Marc Niethammer
CARES: A Comprehensive Benchmark of Trustworthiness in Medical Vision Language Models
Jun 10, 2024



Abstract:Artificial intelligence has significantly impacted medical applications, particularly with the advent of Medical Large Vision Language Models (Med-LVLMs), sparking optimism for the future of automated and personalized healthcare. However, the trustworthiness of Med-LVLMs remains unverified, posing significant risks for future model deployment. In this paper, we introduce CARES and aim to comprehensively evaluate the Trustworthiness of Med-LVLMs across the medical domain. We assess the trustworthiness of Med-LVLMs across five dimensions, including trustfulness, fairness, safety, privacy, and robustness. CARES comprises about 41K question-answer pairs in both closed and open-ended formats, covering 16 medical image modalities and 27 anatomical regions. Our analysis reveals that the models consistently exhibit concerns regarding trustworthiness, often displaying factual inaccuracies and failing to maintain fairness across different demographic groups. Furthermore, they are vulnerable to attacks and demonstrate a lack of privacy awareness. We publicly release our benchmark and code in https://github.com/richard-peng-xia/CARES.
CARL: A Framework for Equivariant Image Registration
May 28, 2024



Abstract:Image registration estimates spatial correspondences between a pair of images. These estimates are typically obtained via numerical optimization or regression by a deep network. A desirable property of such estimators is that a correspondence estimate (e.g., the true oracle correspondence) for an image pair is maintained under deformations of the input images. Formally, the estimator should be equivariant to a desired class of image transformations. In this work, we present careful analyses of the desired equivariance properties in the context of multi-step deep registration networks. Based on these analyses we 1) introduce the notions of $[U,U]$ equivariance (network equivariance to the same deformations of the input images) and $[W,U]$ equivariance (where input images can undergo different deformations); we 2) show that in a suitable multi-step registration setup it is sufficient for overall $[W,U]$ equivariance if the first step has $[W,U]$ equivariance and all others have $[U,U]$ equivariance; we 3) show that common displacement-predicting networks only exhibit $[U,U]$ equivariance to translations instead of the more powerful $[W,U]$ equivariance; and we 4) show how to achieve multi-step $[W,U]$ equivariance via a coordinate-attention mechanism combined with displacement-predicting refinement layers (CARL). Overall, our approach obtains excellent practical registration performance on several 3D medical image registration tasks and outperforms existing unsupervised approaches for the challenging problem of abdomen registration.
Rethinking Interactive Image Segmentation with Low Latency, High Quality, and Diverse Prompts
Mar 31, 2024Abstract:The goal of interactive image segmentation is to delineate specific regions within an image via visual or language prompts. Low-latency and high-quality interactive segmentation with diverse prompts remain challenging for existing specialist and generalist models. Specialist models, with their limited prompts and task-specific designs, experience high latency because the image must be recomputed every time the prompt is updated, due to the joint encoding of image and visual prompts. Generalist models, exemplified by the Segment Anything Model (SAM), have recently excelled in prompt diversity and efficiency, lifting image segmentation to the foundation model era. However, for high-quality segmentations, SAM still lags behind state-of-the-art specialist models despite SAM being trained with x100 more segmentation masks. In this work, we delve deep into the architectural differences between the two types of models. We observe that dense representation and fusion of visual prompts are the key design choices contributing to the high segmentation quality of specialist models. In light of this, we reintroduce this dense design into the generalist models, to facilitate the development of generalist models with high segmentation quality. To densely represent diverse visual prompts, we propose to use a dense map to capture five types: clicks, boxes, polygons, scribbles, and masks. Thus, we propose SegNext, a next-generation interactive segmentation approach offering low latency, high quality, and diverse prompt support. Our method outperforms current state-of-the-art methods on HQSeg-44K and DAVIS, both quantitatively and qualitatively.
Leveraging Near-Field Lighting for Monocular Depth Estimation from Endoscopy Videos
Mar 26, 2024



Abstract:Monocular depth estimation in endoscopy videos can enable assistive and robotic surgery to obtain better coverage of the organ and detection of various health issues. Despite promising progress on mainstream, natural image depth estimation, techniques perform poorly on endoscopy images due to a lack of strong geometric features and challenging illumination effects. In this paper, we utilize the photometric cues, i.e., the light emitted from an endoscope and reflected by the surface, to improve monocular depth estimation. We first create two novel loss functions with supervised and self-supervised variants that utilize a per-pixel shading representation. We then propose a novel depth refinement network (PPSNet) that leverages the same per-pixel shading representation. Finally, we introduce teacher-student transfer learning to produce better depth maps from both synthetic data with supervision and clinical data with self-supervision. We achieve state-of-the-art results on the C3VD dataset while estimating high-quality depth maps from clinical data. Our code, pre-trained models, and supplementary materials can be found on our project page: https://ppsnet.github.io/
A Unified Model for Longitudinal Multi-Modal Multi-View Prediction with Missingness
Mar 22, 2024



Abstract:Medical records often consist of different modalities, such as images, text, and tabular information. Integrating all modalities offers a holistic view of a patient's condition, while analyzing them longitudinally provides a better understanding of disease progression. However, real-world longitudinal medical records present challenges: 1) patients may lack some or all of the data for a specific timepoint, and 2) certain modalities or views might be absent for all patients during a particular period. In this work, we introduce a unified model for longitudinal multi-modal multi-view prediction with missingness. Our method allows as many timepoints as desired for input, and aims to leverage all available data, regardless of their availability. We conduct extensive experiments on the knee osteoarthritis dataset from the Osteoarthritis Initiative for pain and Kellgren-Lawrence grade prediction at a future timepoint. We demonstrate the effectiveness of our method by comparing results from our unified model to specific models that use the same modality and view combinations during training and evaluation. We also show the benefit of having extended temporal data and provide post-hoc analysis for a deeper understanding of each modality/view's importance for different tasks.
NeuralOCT: Airway OCT Analysis via Neural Fields
Mar 15, 2024



Abstract:Optical coherence tomography (OCT) is a popular modality in ophthalmology and is also used intravascularly. Our interest in this work is OCT in the context of airway abnormalities in infants and children where the high resolution of OCT and the fact that it is radiation-free is important. The goal of airway OCT is to provide accurate estimates of airway geometry (in 2D and 3D) to assess airway abnormalities such as subglottic stenosis. We propose $\texttt{NeuralOCT}$, a learning-based approach to process airway OCT images. Specifically, $\texttt{NeuralOCT}$ extracts 3D geometries from OCT scans by robustly bridging two steps: point cloud extraction via 2D segmentation and 3D reconstruction from point clouds via neural fields. Our experiments show that $\texttt{NeuralOCT}$ produces accurate and robust 3D airway reconstructions with an average A-line error smaller than 70 micrometer. Our code will cbe available on GitHub.
uniGradICON: A Foundation Model for Medical Image Registration
Mar 09, 2024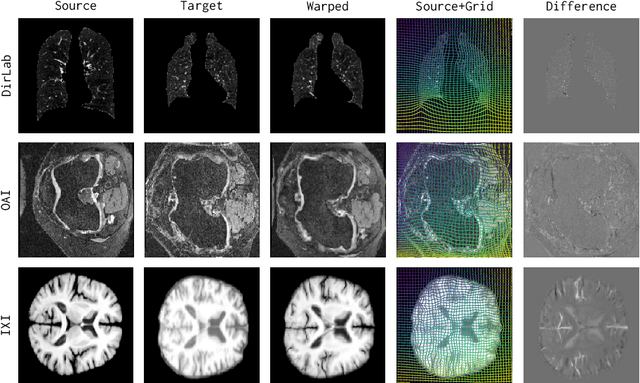
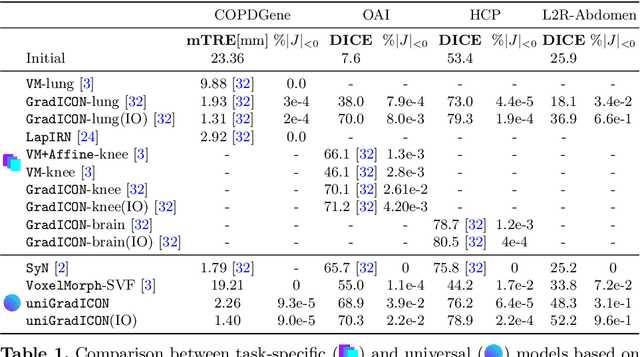

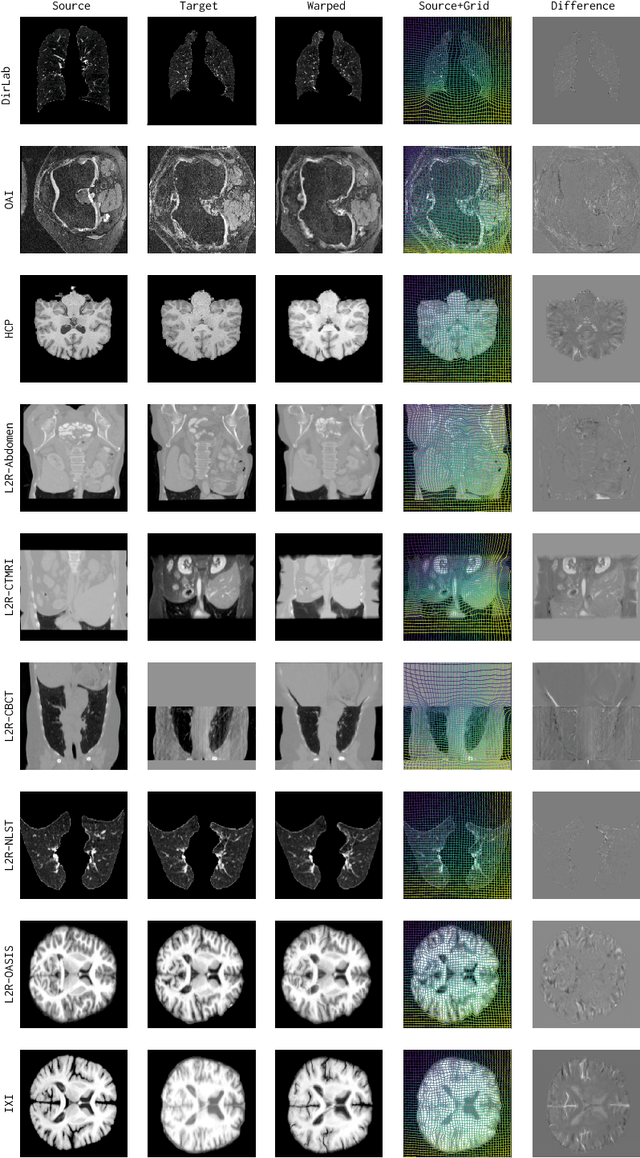
Abstract:Conventional medical image registration approaches directly optimize over the parameters of a transformation model. These approaches have been highly successful and are used generically for registrations of different anatomical regions. Recent deep registration networks are incredibly fast and accurate but are only trained for specific tasks. Hence, they are no longer generic registration approaches. We therefore propose uniGradICON, a first step toward a foundation model for registration providing 1) great performance \emph{across} multiple datasets which is not feasible for current learning-based registration methods, 2) zero-shot capabilities for new registration tasks suitable for different acquisitions, anatomical regions, and modalities compared to the training dataset, and 3) a strong initialization for finetuning on out-of-distribution registration tasks. UniGradICON unifies the speed and accuracy benefits of learning-based registration algorithms with the generic applicability of conventional non-deep-learning approaches. We extensively trained and evaluated uniGradICON on twelve different public datasets. Our code and the uniGradICON model are available at https://github.com/uncbiag/uniGradICON.
SAME++: A Self-supervised Anatomical eMbeddings Enhanced medical image registration framework using stable sampling and regularized transformation
Nov 25, 2023Abstract:Image registration is a fundamental medical image analysis task. Ideally, registration should focus on aligning semantically corresponding voxels, i.e., the same anatomical locations. However, existing methods often optimize similarity measures computed directly on intensities or on hand-crafted features, which lack anatomical semantic information. These similarity measures may lead to sub-optimal solutions where large deformations, complex anatomical differences, or cross-modality imagery exist. In this work, we introduce a fast and accurate method for unsupervised 3D medical image registration building on top of a Self-supervised Anatomical eMbedding (SAM) algorithm, which is capable of computing dense anatomical correspondences between two images at the voxel level. We name our approach SAM-Enhanced registration (SAME++), which decomposes image registration into four steps: affine transformation, coarse deformation, deep non-parametric transformation, and instance optimization. Using SAM embeddings, we enhance these steps by finding more coherent correspondence and providing features with better semantic guidance. We extensively evaluated SAME++ using more than 50 labeled organs on three challenging inter-subject registration tasks of different body parts. As a complete registration framework, SAME++ markedly outperforms leading methods by $4.2\%$ - $8.2\%$ in terms of Dice score while being orders of magnitude faster than numerical optimization-based methods. Code is available at \url{https://github.com/alibaba-damo-academy/same}.
Joint Depth Prediction and Semantic Segmentation with Multi-View SAM
Oct 31, 2023



Abstract:Multi-task approaches to joint depth and segmentation prediction are well-studied for monocular images. Yet, predictions from a single-view are inherently limited, while multiple views are available in many robotics applications. On the other end of the spectrum, video-based and full 3D methods require numerous frames to perform reconstruction and segmentation. With this work we propose a Multi-View Stereo (MVS) technique for depth prediction that benefits from rich semantic features of the Segment Anything Model (SAM). This enhanced depth prediction, in turn, serves as a prompt to our Transformer-based semantic segmentation decoder. We report the mutual benefit that both tasks enjoy in our quantitative and qualitative studies on the ScanNet dataset. Our approach consistently outperforms single-task MVS and segmentation models, along with multi-task monocular methods.
$\texttt{NePhi}$: Neural Deformation Fields for Approximately Diffeomorphic Medical Image Registration
Sep 13, 2023



Abstract:This work proposes $\texttt{NePhi}$, a neural deformation model which results in approximately diffeomorphic transformations. In contrast to the predominant voxel-based approaches, $\texttt{NePhi}$ represents deformations functionally which allows for memory-efficient training and inference. This is of particular importance for large volumetric registrations. Further, while medical image registration approaches representing transformation maps via multi-layer perceptrons have been proposed, $\texttt{NePhi}$ facilitates both pairwise optimization-based registration $\textit{as well as}$ learning-based registration via predicted or optimized global and local latent codes. Lastly, as deformation regularity is a highly desirable property for most medical image registration tasks, $\texttt{NePhi}$ makes use of gradient inverse consistency regularization which empirically results in approximately diffeomorphic transformations. We show the performance of $\texttt{NePhi}$ on two 2D synthetic datasets as well as on real 3D lung registration. Our results show that $\texttt{NePhi}$ can achieve similar accuracies as voxel-based representations in a single-resolution registration setting while using less memory and allowing for faster instance-optimization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge