Jizhou Li
LamiGauss: Pitching Radiative Gaussian for Sparse-View X-ray Laminography Reconstruction
Sep 17, 2025
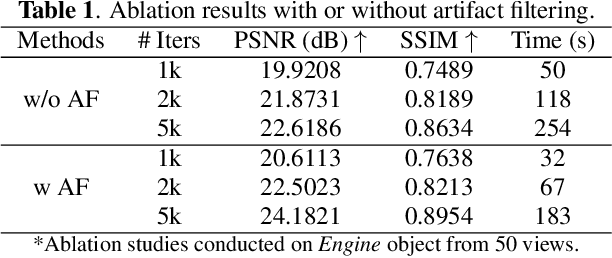


Abstract:X-ray Computed Laminography (CL) is essential for non-destructive inspection of plate-like structures in applications such as microchips and composite battery materials, where traditional computed tomography (CT) struggles due to geometric constraints. However, reconstructing high-quality volumes from laminographic projections remains challenging, particularly under highly sparse-view acquisition conditions. In this paper, we propose a reconstruction algorithm, namely LamiGauss, that combines Gaussian Splatting radiative rasterization with a dedicated detector-to-world transformation model incorporating the laminographic tilt angle. LamiGauss leverages an initialization strategy that explicitly filters out common laminographic artifacts from the preliminary reconstruction, preventing redundant Gaussians from being allocated to false structures and thereby concentrating model capacity on representing the genuine object. Our approach effectively optimizes directly from sparse projections, enabling accurate and efficient reconstruction with limited data. Extensive experiments on both synthetic and real datasets demonstrate the effectiveness and superiority of the proposed method over existing techniques. LamiGauss uses only 3$\%$ of full views to achieve superior performance over the iterative method optimized on a full dataset.
Learning neural representations for X-ray ptychography reconstruction with unknown probes
Sep 04, 2025



Abstract:X-ray ptychography provides exceptional nanoscale resolution and is widely applied in materials science, biology, and nanotechnology. However, its full potential is constrained by the critical challenge of accurately reconstructing images when the illuminating probe is unknown. Conventional iterative methods and deep learning approaches are often suboptimal, particularly under the low-signal conditions inherent to low-dose and high-speed experiments. These limitations compromise reconstruction fidelity and restrict the broader adoption of the technique. In this work, we introduce the Ptychographic Implicit Neural Representation (PtyINR), a self-supervised framework that simultaneously addresses the object and probe recovery problem. By parameterizing both as continuous neural representations, PtyINR performs end-to-end reconstruction directly from raw diffraction patterns without requiring any pre-characterization of the probe. Extensive evaluations demonstrate that PtyINR achieves superior reconstruction quality on both simulated and experimental data, with remarkable robustness under challenging low-signal conditions. Furthermore, PtyINR offers a generalizable, physics-informed framework for addressing probe-dependent inverse problems, making it applicable to a wide range of computational microscopy problems.
Implicit Regression in Subspace for High-Sensitivity CEST Imaging
Jul 09, 2024



Abstract:Chemical Exchange Saturation Transfer (CEST) MRI demonstrates its capability in significantly enhancing the detection of proteins and metabolites with low concentrations through exchangeable protons. The clinical application of CEST, however, is constrained by its low contrast and low signal-to-noise ratio (SNR) in the acquired data. Denoising, as one of the post-processing stages for CEST data, can effectively improve the accuracy of CEST quantification. In this work, by modeling spatial variant z-spectrums into low-dimensional subspace, we introduce Implicit Regression in Subspace (IRIS), which is an unsupervised denoising algorithm utilizing the excellent property of implicit neural representation for continuous mapping. Experiments conducted on both synthetic and in-vivo data demonstrate that our proposed method surpasses other CEST denoising methods regarding both qualitative and quantitative performance.
Generalized Robust Fundus Photography-based Vision Loss Estimation for High Myopia
Jul 04, 2024



Abstract:High myopia significantly increases the risk of irreversible vision loss. Traditional perimetry-based visual field (VF) assessment provides systematic quantification of visual loss but it is subjective and time-consuming. Consequently, machine learning models utilizing fundus photographs to estimate VF have emerged as promising alternatives. However, due to the high variability and the limited availability of VF data, existing VF estimation models fail to generalize well, particularly when facing out-of-distribution data across diverse centers and populations. To tackle this challenge, we propose a novel, parameter-efficient framework to enhance the generalized robustness of VF estimation on both in- and out-of-distribution data. Specifically, we design a Refinement-by-Denoising (RED) module for feature refinement and adaptation from pretrained vision models, aiming to learn high-entropy feature representations and to mitigate the domain gap effectively and efficiently. Through independent validation on two distinct real-world datasets from separate centers, our method significantly outperforms existing approaches in RMSE, MAE and correlation coefficient for both internal and external validation. Our proposed framework benefits both in- and out-of-distribution VF estimation, offering significant clinical implications and potential utility in real-world ophthalmic practices.
Boosting of Implicit Neural Representation-based Image Denoiser
Jan 03, 2024Abstract:Implicit Neural Representation (INR) has emerged as an effective method for unsupervised image denoising. However, INR models are typically overparameterized; consequently, these models are prone to overfitting during learning, resulting in suboptimal results, even noisy ones. To tackle this problem, we propose a general recipe for regularizing INR models in image denoising. In detail, we propose to iteratively substitute the supervision signal with the mean value derived from both the prediction and supervision signal during the learning process. We theoretically prove that such a simple iterative substitute can gradually enhance the signal-to-noise ratio of the supervision signal, thereby benefiting INR models during the learning process. Our experimental results demonstrate that INR models can be effectively regularized by the proposed approach, relieving overfitting and boosting image denoising performance.
Coordinate-based Neural Network for Fourier Phase Retrieval
Nov 25, 2023



Abstract:Fourier phase retrieval is essential for high-definition imaging of nanoscale structures across diverse fields, notably coherent diffraction imaging. This study presents the Single impliCit neurAl Network (SCAN), a tool built upon coordinate neural networks meticulously designed for enhanced phase retrieval performance. Bypassing the pitfalls of conventional iterative methods, which frequently face high computational loads and are prone to noise interference, SCAN adeptly connects object coordinates to their amplitude and phase within a unified network in an unsupervised manner. While many existing methods primarily use Fourier magnitude in their loss function, our approach incorporates both the predicted magnitude and phase, enhancing retrieval accuracy. Comprehensive tests validate SCAN's superiority over traditional and other deep learning models regarding accuracy and noise robustness. We also demonstrate that SCAN excels in the ptychography setting.
Robust retrieval of material chemical states in X-ray microspectroscopy
Aug 08, 2023Abstract:X-ray microspectroscopic techniques are essential for studying morphological and chemical changes in materials, providing high-resolution structural and spectroscopic information. However, its practical data analysis for reliably retrieving the chemical states remains a major obstacle to accelerating the fundamental understanding of materials in many research fields. In this work, we propose a novel data formulation model for X-ray microspectroscopy and develop a dedicated unmixing framework to solve this problem, which is robust to noise and spectral variability. Moreover, this framework is not limited to the analysis of two-state material chemistry, making it an effective alternative to conventional and widely-used methods. In addition, an alternative directional multiplier method with provable convergence is applied to obtain the solution efficiently. Our framework can accurately identify and characterize chemical states in complex and heterogeneous samples, even under challenging conditions such as low signal-to-noise ratios and overlapping spectral features. Extensive experimental results on simulated and real datasets demonstrate its effectiveness and reliability.
Learning to Deblur using Light Field Generated and Real Defocus Images
Apr 01, 2022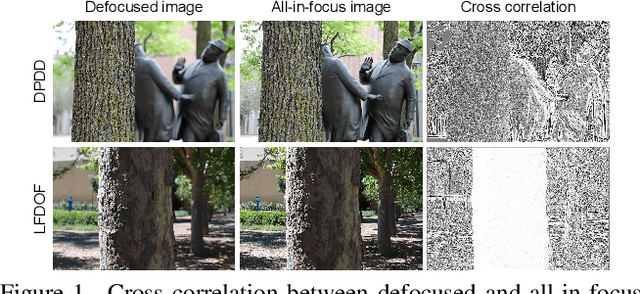

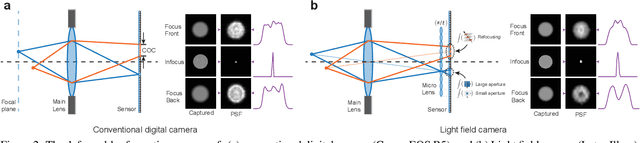
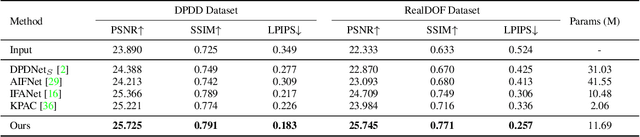
Abstract:Defocus deblurring is a challenging task due to the spatially varying nature of defocus blur. While deep learning approach shows great promise in solving image restoration problems, defocus deblurring demands accurate training data that consists of all-in-focus and defocus image pairs, which is difficult to collect. Naive two-shot capturing cannot achieve pixel-wise correspondence between the defocused and all-in-focus image pairs. Synthetic aperture of light fields is suggested to be a more reliable way to generate accurate image pairs. However, the defocus blur generated from light field data is different from that of the images captured with a traditional digital camera. In this paper, we propose a novel deep defocus deblurring network that leverages the strength and overcomes the shortcoming of light fields. We first train the network on a light field-generated dataset for its highly accurate image correspondence. Then, we fine-tune the network using feature loss on another dataset collected by the two-shot method to alleviate the differences between the defocus blur exists in the two domains. This strategy is proved to be highly effective and able to achieve the state-of-the-art performance both quantitatively and qualitatively on multiple test sets. Extensive ablation studies have been conducted to analyze the effect of each network module to the final performance.
Subspace modeling for fast and high-sensitivity X-ray chemical imaging
Jan 01, 2022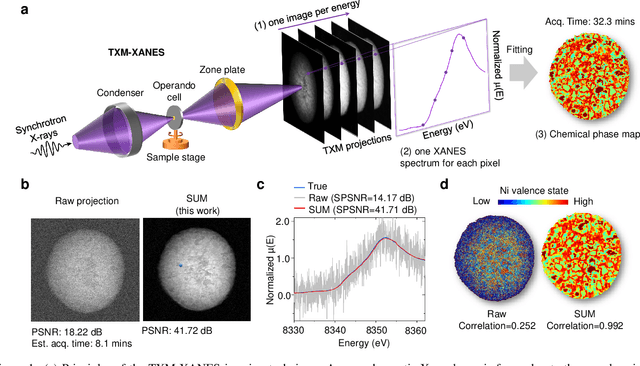
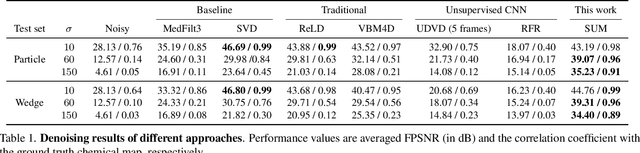
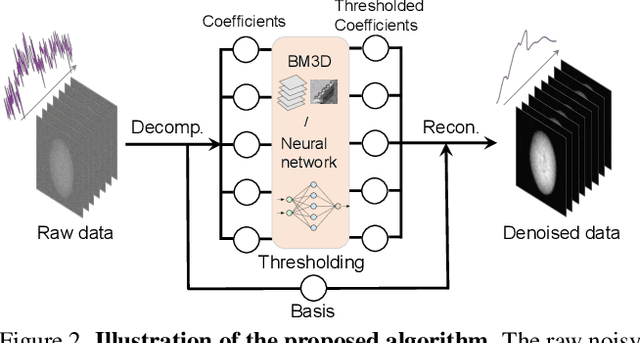
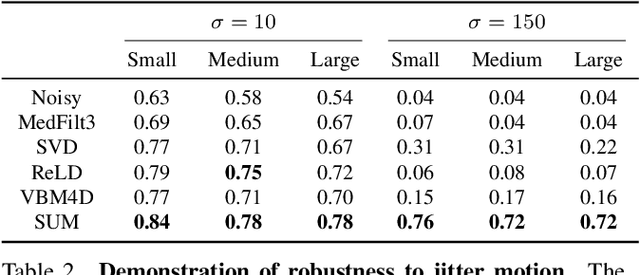
Abstract:Resolving morphological chemical phase transformations at the nanoscale is of vital importance to many scientific and industrial applications across various disciplines. The TXM-XANES imaging technique, by combining full field transmission X-ray microscopy (TXM) and X-ray absorption near edge structure (XANES), has been an emerging tool which operates by acquiring a series of microscopy images with multi-energy X-rays and fitting to obtain the chemical map. Its capability, however, is limited by the poor signal-to-noise ratios due to the system errors and low exposure illuminations for fast acquisition. In this work, by exploiting the intrinsic properties and subspace modeling of the TXM-XANES imaging data, we introduce a simple and robust denoising approach to improve the image quality, which enables fast and high-sensitivity chemical imaging. Extensive experiments on both synthetic and real datasets demonstrate the superior performance of the proposed method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge