Mohammadreza Armandpour
TiC-LM: A Web-Scale Benchmark for Time-Continual LLM Pretraining
Apr 02, 2025Abstract:Large Language Models (LLMs) trained on historical web data inevitably become outdated. We investigate evaluation strategies and update methods for LLMs as new data becomes available. We introduce a web-scale dataset for time-continual pretraining of LLMs derived from 114 dumps of Common Crawl (CC) - orders of magnitude larger than previous continual language modeling benchmarks. We also design time-stratified evaluations across both general CC data and specific domains (Wikipedia, StackExchange, and code documentation) to assess how well various continual learning methods adapt to new data while retaining past knowledge. Our findings demonstrate that, on general CC data, autoregressive meta-schedules combined with a fixed-ratio replay of older data can achieve comparable held-out loss to re-training from scratch, while requiring significantly less computation (2.6x). However, the optimal balance between incorporating new data and replaying old data differs as replay is crucial to avoid forgetting on generic web data but less so on specific domains.
Re-imagine the Negative Prompt Algorithm: Transform 2D Diffusion into 3D, alleviate Janus problem and Beyond
Apr 26, 2023



Abstract:Although text-to-image diffusion models have made significant strides in generating images from text, they are sometimes more inclined to generate images like the data on which the model was trained rather than the provided text. This limitation has hindered their usage in both 2D and 3D applications. To address this problem, we explored the use of negative prompts but found that the current implementation fails to produce desired results, particularly when there is an overlap between the main and negative prompts. To overcome this issue, we propose Perp-Neg, a new algorithm that leverages the geometrical properties of the score space to address the shortcomings of the current negative prompts algorithm. Perp-Neg does not require any training or fine-tuning of the model. Moreover, we experimentally demonstrate that Perp-Neg provides greater flexibility in generating images by enabling users to edit out unwanted concepts from the initially generated images in 2D cases. Furthermore, to extend the application of Perp-Neg to 3D, we conducted a thorough exploration of how Perp-Neg can be used in 2D to condition the diffusion model to generate desired views, rather than being biased toward the canonical views. Finally, we applied our 2D intuition to integrate Perp-Neg with the state-of-the-art text-to-3D (DreamFusion) method, effectively addressing its Janus (multi-head) problem. Our project page is available at https://Perp-Neg.github.io/
Gluformer: Transformer-Based Personalized Glucose Forecasting with Uncertainty Quantification
Sep 09, 2022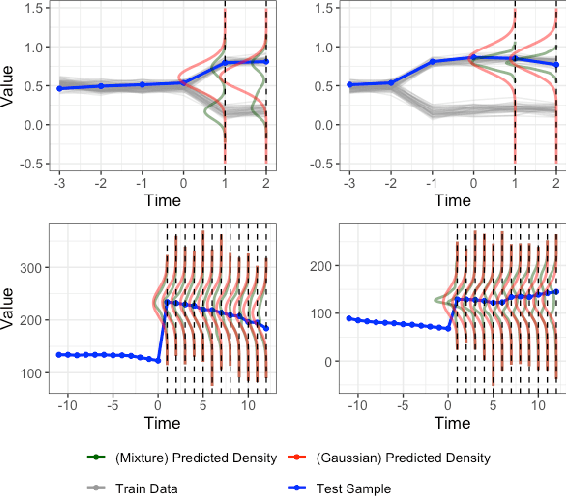
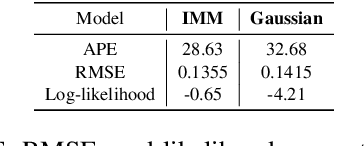
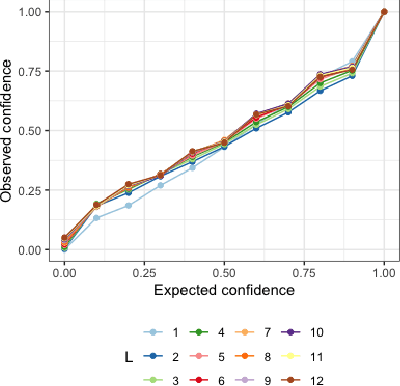
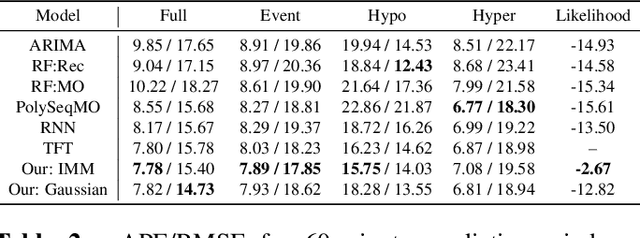
Abstract:Deep learning models achieve state-of-the art results in predicting blood glucose trajectories, with a wide range of architectures being proposed. However, the adaptation of such models in clinical practice is slow, largely due to the lack of uncertainty quantification of provided predictions. In this work, we propose to model the future glucose trajectory conditioned on the past as an infinite mixture of basis distributions (i.e., Gaussian, Laplace, etc.). This change allows us to learn the uncertainty and predict more accurately in the cases when the trajectory has a heterogeneous or multi-modal distribution. To estimate the parameters of the predictive distribution, we utilize the Transformer architecture. We empirically demonstrate the superiority of our method over existing state-of-the-art techniques both in terms of accuracy and uncertainty on the synthetic and benchmark glucose data sets.
Bayesian Graph Contrastive Learning
Jan 22, 2022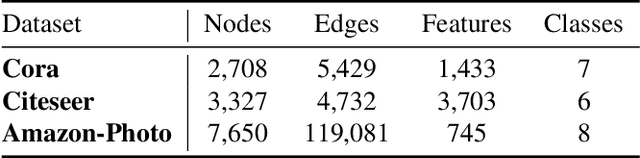
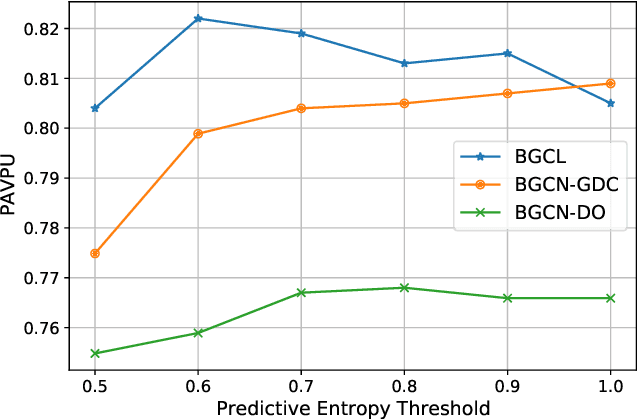
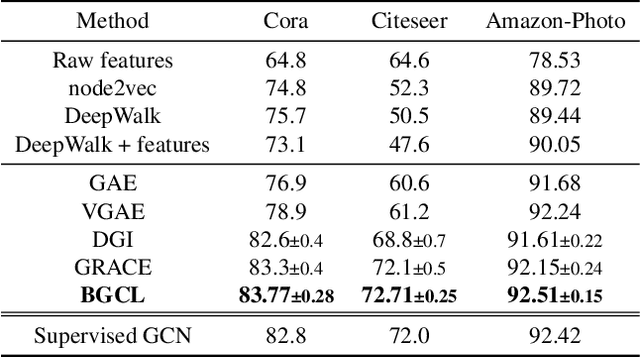
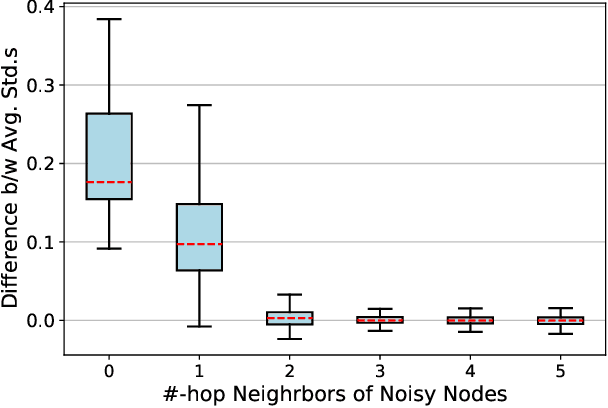
Abstract:Contrastive learning has become a key component of self-supervised learning approaches for graph-structured data. However, despite their success, existing graph contrastive learning methods are incapable of uncertainty quantification for node representations or their downstream tasks, limiting their application in high-stakes domains. In this paper, we propose a novel Bayesian perspective of graph contrastive learning methods showing random augmentations leads to stochastic encoders. As a result, our proposed method represents each node by a distribution in the latent space in contrast to existing techniques which embed each node to a deterministic vector. By learning distributional representations, we provide uncertainty estimates in downstream graph analytics tasks and increase the expressive power of the predictive model. In addition, we propose a Bayesian framework to infer the probability of perturbations in each view of the contrastive model, eliminating the need for a computationally expensive search for hyperparameter tuning. We empirically show a considerable improvement in performance compared to existing state-of-the-art methods on several benchmark datasets.
Synt++: Utilizing Imperfect Synthetic Data to Improve Speech Recognition
Oct 21, 2021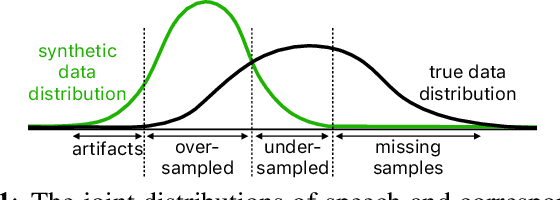
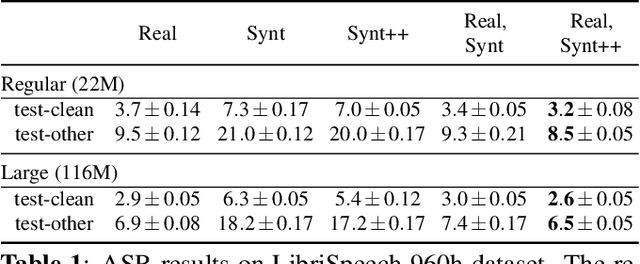
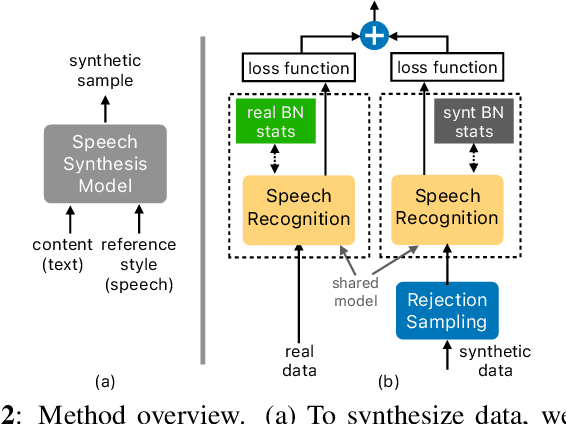
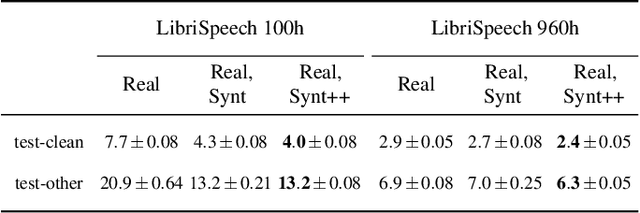
Abstract:With recent advances in speech synthesis, synthetic data is becoming a viable alternative to real data for training speech recognition models. However, machine learning with synthetic data is not trivial due to the gap between the synthetic and the real data distributions. Synthetic datasets may contain artifacts that do not exist in real data such as structured noise, content errors, or unrealistic speaking styles. Moreover, the synthesis process may introduce a bias due to uneven sampling of the data manifold. We propose two novel techniques during training to mitigate the problems due to the distribution gap: (i) a rejection sampling algorithm and (ii) using separate batch normalization statistics for the real and the synthetic samples. We show that these methods significantly improve the training of speech recognition models using synthetic data. We evaluate the proposed approach on keyword detection and Automatic Speech Recognition (ASR) tasks, and observe up to 18% and 13% relative error reduction, respectively, compared to naively using the synthetic data.
Deep Spatio-Temporal Wind Power Forecasting
Oct 07, 2021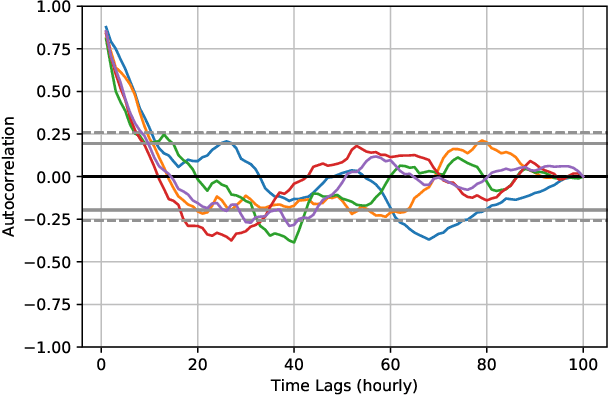
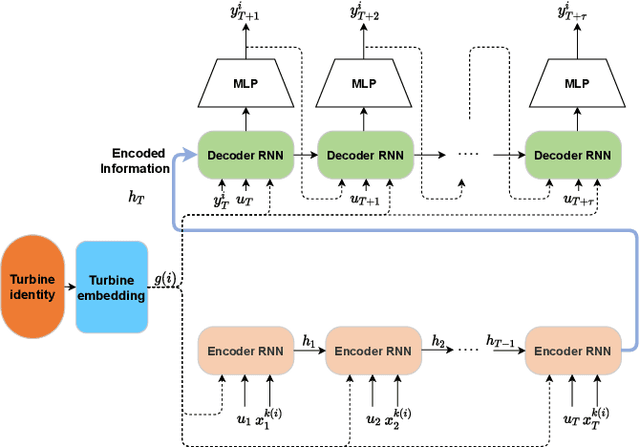

Abstract:Wind power forecasting has drawn increasing attention among researchers as the consumption of renewable energy grows. In this paper, we develop a deep learning approach based on encoder-decoder structure. Our model forecasts wind power generated by a wind turbine using its spatial location relative to other turbines and historical wind speed data. In this way, we effectively integrate spatial dependency and temporal trends to make turbine-specific predictions. The advantages of our method over existing work can be summarized as 1) it directly predicts wind power based on historical wind speed, without the need for prediction of wind speed first, and then using a transformation; 2) it can effectively capture long-term dependency 3) our model is more scalable and efficient compared with other deep learning based methods. We demonstrate the efficacy of our model on the benchmark real-world datasets.
Deep Personalized Glucose Level Forecasting Using Attention-based Recurrent Neural Networks
Jun 02, 2021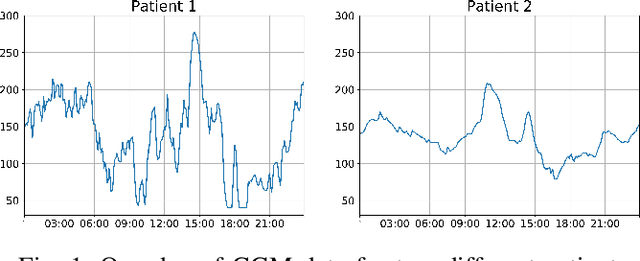
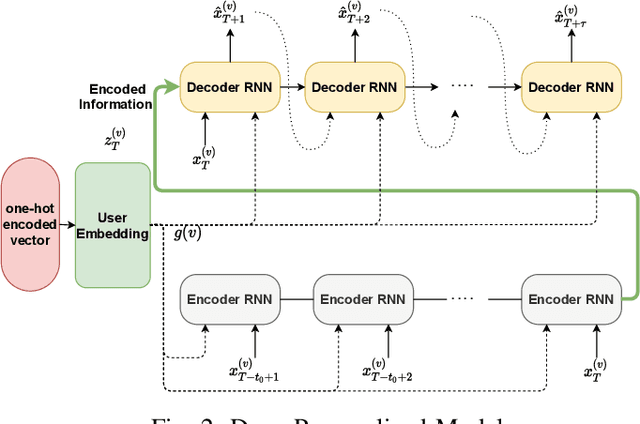
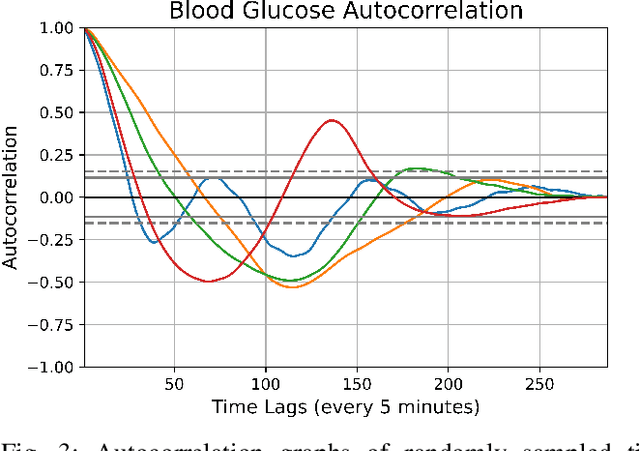
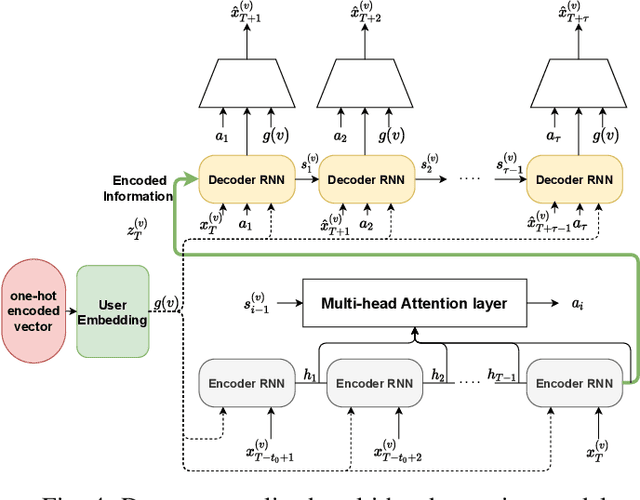
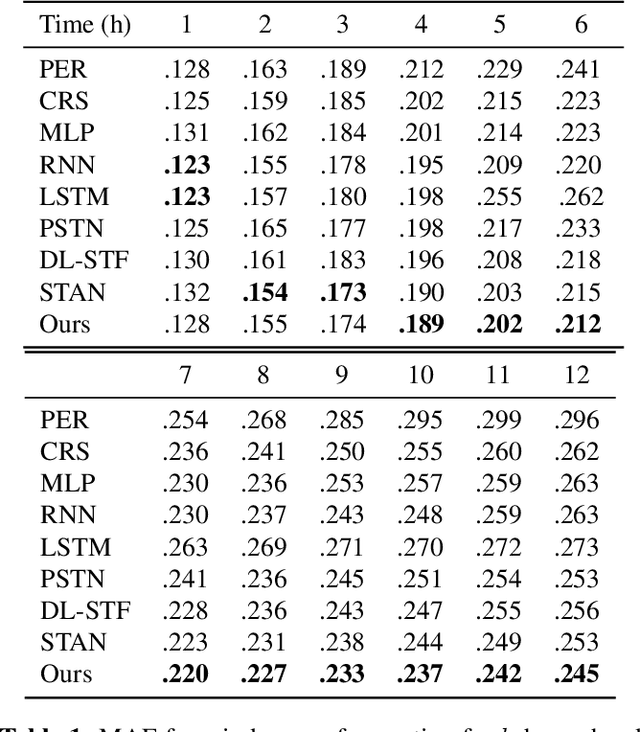
Abstract:In this paper, we study the problem of blood glucose forecasting and provide a deep personalized solution. Predicting blood glucose level in people with diabetes has significant value because health complications of abnormal glucose level are serious, sometimes even leading to death. Therefore, having a model that can accurately and quickly warn patients of potential problems is essential. To develop a better deep model for blood glucose forecasting, we analyze the data and detect important patterns. These observations helped us to propose a method that has several key advantages over existing methods: 1- it learns a personalized model for each patient as well as a global model; 2- it uses an attention mechanism and extracted time features to better learn long-term dependencies in the data; 3- it introduces a new, robust training procedure for time series data. We empirically show the efficacy of our model on a real dataset.
Partition-Guided GANs
Apr 02, 2021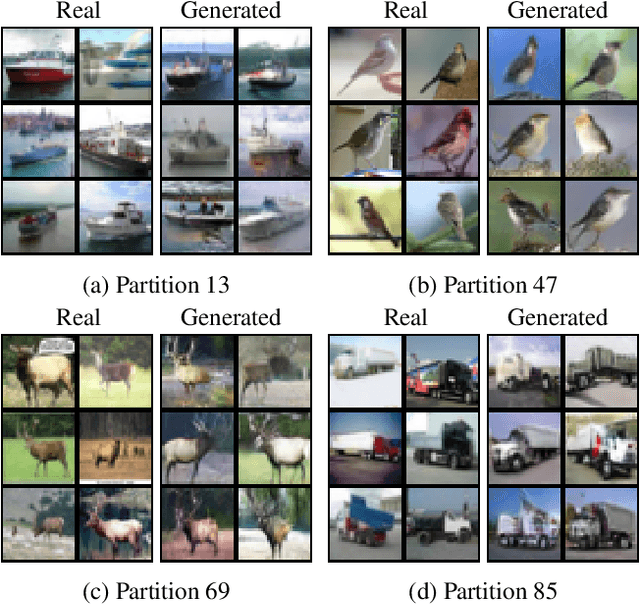
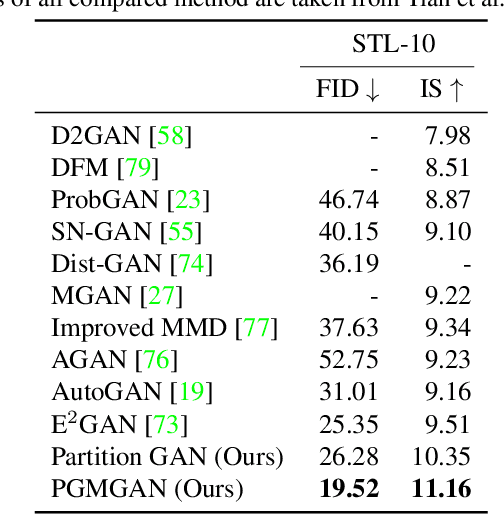
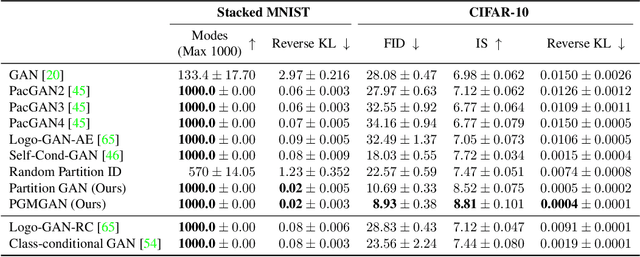
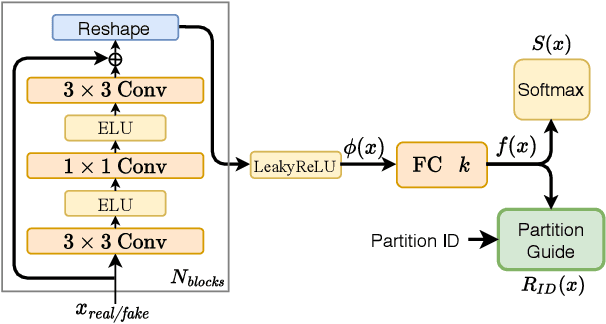

Abstract:Despite the success of Generative Adversarial Networks (GANs), their training suffers from several well-known problems, including mode collapse and difficulties learning a disconnected set of manifolds. In this paper, we break down the challenging task of learning complex high dimensional distributions, supporting diverse data samples, to simpler sub-tasks. Our solution relies on designing a partitioner that breaks the space into smaller regions, each having a simpler distribution, and training a different generator for each partition. This is done in an unsupervised manner without requiring any labels. We formulate two desired criteria for the space partitioner that aid the training of our mixture of generators: 1) to produce connected partitions and 2) provide a proxy of distance between partitions and data samples, along with a direction for reducing that distance. These criteria are developed to avoid producing samples from places with non-existent data density, and also facilitate training by providing additional direction to the generators. We develop theoretical constraints for a space partitioner to satisfy the above criteria. Guided by our theoretical analysis, we design an effective neural architecture for the space partitioner that empirically assures these conditions. Experimental results on various standard benchmarks show that the proposed unsupervised model outperforms several recent methods.
ChronoR: Rotation Based Temporal Knowledge Graph Embedding
Mar 18, 2021
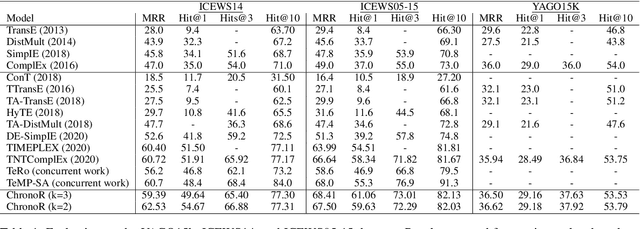
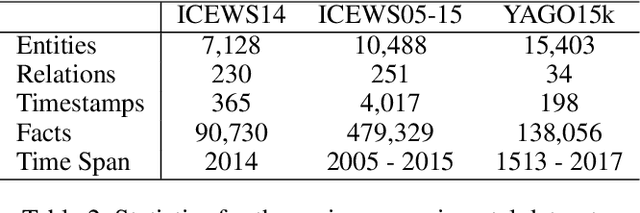

Abstract:Despite the importance and abundance of temporal knowledge graphs, most of the current research has been focused on reasoning on static graphs. In this paper, we study the challenging problem of inference over temporal knowledge graphs. In particular, the task of temporal link prediction. In general, this is a difficult task due to data non-stationarity, data heterogeneity, and its complex temporal dependencies. We propose Chronological Rotation embedding (ChronoR), a novel model for learning representations for entities, relations, and time. Learning dense representations is frequently used as an efficient and versatile method to perform reasoning on knowledge graphs. The proposed model learns a k-dimensional rotation transformation parametrized by relation and time, such that after each fact's head entity is transformed using the rotation, it falls near its corresponding tail entity. By using high dimensional rotation as its transformation operator, ChronoR captures rich interaction between the temporal and multi-relational characteristics of a Temporal Knowledge Graph. Experimentally, we show that ChronoR is able to outperform many of the state-of-the-art methods on the benchmark datasets for temporal knowledge graph link prediction.
Convex Polytope Trees
Oct 21, 2020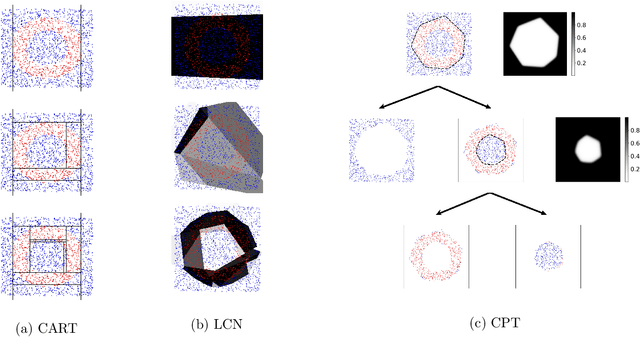

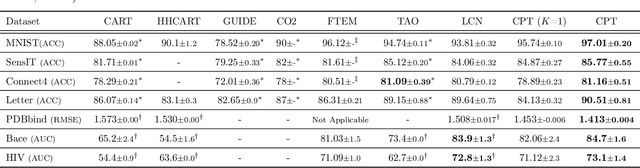
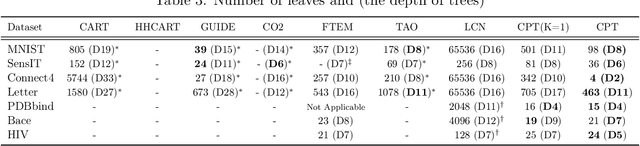
Abstract:A decision tree is commonly restricted to use a single hyperplane to split the covariate space at each of its internal nodes. It often requires a large number of nodes to achieve high accuracy, hurting its interpretability. In this paper, we propose convex polytope trees (CPT) to expand the family of decision trees by an interpretable generalization of their decision boundary. The splitting function at each node of CPT is based on the logical disjunction of a community of differently weighted probabilistic linear decision-makers, which also geometrically corresponds to a convex polytope in the covariate space. We use a nonparametric Bayesian prior at each node to infer the community's size, encouraging simpler decision boundaries by shrinking the number of polytope facets. We develop a greedy method to efficiently construct CPT and scalable end-to-end training algorithms for the tree parameters when the tree structure is given. We empirically demonstrate the efficiency of CPT over existing state-of-the-art decision trees in several real-world classification and regression tasks from diverse domains.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge