Konstantinos Kamnitsas
Analyzing Overfitting under Class Imbalance in Neural Networks for Image Segmentation
Feb 20, 2021
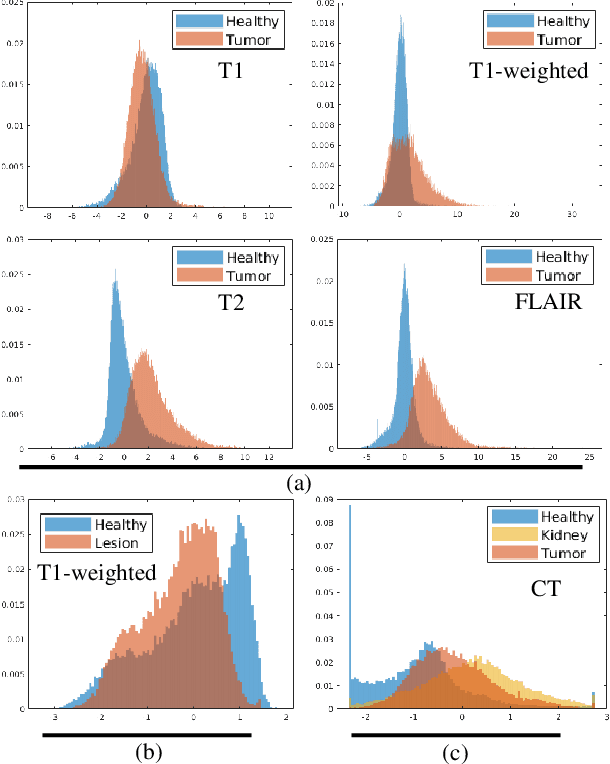
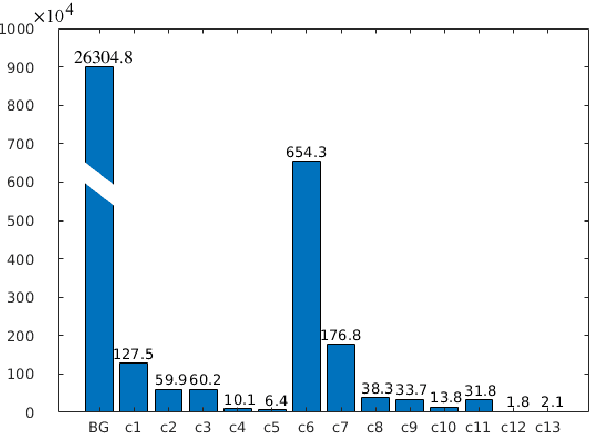
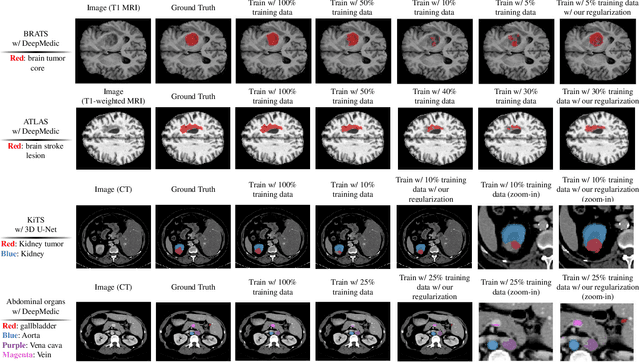
Abstract:Class imbalance poses a challenge for developing unbiased, accurate predictive models. In particular, in image segmentation neural networks may overfit to the foreground samples from small structures, which are often heavily under-represented in the training set, leading to poor generalization. In this study, we provide new insights on the problem of overfitting under class imbalance by inspecting the network behavior. We find empirically that when training with limited data and strong class imbalance, at test time the distribution of logit activations may shift across the decision boundary, while samples of the well-represented class seem unaffected. This bias leads to a systematic under-segmentation of small structures. This phenomenon is consistently observed for different databases, tasks and network architectures. To tackle this problem, we introduce new asymmetric variants of popular loss functions and regularization techniques including a large margin loss, focal loss, adversarial training, mixup and data augmentation, which are explicitly designed to counter logit shift of the under-represented classes. Extensive experiments are conducted on several challenging segmentation tasks. Our results demonstrate that the proposed modifications to the objective function can lead to significantly improved segmentation accuracy compared to baselines and alternative approaches.
Stochastic Segmentation Networks: Modelling Spatially Correlated Aleatoric Uncertainty
Jun 10, 2020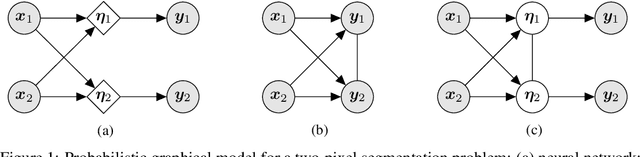
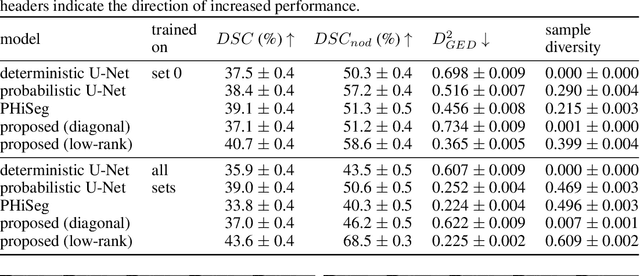

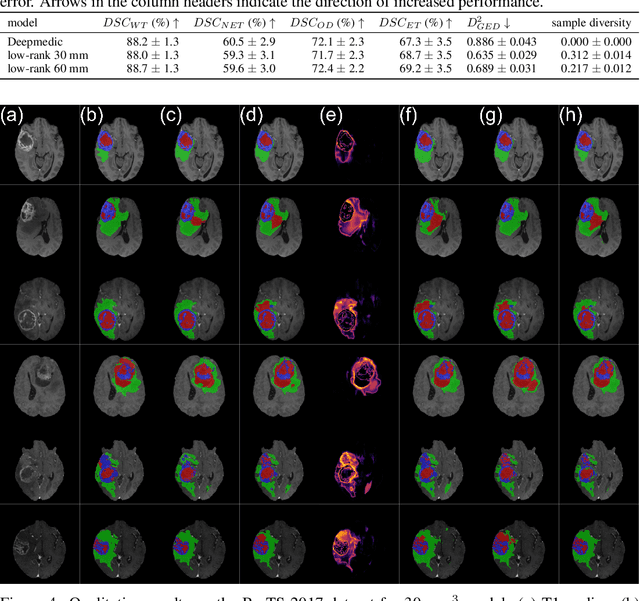
Abstract:In image segmentation, there is often more than one plausible solution for a given input. In medical imaging, for example, experts will often disagree about the exact location of object boundaries. Estimating this inherent uncertainty and predicting multiple plausible hypotheses is of great interest in many applications, yet this ability is lacking in most current deep learning methods. In this paper, we introduce stochastic segmentation networks (SSNs), an efficient probabilistic method for modelling aleatoric uncertainty with any image segmentation network architecture. In contrast to approaches that produce pixel-wise estimates, SSNs model joint distributions over entire label maps and thus can generate multiple spatially coherent hypotheses for a single image. By using a low-rank multivariate normal distribution over the logit space to model the probability of the label map given the image, we obtain a spatially consistent probability distribution that can be efficiently computed by a neural network without any changes to the underlying architecture. We tested our method on the segmentation of real-world medical data, including lung nodules in 2D CT and brain tumours in 3D multimodal MRI scans. SSNs outperform state-of-the-art for modelling correlated uncertainty in ambiguous images while being much simpler, more flexible, and more efficient.
Domain Generalization via Model-Agnostic Learning of Semantic Features
Oct 29, 2019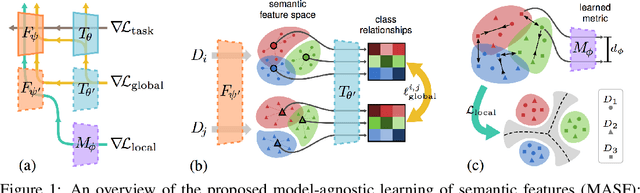
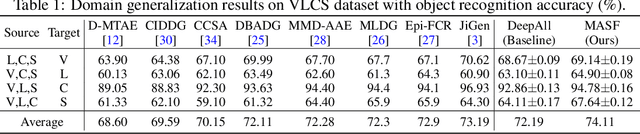
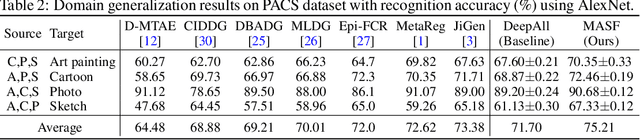
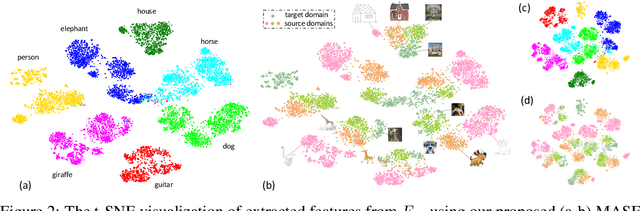
Abstract:Generalization capability to unseen domains is crucial for machine learning models when deploying to real-world conditions. We investigate the challenging problem of domain generalization, i.e., training a model on multi-domain source data such that it can directly generalize to target domains with unknown statistics. We adopt a model-agnostic learning paradigm with gradient-based meta-train and meta-test procedures to expose the optimization to domain shift. Further, we introduce two complementary losses which explicitly regularize the semantic structure of the feature space. Globally, we align a derived soft confusion matrix to preserve general knowledge about inter-class relationships. Locally, we promote domain-independent class-specific cohesion and separation of sample features with a metric-learning component. The effectiveness of our method is demonstrated with new state-of-the-art results on two common object recognition benchmarks. Our method also shows consistent improvement on a medical image segmentation task.
Data Efficient Unsupervised Domain Adaptation for Cross-Modality Image Segmentation
Aug 12, 2019
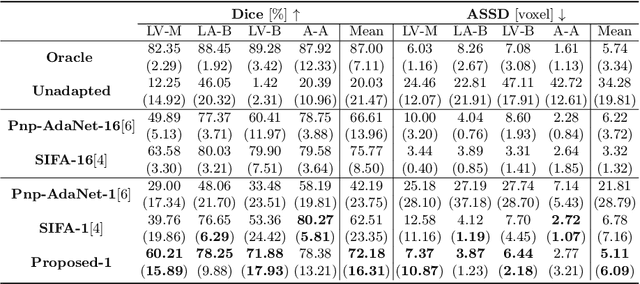
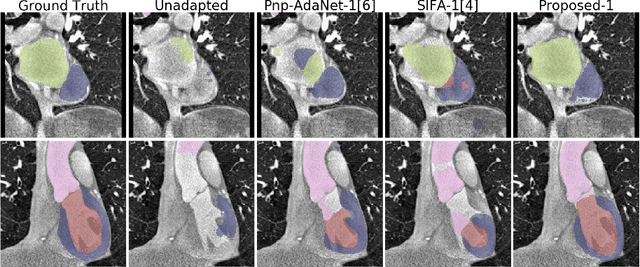
Abstract:Deep learning models trained on medical images from a source domain (e.g. imaging modality) often fail when deployed on images from a different target domain, despite imaging common anatomical structures. Deep unsupervised domain adaptation (UDA) aims to improve the performance of a deep neural network model on a target domain, using solely unlabelled target domain data and labelled source domain data. However, current state-of-the-art methods exhibit reduced performance when target data is scarce. In this work, we introduce a new data efficient UDA method for multi-domain medical image segmentation. The proposed method combines a novel VAE-based feature prior matching, which is data-efficient, and domain adversarial training to learn a shared domain-invariant latent space which is exploited during segmentation. Our method is evaluated on a public multi-modality cardiac image segmentation dataset by adapting from the labelled source domain (3D MRI) to the unlabelled target domain (3D CT). We show that by using only one single unlabelled 3D CT scan, the proposed architecture outperforms the state-of-the-art in the same setting. Finally, we perform ablation studies on prior matching and domain adversarial training to shed light on the theoretical grounding of the proposed method.
Overfitting of neural nets under class imbalance: Analysis and improvements for segmentation
Jul 25, 2019
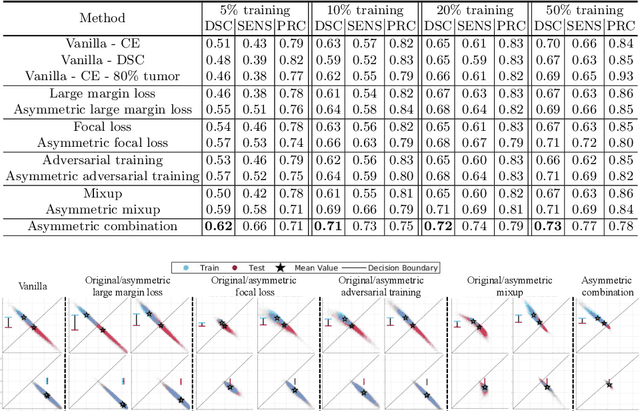
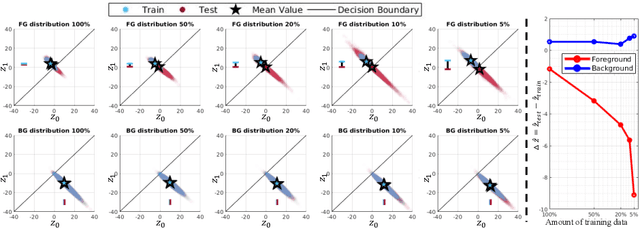
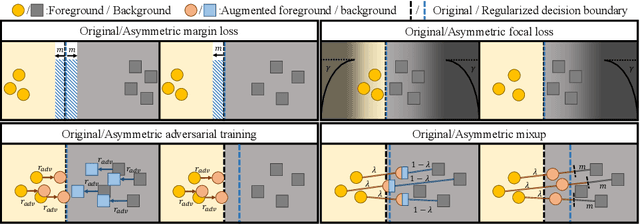
Abstract:Overfitting in deep learning has been the focus of a number of recent works, yet its exact impact on the behavior of neural networks is not well understood. This study analyzes overfitting by examining how the distribution of logits alters in relation to how much the model overfits. Specifically, we find that when training with few data samples, the distribution of logit activations when processing unseen test samples of an under-represented class tends to shift towards and even across the decision boundary, while the over-represented class seems unaffected. In image segmentation, foreground samples are often heavily under-represented. We observe that sensitivity of the model drops as a result of overfitting, while precision remains mostly stable. Based on our analysis, we derive asymmetric modifications of existing loss functions and regularizers including a large margin loss, focal loss, adversarial training and mixup, which specifically aim at reducing the shift observed when embedding unseen samples of the under-represented class. We study the case of binary segmentation of brain tumor core and show that our proposed simple modifications lead to significantly improved segmentation performance over the symmetric variants.
Multiple Landmark Detection using Multi-Agent Reinforcement Learning
Jul 22, 2019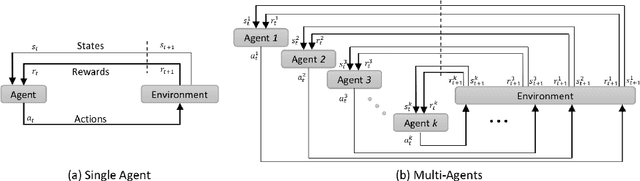
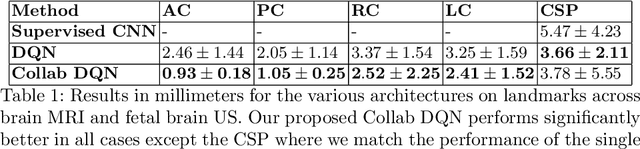
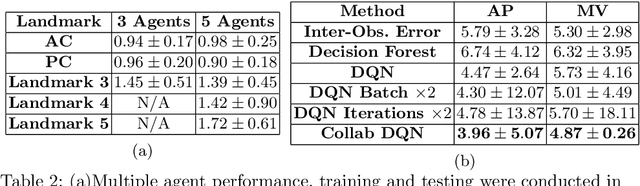

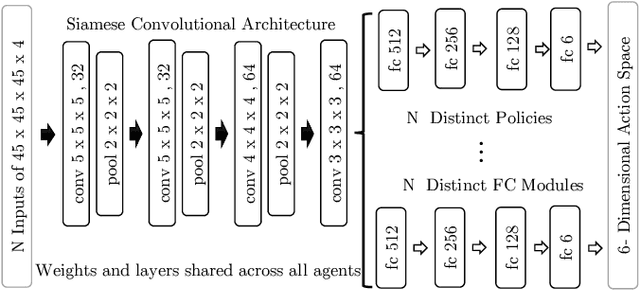

Abstract:The detection of anatomical landmarks is a vital step for medical image analysis and applications for diagnosis, interpretation and guidance. Manual annotation of landmarks is a tedious process that requires domain-specific expertise and introduces inter-observer variability. This paper proposes a new detection approach for multiple landmarks based on multi-agent reinforcement learning. Our hypothesis is that the position of all anatomical landmarks is interdependent and non-random within the human anatomy, thus finding one landmark can help to deduce the location of others. Using a Deep Q-Network (DQN) architecture we construct an environment and agent with implicit inter-communication such that we can accommodate K agents acting and learning simultaneously, while they attempt to detect K different landmarks. During training the agents collaborate by sharing their accumulated knowledge for a collective gain. We compare our approach with state-of-the-art architectures and achieve significantly better accuracy by reducing the detection error by 50%, while requiring fewer computational resources and time to train compared to the naive approach of training K agents separately.
Explainable Shape Analysis through Deep Hierarchical Generative Models: Application to Cardiac Remodeling
Jun 28, 2019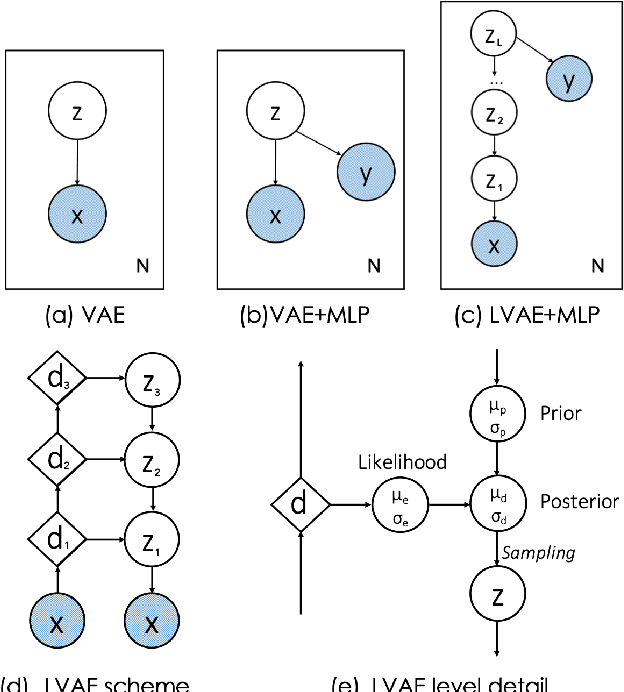
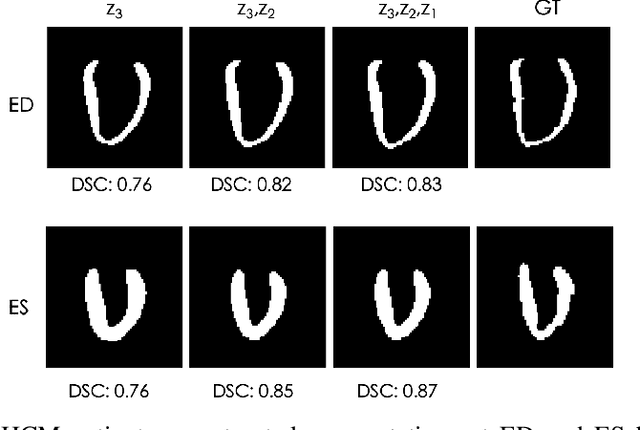
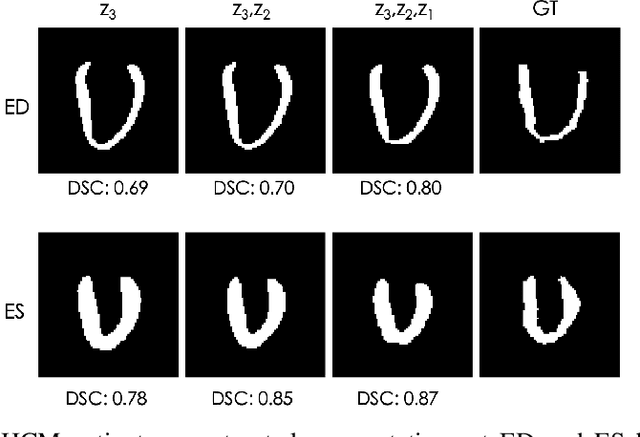
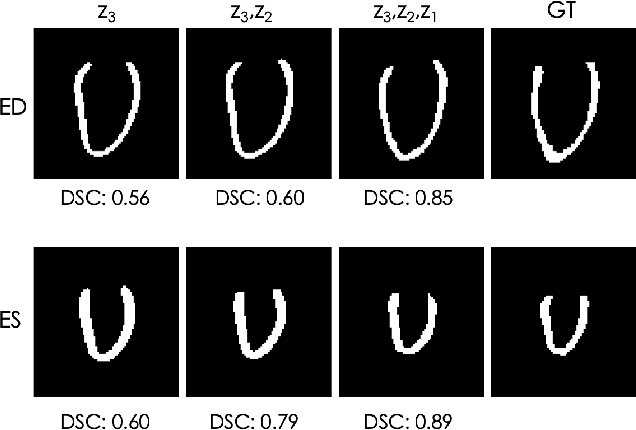


Abstract:Quantification of anatomical shape changes still relies on scalar global indexes which are largely insensitive to regional or asymmetric modifications. Accurate assessment of pathology-driven anatomical remodeling is a crucial step for the diagnosis and treatment of heart conditions. Deep learning approaches have recently achieved wide success in the analysis of medical images, but they lack interpretability in the feature extraction and decision processes. In this work, we propose a new interpretable deep learning model for shape analysis. In particular, we exploit deep generative networks to model a population of anatomical segmentations through a hierarchy of conditional latent variables. At the highest level of this hierarchy, a two-dimensional latent space is simultaneously optimised to discriminate distinct clinical conditions, enabling the direct visualisation of the classification space. Moreover, the anatomical variability encoded by this discriminative latent space can be visualised in the segmentation space thanks to the generative properties of the model, making the classification task transparent. This approach yielded high accuracy in the categorisation of healthy and remodelled hearts when tested on unseen segmentations from our own multi-centre dataset as well as in an external validation set. More importantly, it enabled the visualisation in three-dimensions of the most discriminative anatomical features between the two conditions. The proposed approach scales effectively to large populations, facilitating high-throughput analysis of normal anatomy and pathology in large-scale studies of volumetric imaging.
Towards continual learning in medical imaging
Nov 06, 2018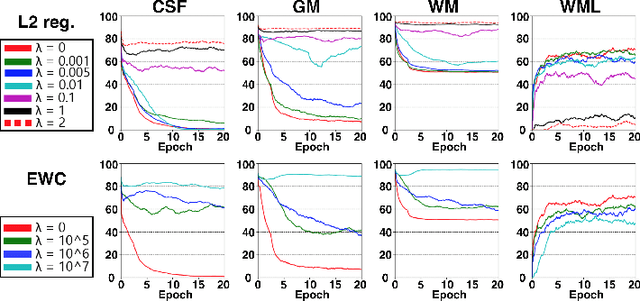
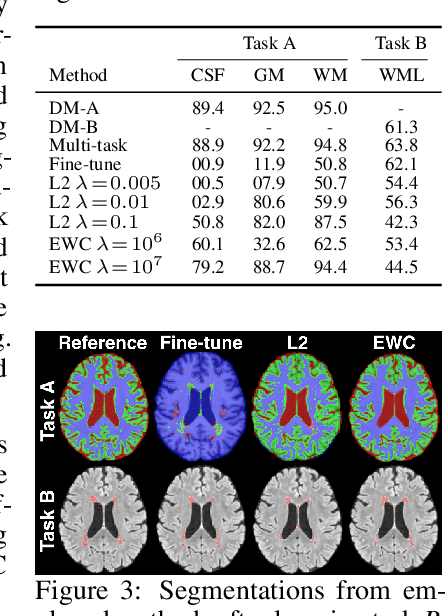
Abstract:This work investigates continual learning of two segmentation tasks in brain MRI with neural networks. To explore in this context the capabilities of current methods for countering catastrophic forgetting of the first task when a new one is learned, we investigate elastic weight consolidation, a recently proposed method based on Fisher information, originally evaluated on reinforcement learning of Atari games. We use it to sequentially learn segmentation of normal brain structures and then segmentation of white matter lesions. Our findings show this recent method reduces catastrophic forgetting, while large room for improvement exists in these challenging settings for continual learning.
Generative adversarial networks and adversarial methods in biomedical image analysis
Oct 24, 2018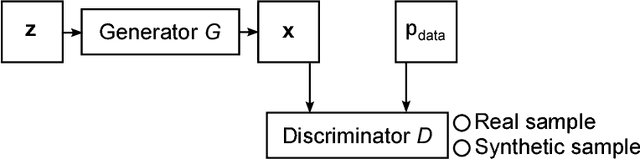
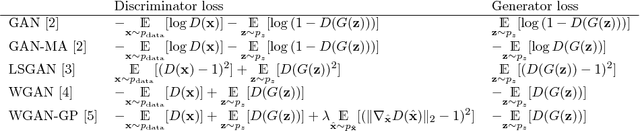
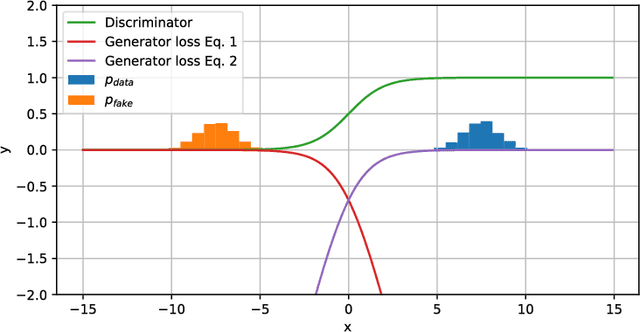
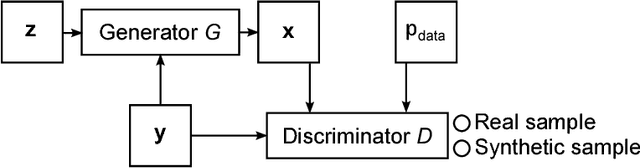
Abstract:Generative adversarial networks (GANs) and other adversarial methods are based on a game-theoretical perspective on joint optimization of two neural networks as players in a game. Adversarial techniques have been extensively used to synthesize and analyze biomedical images. We provide an introduction to GANs and adversarial methods, with an overview of biomedical image analysis tasks that have benefited from such methods. We conclude with a discussion of strengths and limitations of adversarial methods in biomedical image analysis, and propose potential future research directions.
Semi-Supervised Learning via Compact Latent Space Clustering
Jul 29, 2018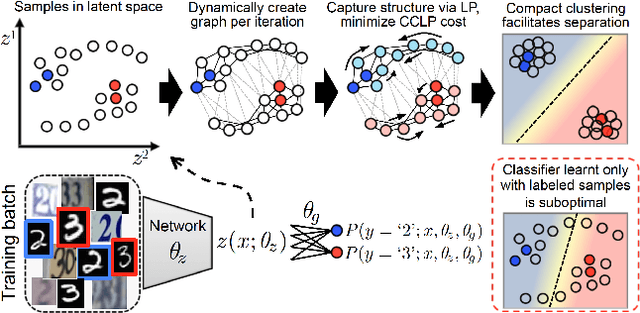
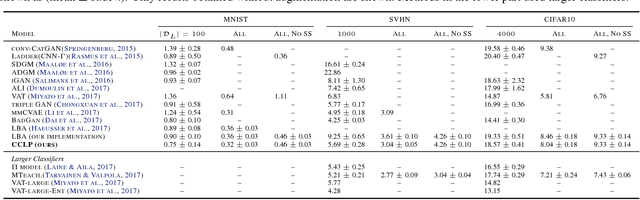
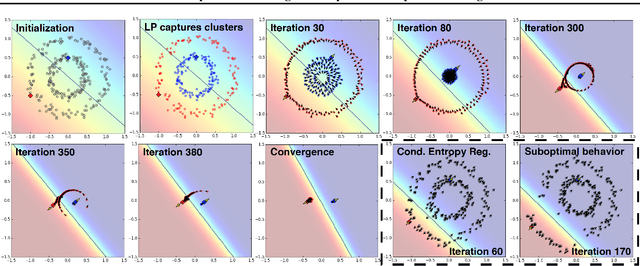
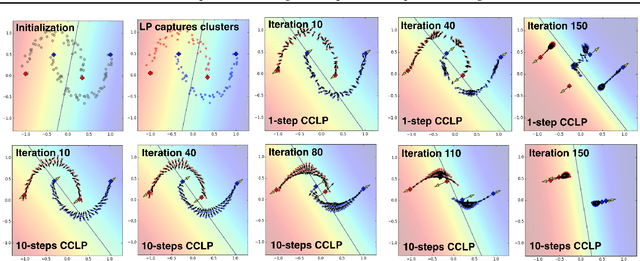
Abstract:We present a novel cost function for semi-supervised learning of neural networks that encourages compact clustering of the latent space to facilitate separation. The key idea is to dynamically create a graph over embeddings of labeled and unlabeled samples of a training batch to capture underlying structure in feature space, and use label propagation to estimate its high and low density regions. We then devise a cost function based on Markov chains on the graph that regularizes the latent space to form a single compact cluster per class, while avoiding to disturb existing clusters during optimization. We evaluate our approach on three benchmarks and compare to state-of-the art with promising results. Our approach combines the benefits of graph-based regularization with efficient, inductive inference, does not require modifications to a network architecture, and can thus be easily applied to existing networks to enable an effective use of unlabeled data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge