Krzysztof J. Geras
Globally-Aware Multiple Instance Classifier for Breast Cancer Screening
Jun 07, 2019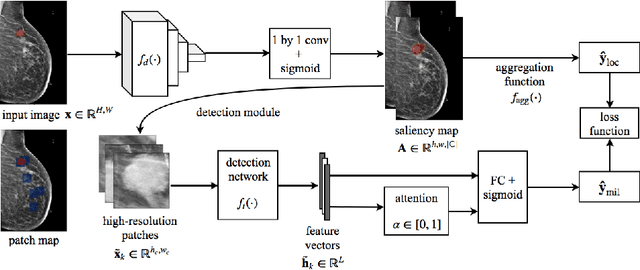
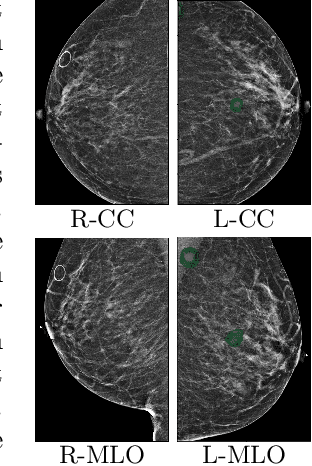
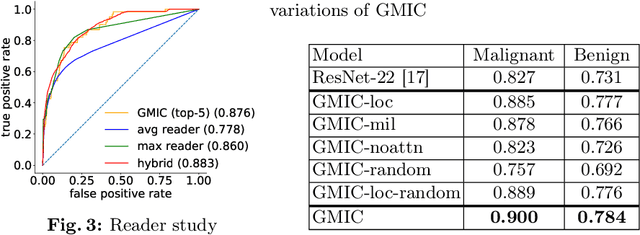
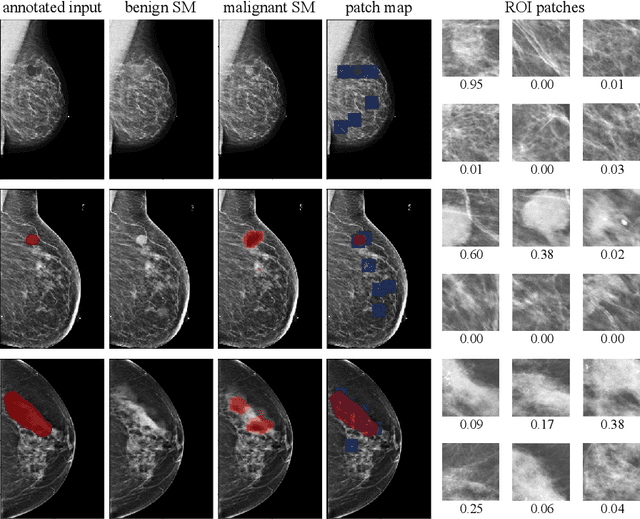
Abstract:Deep learning models designed for visual classification tasks on natural images have become prevalent in medical image analysis. However, medical images differ from typical natural images in many ways, such as significantly higher resolutions and smaller regions of interest. Moreover, both the global structure and local details play important roles in medical image analysis tasks. To address these unique properties of medical images, we propose a neural network that is able to classify breast cancer lesions utilizing information from both a global saliency map and multiple local patches. The proposed model outperforms the ResNet-based baseline and achieves radiologist-level performance in the interpretation of screening mammography. Although our model is trained only with image-level labels, it is able to generate pixel-level saliency maps that provide localization of possible malignant findings.
Deep Neural Networks Improve Radiologists' Performance in Breast Cancer Screening
Mar 20, 2019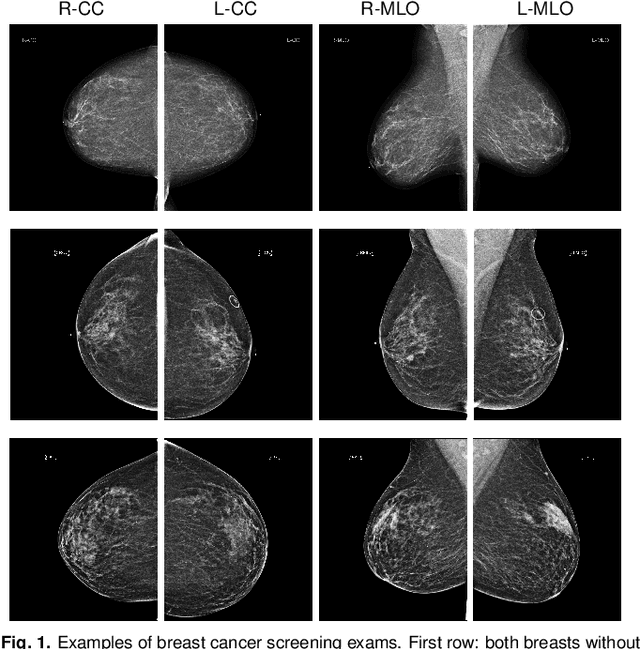
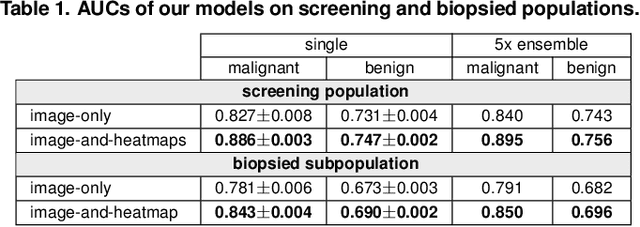
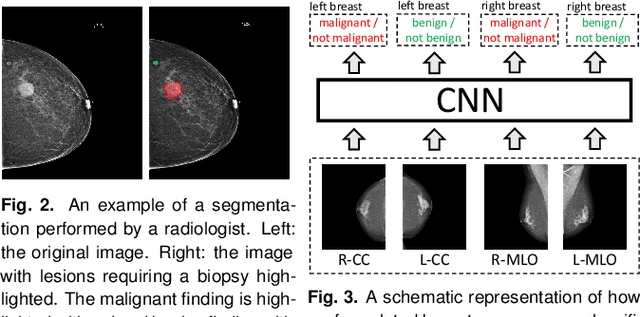
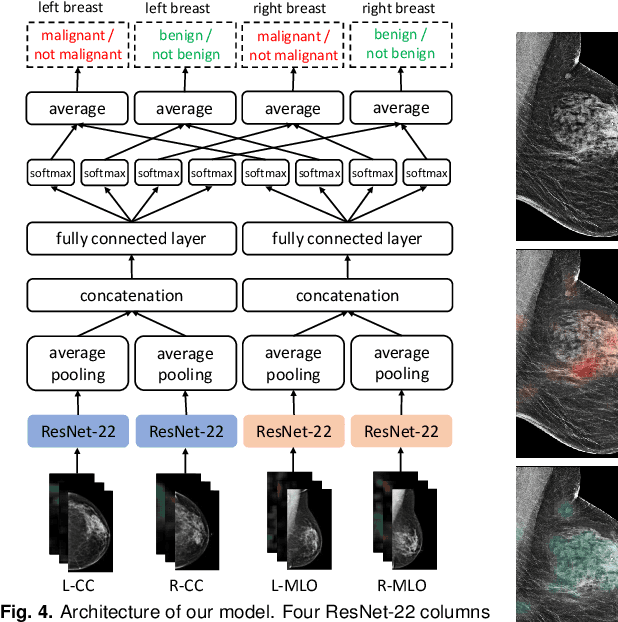
Abstract:We present a deep convolutional neural network for breast cancer screening exam classification, trained and evaluated on over 200,000 exams (over 1,000,000 images). Our network achieves an AUC of 0.895 in predicting whether there is a cancer in the breast, when tested on the screening population. We attribute the high accuracy of our model to a two-stage training procedure, which allows us to use a very high-capacity patch-level network to learn from pixel-level labels alongside a network learning from macroscopic breast-level labels. To validate our model, we conducted a reader study with 14 readers, each reading 720 screening mammogram exams, and find our model to be as accurate as experienced radiologists when presented with the same data. Finally, we show that a hybrid model, averaging probability of malignancy predicted by a radiologist with a prediction of our neural network, is more accurate than either of the two separately. To better understand our results, we conduct a thorough analysis of our network's performance on different subpopulations of the screening population, model design, training procedure, errors, and properties of its internal representations.
fastMRI: An Open Dataset and Benchmarks for Accelerated MRI
Nov 21, 2018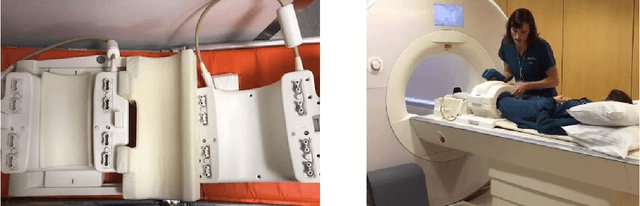
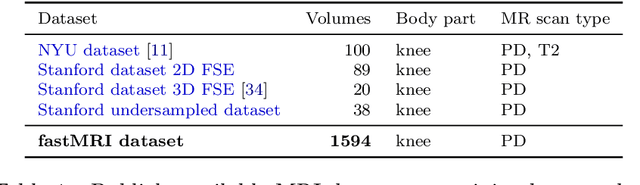
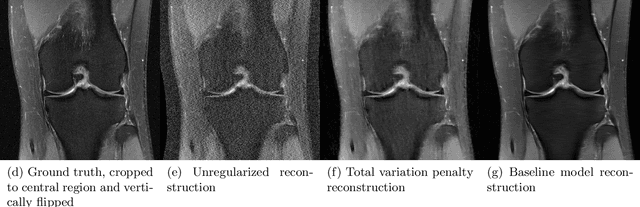
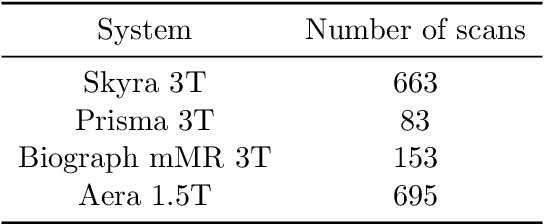
Abstract:Accelerating Magnetic Resonance Imaging (MRI) by taking fewer measurements has the potential to reduce medical costs, minimize stress to patients and make MRI possible in applications where it is currently prohibitively slow or expensive. We introduce the fastMRI dataset, a large-scale collection of both raw MR measurements and clinical MR images, that can be used for training and evaluation of machine-learning approaches to MR image reconstruction. By introducing standardized evaluation criteria and a freely-accessible dataset, our goal is to help the community make rapid advances in the state of the art for MR image reconstruction. We also provide a self-contained introduction to MRI for machine learning researchers with no medical imaging background.
Classifier-agnostic saliency map extraction
Oct 02, 2018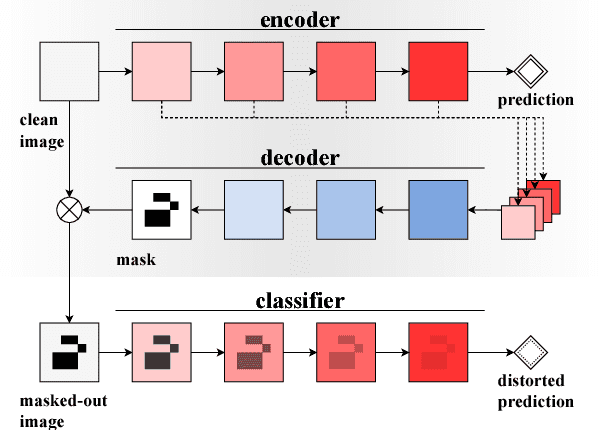
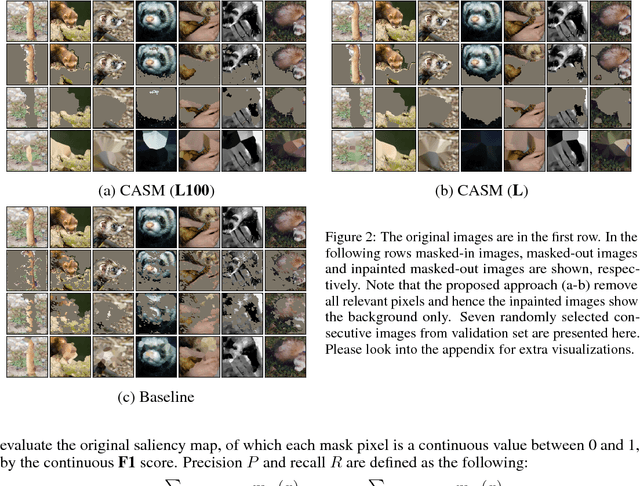
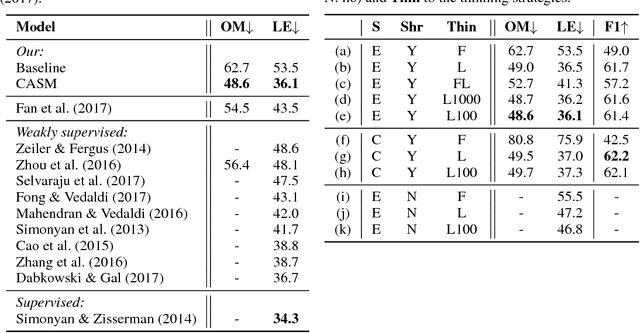

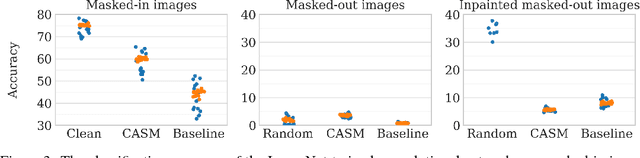
Abstract:Extracting saliency maps, which indicate parts of the image important to classification, requires many tricks to achieve satisfactory performance when using classifier-dependent methods. Instead, we propose classifier-agnostic saliency map extraction, which finds all parts of the image that any classifier could use, not just one given in advance. We observe that the proposed approach extracts higher quality saliency maps and outperforms existing weakly-supervised localization techniques, setting the new state of the art result on the ImageNet dataset. We made our code publicly available at https://github.com/kondiz/casme .
High-Resolution Breast Cancer Screening with Multi-View Deep Convolutional Neural Networks
Jun 28, 2018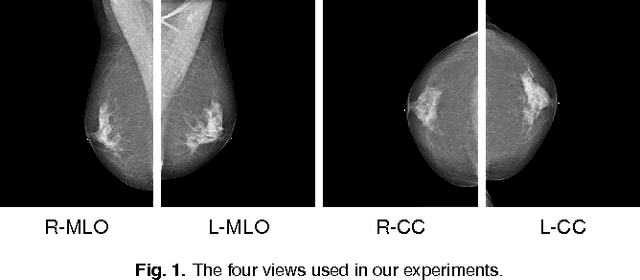
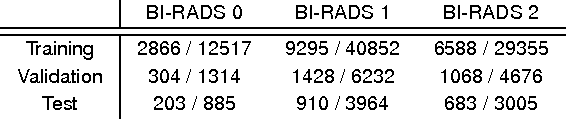
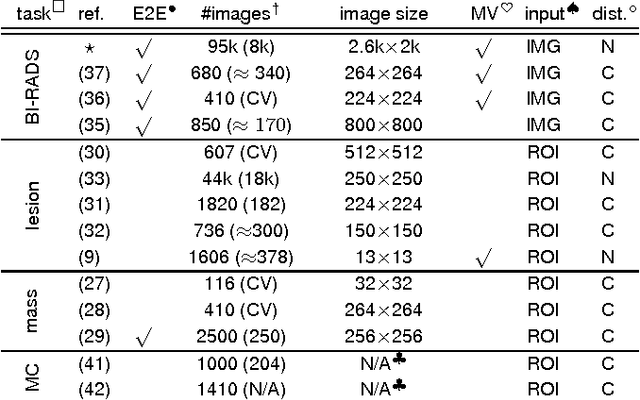
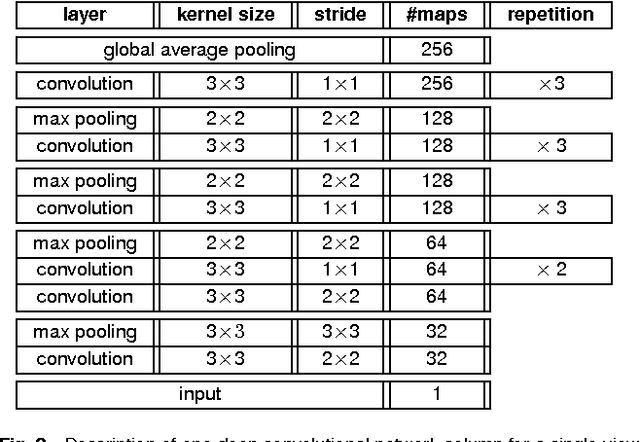

Abstract:Advances in deep learning for natural images have prompted a surge of interest in applying similar techniques to medical images. The majority of the initial attempts focused on replacing the input of a deep convolutional neural network with a medical image, which does not take into consideration the fundamental differences between these two types of images. Specifically, fine details are necessary for detection in medical images, unlike in natural images where coarse structures matter most. This difference makes it inadequate to use the existing network architectures developed for natural images, because they work on heavily downscaled images to reduce the memory requirements. This hides details necessary to make accurate predictions. Additionally, a single exam in medical imaging often comes with a set of views which must be fused in order to reach a correct conclusion. In our work, we propose to use a multi-view deep convolutional neural network that handles a set of high-resolution medical images. We evaluate it on large-scale mammography-based breast cancer screening (BI-RADS prediction) using 886,000 images. We focus on investigating the impact of the training set size and image size on the prediction accuracy. Our results highlight that performance increases with the size of training set, and that the best performance can only be achieved using the original resolution. In the reader study, performed on a random subset of the test set, we confirmed the efficacy of our model, which achieved performance comparable to a committee of radiologists when presented with the same data.
Breast density classification with deep convolutional neural networks
Nov 10, 2017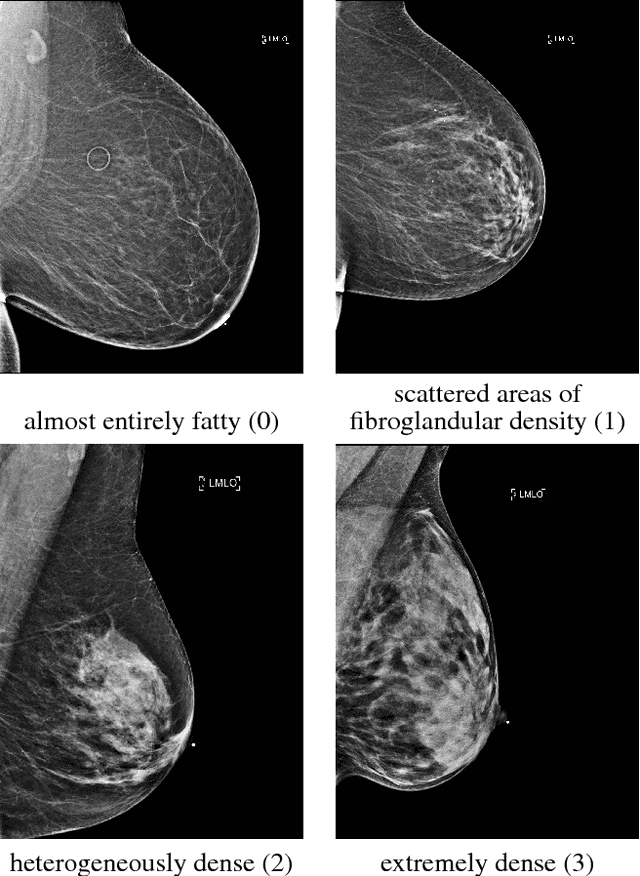
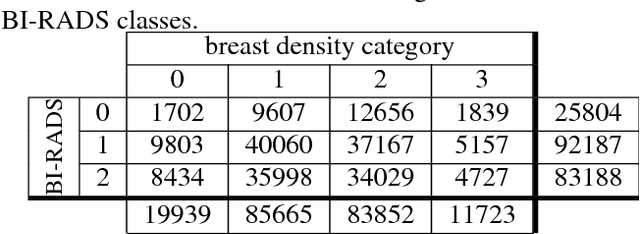
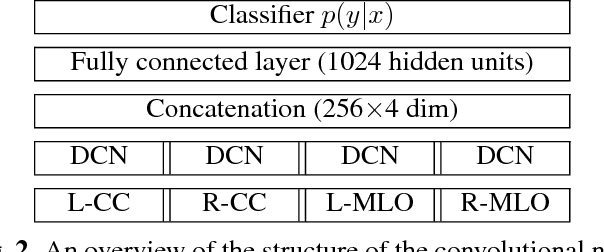
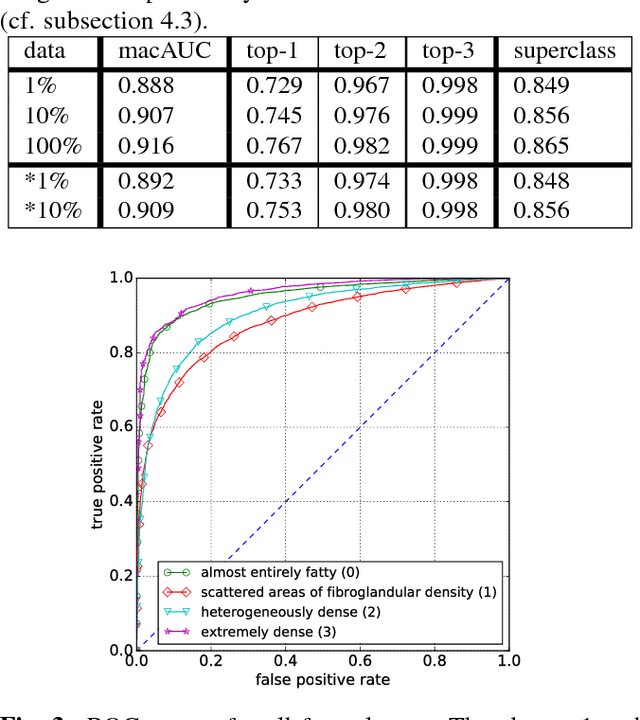
Abstract:Breast density classification is an essential part of breast cancer screening. Although a lot of prior work considered this problem as a task for learning algorithms, to our knowledge, all of them used small and not clinically realistic data both for training and evaluation of their models. In this work, we explore the limits of this task with a data set coming from over 200,000 breast cancer screening exams. We use this data to train and evaluate a strong convolutional neural network classifier. In a reader study, we find that our model can perform this task comparably to a human expert.
Do Deep Convolutional Nets Really Need to be Deep and Convolutional?
Mar 04, 2017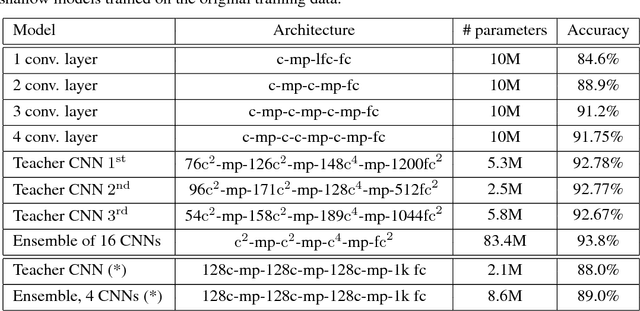
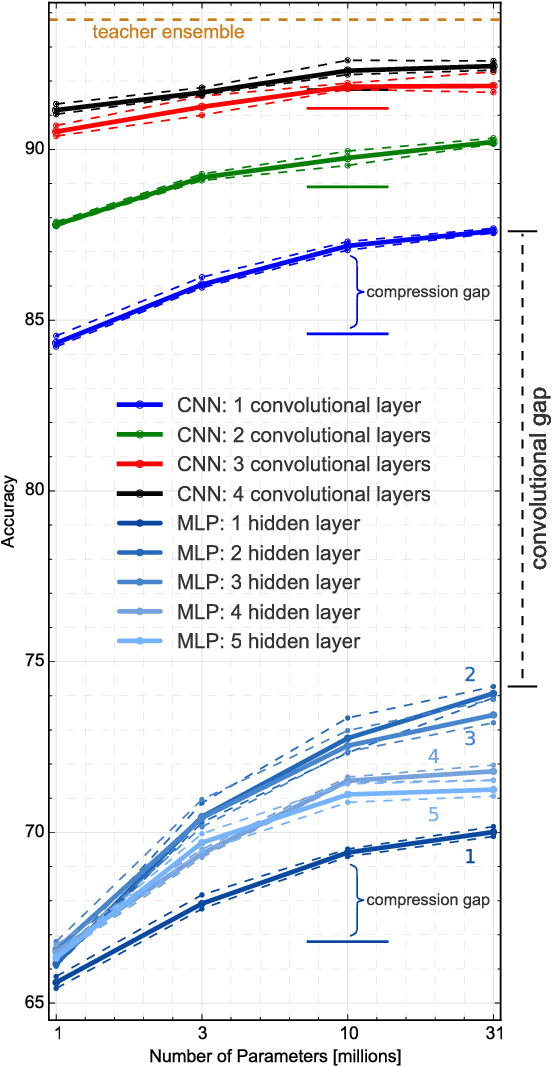
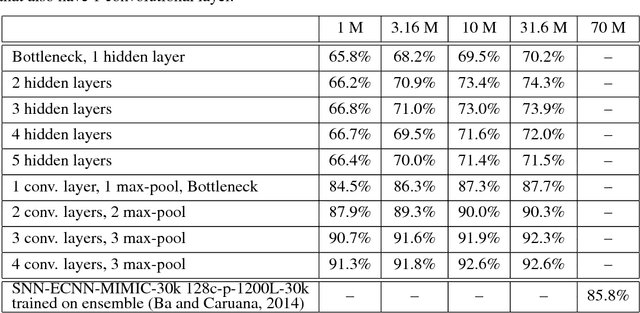
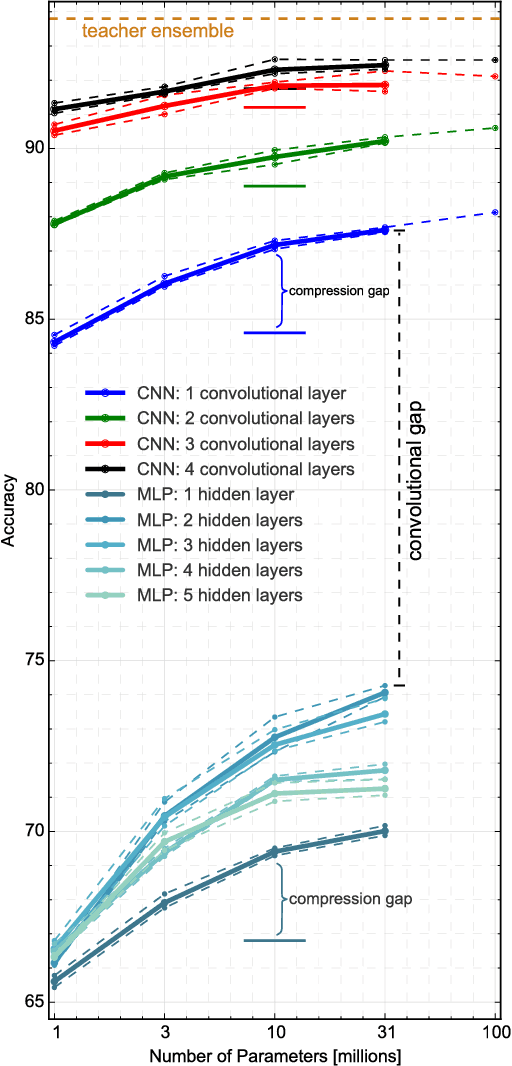
Abstract:Yes, they do. This paper provides the first empirical demonstration that deep convolutional models really need to be both deep and convolutional, even when trained with methods such as distillation that allow small or shallow models of high accuracy to be trained. Although previous research showed that shallow feed-forward nets sometimes can learn the complex functions previously learned by deep nets while using the same number of parameters as the deep models they mimic, in this paper we demonstrate that the same methods cannot be used to train accurate models on CIFAR-10 unless the student models contain multiple layers of convolution. Although the student models do not have to be as deep as the teacher model they mimic, the students need multiple convolutional layers to learn functions of comparable accuracy as the deep convolutional teacher.
Blending LSTMs into CNNs
Sep 14, 2016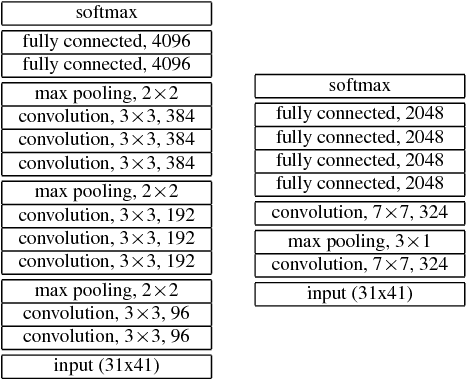
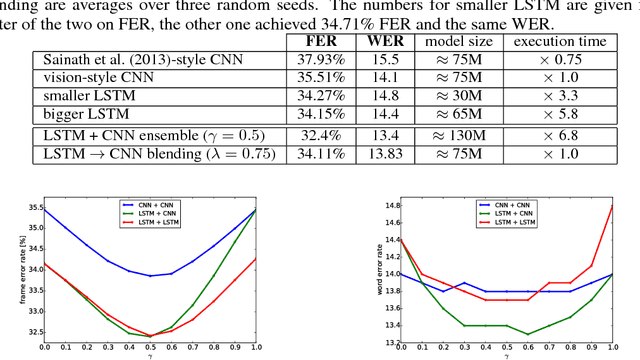
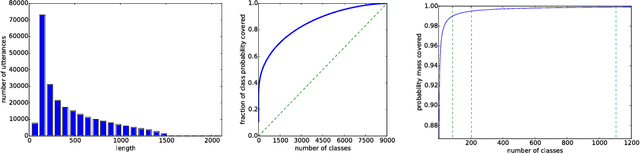

Abstract:We consider whether deep convolutional networks (CNNs) can represent decision functions with similar accuracy as recurrent networks such as LSTMs. First, we show that a deep CNN with an architecture inspired by the models recently introduced in image recognition can yield better accuracy than previous convolutional and LSTM networks on the standard 309h Switchboard automatic speech recognition task. Then we show that even more accurate CNNs can be trained under the guidance of LSTMs using a variant of model compression, which we call model blending because the teacher and student models are similar in complexity but different in inductive bias. Blending further improves the accuracy of our CNN, yielding a computationally efficient model of accuracy higher than any of the other individual models. Examining the effect of "dark knowledge" in this model compression task, we find that less than 1% of the highest probability labels are needed for accurate model compression.
Scheduled denoising autoencoders
Apr 10, 2015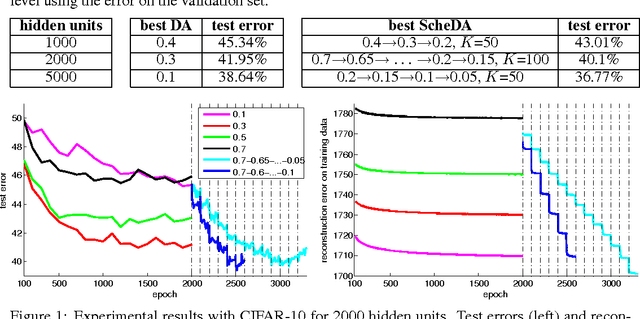
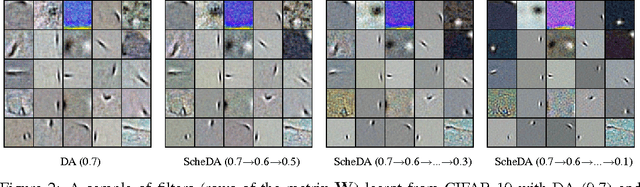
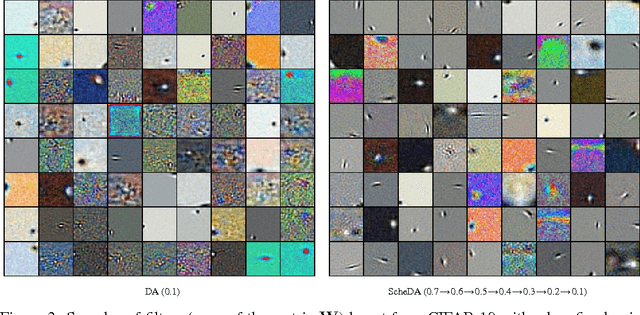



Abstract:We present a representation learning method that learns features at multiple different levels of scale. Working within the unsupervised framework of denoising autoencoders, we observe that when the input is heavily corrupted during training, the network tends to learn coarse-grained features, whereas when the input is only slightly corrupted, the network tends to learn fine-grained features. This motivates the scheduled denoising autoencoder, which starts with a high level of noise that lowers as training progresses. We find that the resulting representation yields a significant boost on a later supervised task compared to the original input, or to a standard denoising autoencoder trained at a single noise level. After supervised fine-tuning our best model achieves the lowest ever reported error on the CIFAR-10 data set among permutation-invariant methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge