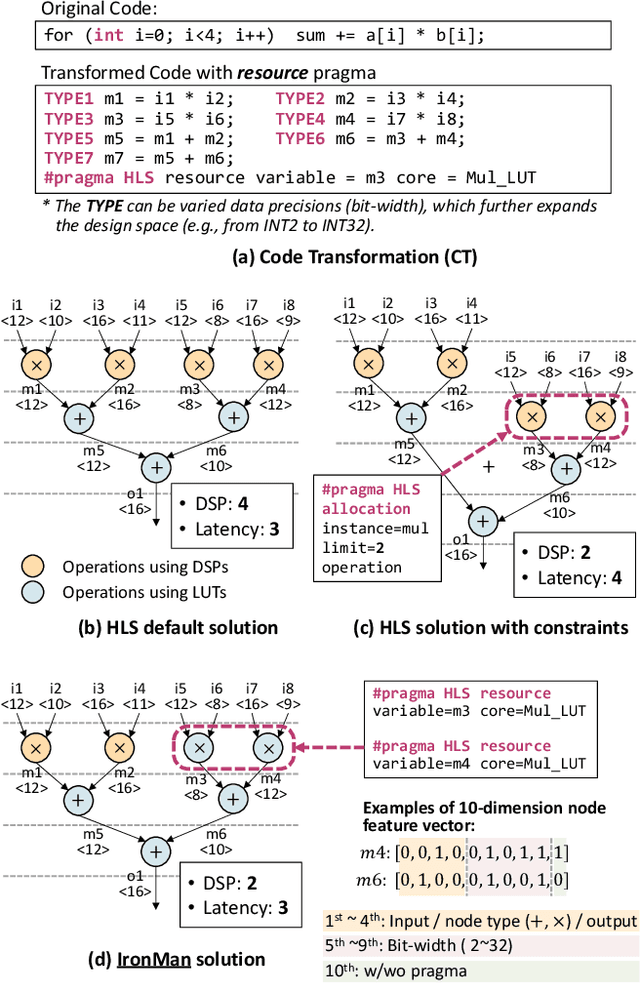
Abstract:Despite the great success of High-Level Synthesis (HLS) tools, we observe several unresolved challenges: 1) the high-level abstraction of programming styles in HLS sometimes conceals optimization opportunities; 2) existing HLS tools do not provide flexible trade-off (Pareto) solutions among different objectives and constraints; 3) the actual quality of the resulting RTL designs is hard to predict. To address these challenges, we propose an end-to-end framework, namelyIronMan. The primary goal is to enable a flexible and automated design space exploration (DSE), to provide either optimal solutions under user-specified constraints, or various trade-offs among different objectives (such as different types of resources, area, and latency). Such DSE either requires tedious manual efforts or is not achievable to attain these goals through existing HLS tools. There are three components in IronMan: 1) GPP, a highly accurate graph-neural-network-based performance and resource predictor; 2) RLMD, a reinforcement-learning-based multi-objective DSE engine that explores the optimal resource allocation strategy, to provide Pareto solutions between different objectives; 3) CT, a code transformer to assist RLMD and GPP, which extracts the data flow graph from original HLS C/C++ and automatically generates synthesizable code with HLS directives. The experimental results show that: 1) GPP achieves high prediction accuracy, reducing prediction errors of HLS tools by 10.9x in resource utilization and 5.7x in timing; 2) RLMD obtains optimal or Pareto solutions that outperform the genetic algorithm and simulated annealing by 12.7% and 12.9%, respectively; 3) IronMan is able to find optimized solutions perfectly matching various DSP constraints, with 2.54x fewer DSPs and up to 6x shorter latency than those of HLS tools while being up to 400x faster than the heuristic algorithms and HLS tools.
A Survey of Machine Learning for Computer Architecture and Systems
Feb 16, 2021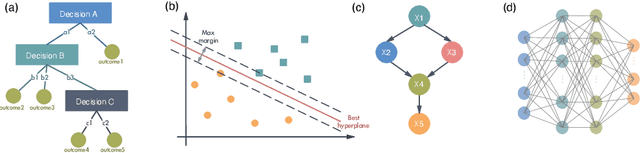
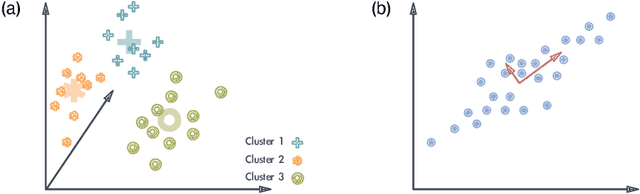
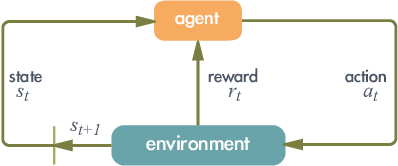
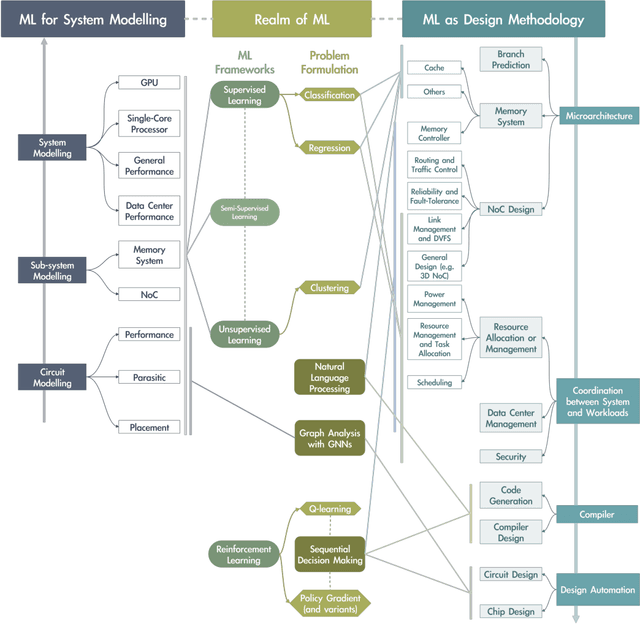
Abstract:It has been a long time that computer architecture and systems are optimized to enable efficient execution of machine learning (ML) algorithms or models. Now, it is time to reconsider the relationship between ML and systems, and let ML transform the way that computer architecture and systems are designed. This embraces a twofold meaning: the improvement of designers' productivity, and the completion of the virtuous cycle. In this paper, we present a comprehensive review of work that applies ML for system design, which can be grouped into two major categories, ML-based modelling that involves predictions of performance metrics or some other criteria of interest, and ML-based design methodology that directly leverages ML as the design tool. For ML-based modelling, we discuss existing studies based on their target level of system, ranging from the circuit level to the architecture/system level. For ML-based design methodology, we follow a bottom-up path to review current work, with a scope of (micro-)architecture design (memory, branch prediction, NoC), coordination between architecture/system and workload (resource allocation and management, data center management, and security), compiler, and design automation. We further provide a future vision of opportunities and potential directions, and envision that applying ML for computer architecture and systems would thrive in the community.
Eigen-convergence of Gaussian kernelized graph Laplacian by manifold heat interpolation
Jan 25, 2021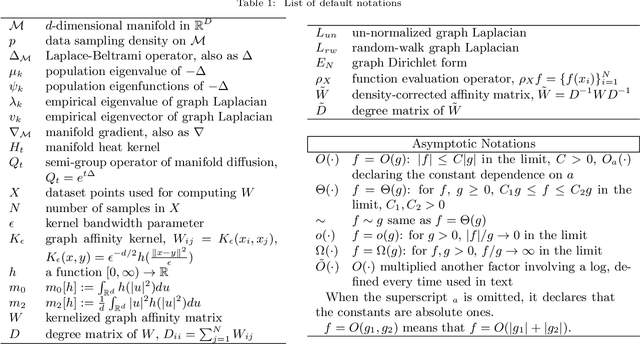
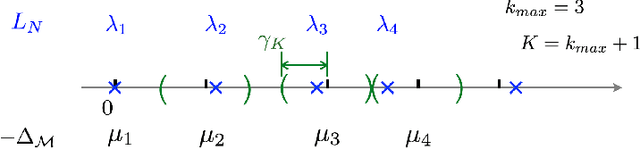
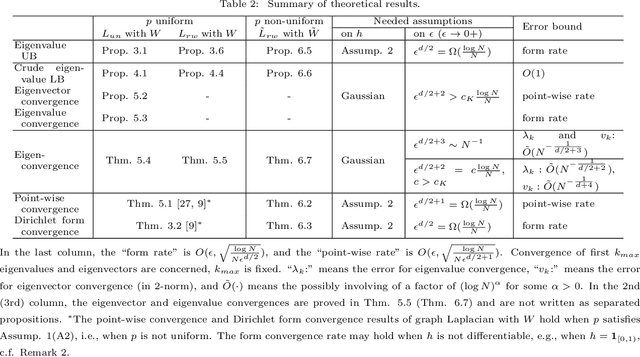
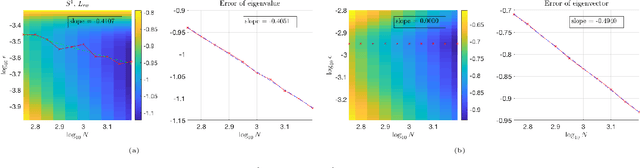
Abstract:This work studies the spectral convergence of graph Laplacian to the Laplace-Beltrami operator when the graph affinity matrix is constructed from $N$ random samples on a $d$-dimensional manifold embedded in a possibly high dimensional space. By analyzing Dirichlet form convergence and constructing candidate approximate eigenfunctions via convolution with manifold heat kernel, we prove that, with Gaussian kernel, one can set the kernel bandwidth parameter $\epsilon \sim (\log N/ N)^{1/(d/2+2)}$ such that the eigenvalue convergence rate is $N^{-1/(d/2+2)}$ and the eigenvector convergence in 2-norm has rate $N^{-1/(d+4)}$; When $\epsilon \sim N^{-1/(d/2+3)}$, both eigenvalue and eigenvector rates are $N^{-1/(d/2+3)}$. These rates are up to a $\log N$ factor and proved for finitely many low-lying eigenvalues. The result holds for un-normalized and random-walk graph Laplacians when data are uniformly sampled on the manifold, as well as the density-corrected graph Laplacian (where the affinity matrix is normalized by the degree matrix from both sides) with non-uniformly sampled data. As an intermediate result, we prove new point-wise and Dirichlet form convergence rates for the density-corrected graph Laplacian. Numerical results are provided to verify the theory.
Phoebe: Reuse-Aware Online Caching with Reinforcement Learning for Emerging Storage Models
Nov 13, 2020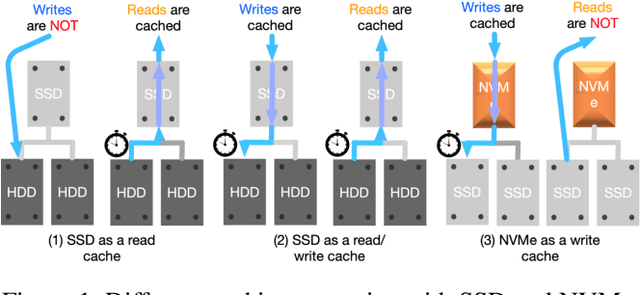
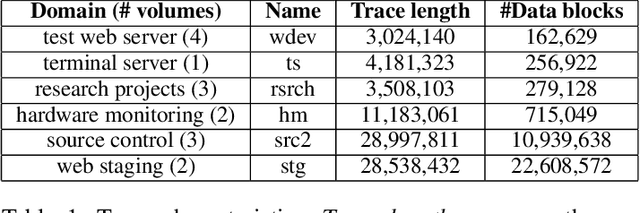
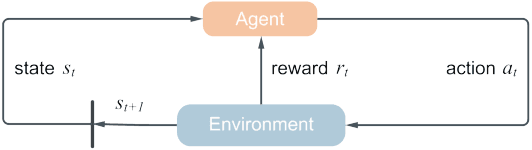
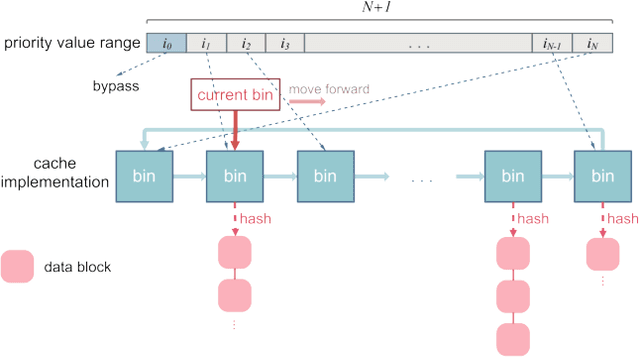
Abstract:With data durability, high access speed, low power efficiency and byte addressability, NVMe and SSD, which are acknowledged representatives of emerging storage technologies, have been applied broadly in many areas. However, one key issue with high-performance adoption of these technologies is how to properly define intelligent cache layers such that the performance gap between emerging technologies and main memory can be well bridged. To this end, we propose Phoebe, a reuse-aware reinforcement learning framework for the optimal online caching that is applicable for a wide range of emerging storage models. By continuous interacting with the cache environment and the data stream, Phoebe is capable to extract critical temporal data dependency and relative positional information from a single trace, becoming ever smarter over time. To reduce training overhead during online learning, we utilize periodical training to amortize costs. Phoebe is evaluated on a set of Microsoft cloud storage workloads. Experiment results show that Phoebe is able to close the gap of cache miss rate from LRU and a state-of-the-art online learning based cache policy to the Belady's optimal policy by 70.3% and 52.6%, respectively.
Diffusion Based Gaussian Processes on Restricted Domains
Oct 14, 2020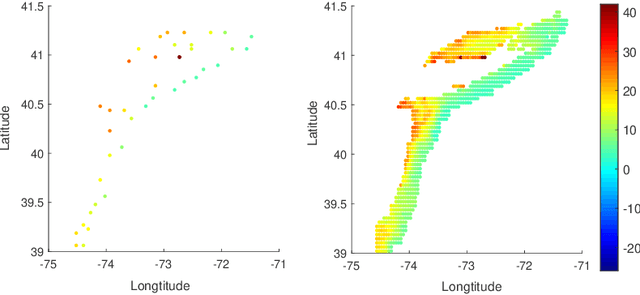
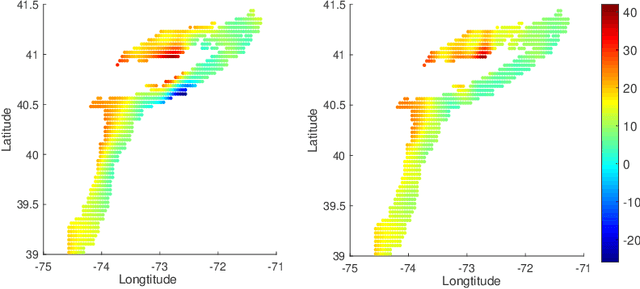
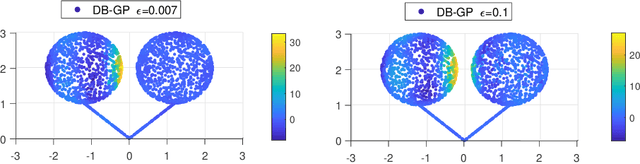
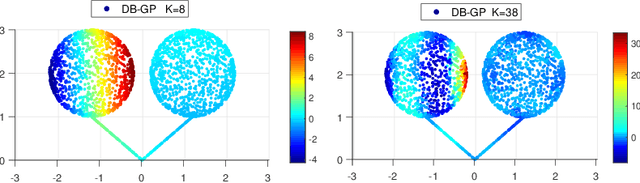
Abstract:In nonparametric regression and spatial process modeling, it is common for the inputs to fall in a restricted subset of Euclidean space. For example, the locations at which spatial data are collected may be restricted to a narrow non-linear subset, such as near the edge of a lake. Typical kernel-based methods that do not take into account the intrinsic geometric of the domain across which observations are collected may produce sub-optimal results. In this article, we focus on solving this problem in the context of Gaussian process (GP) models, proposing a new class of diffusion-based GPs (DB-GPs), which learn a covariance that respects the geometry of the input domain. We use the term `diffusion-based' as the idea is to measure intrinsic distances between inputs in a restricted domain via a diffusion process. As the heat kernel is intractable computationally, we approximate the covariance using finitely-many eigenpairs of the Graph Laplacian (GL). Our proposed algorithm has the same order of computational complexity as current GP algorithms using simple covariance kernels. We provide substantial theoretical support for the DB-GP methodology, and illustrate performance gains through toy examples, simulation studies, and applications to ecology data.
Reducing false-positive biopsies with deep neural networks that utilize local and global information in screening mammograms
Sep 19, 2020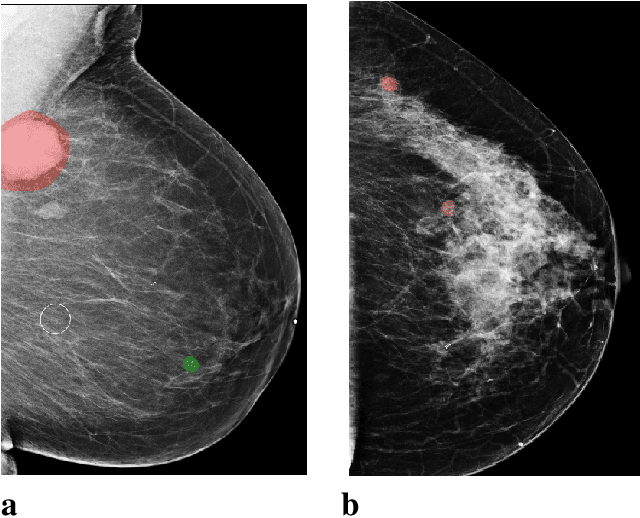
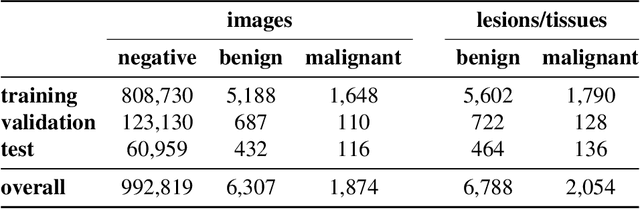
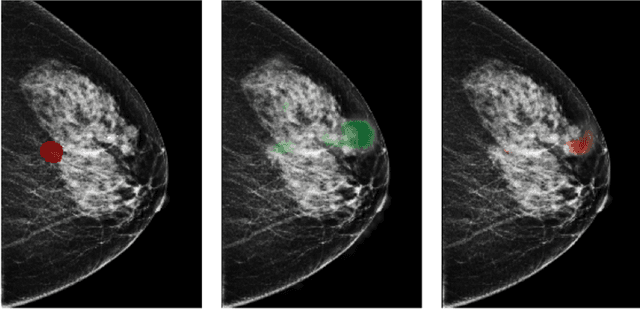
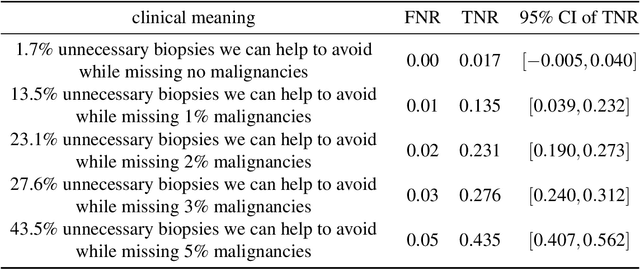
Abstract:Breast cancer is the most common cancer in women, and hundreds of thousands of unnecessary biopsies are done around the world at a tremendous cost. It is crucial to reduce the rate of biopsies that turn out to be benign tissue. In this study, we build deep neural networks (DNNs) to classify biopsied lesions as being either malignant or benign, with the goal of using these networks as second readers serving radiologists to further reduce the number of false positive findings. We enhance the performance of DNNs that are trained to learn from small image patches by integrating global context provided in the form of saliency maps learned from the entire image into their reasoning, similar to how radiologists consider global context when evaluating areas of interest. Our experiments are conducted on a dataset of 229,426 screening mammography exams from 141,473 patients. We achieve an AUC of 0.8 on a test set consisting of 464 benign and 136 malignant lesions.
An artificial intelligence system for predicting the deterioration of COVID-19 patients in the emergency department
Aug 04, 2020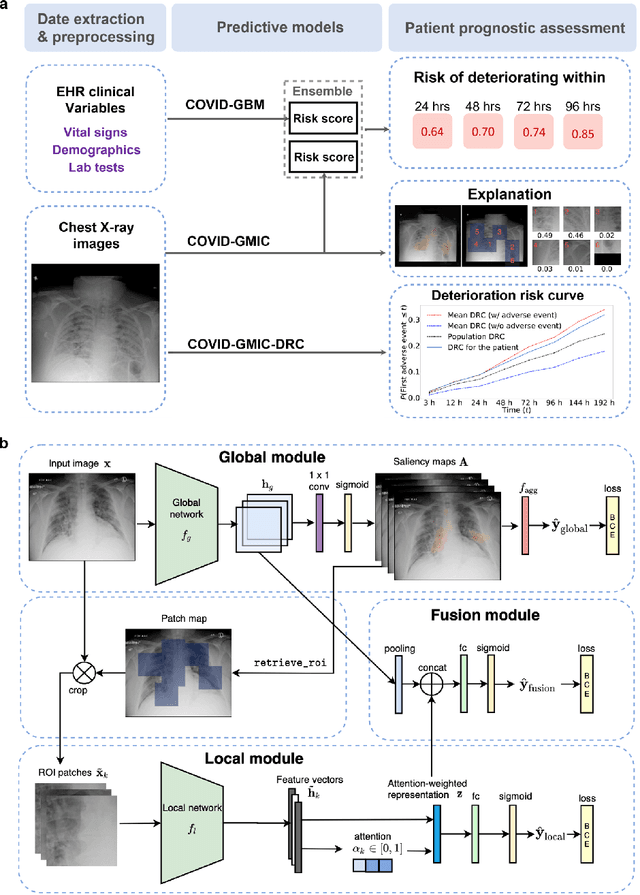
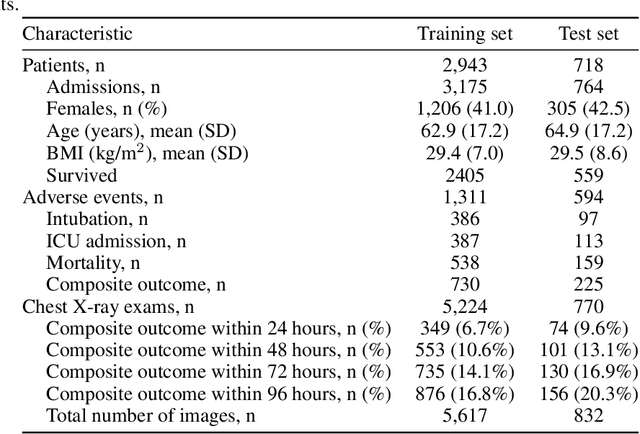
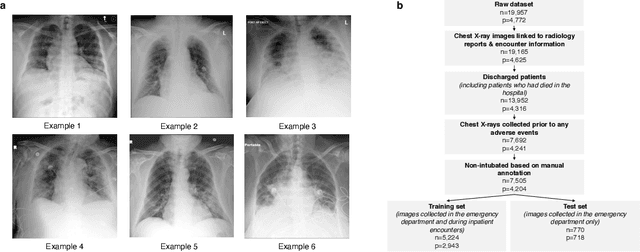
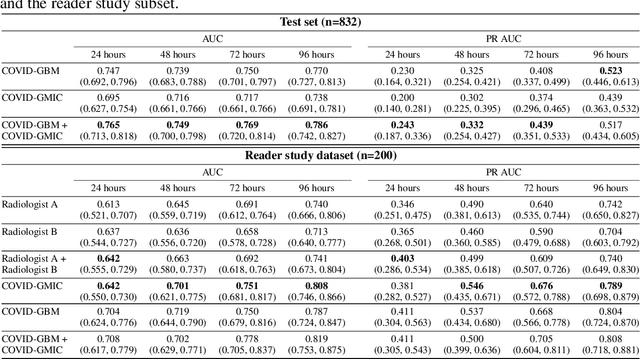
Abstract:During the COVID-19 pandemic, rapid and accurate triage of patients at the emergency department is critical to inform decision-making. We propose a data-driven approach for automatic prediction of deterioration risk using a deep neural network that learns from chest X-ray images, and a gradient boosting model that learns from routine clinical variables. Our AI prognosis system, trained using data from 3,661 patients, achieves an AUC of 0.786 (95% CI: 0.742-0.827) when predicting deterioration within 96 hours. The deep neural network extracts informative areas of chest X-ray images to assist clinicians in interpreting the predictions, and performs comparably to two radiologists in a reader study. In order to verify performance in a real clinical setting, we silently deployed a preliminary version of the deep neural network at NYU Langone Health during the first wave of the pandemic, which produced accurate predictions in real-time. In summary, our findings demonstrate the potential of the proposed system for assisting front-line physicians in the triage of COVID-19 patients.
Strong Uniform Consistency with Rates for Kernel Density Estimators with General Kernels on Manifolds
Jul 13, 2020Abstract:We provide a strong uniform consistency result with the convergence rate for the kernel density estimation on Riemannian manifolds with Riemann integrable kernels (in the ambient Euclidean space). We also provide a strong uniform consistency result for the kernel density estimation on Riemannian manifolds with Lebesgue integrable kernels. The kernels considered in this paper are different from the kernels in the Vapnik-Chervonenkis class that are frequently considered in statistics society. We illustrate the difference when we apply them to estimate probability density function. We also provide the necessary and sufficient condition for a kernel to be Riemann integrable on a submanifold in the Euclidean space.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge