Jinchuan Xiao
BioMiner: A Multi-modal System for Automated Mining of Protein-Ligand Bioactivity Data from Literature
Apr 23, 2026Abstract:Protein-ligand bioactivity data published in the literature are essential for drug discovery, yet manual curation struggles to keep pace with rapidly growing literature. Automated bioactivity extraction remains challenging because it requires not only interpreting biochemical semantics distributed across text, tables, and figures, but also reconstructing chemically exact ligand structures (e.g., Markush structures). To address this bottleneck, we introduce BioMiner, a multi-modal extraction framework that explicitly separates bioactivity semantic interpretation from ligand structure construction. Within BioMiner, bioactivity semantics are inferred through direct reasoning, while chemical structures are resolved via a chemical-structure-grounded visual semantic reasoning paradigm, in which multi-modal large language models operate on chemically grounded visual representations to infer inter-structure relationships, and exact molecular construction is delegated to domain chemistry tools. For rigorous evaluation and method development, we further establish BioVista, a comprehensive benchmark comprising 16,457 bioactivity entries curated from 500 publications. BioMiner validates its extraction ability and provides a quantitative baseline, achieving an F1 score of 0.32 for bioactivity triplets. BioMiner's practical utility is demonstrated via three applications: (1) extracting 82,262 data from 11,683 papers to build a pre-training database that improves downstream models performance by 3.9%; (2) enabling a human-in-the-loop workflow that doubles the number of high-quality NLRP3 bioactivity data, helping 38.6% improvement over 28 QSAR models and identification of 16 hit candidates with novel scaffolds; and (3) accelerating protein-ligand complex bioactivity annotation, achieving a 5.59-fold speed increase and 5.75% accuracy improvement over manual workflows in PoseBusters dataset.
MolMiner: You only look once for chemical structure recognition
May 23, 2022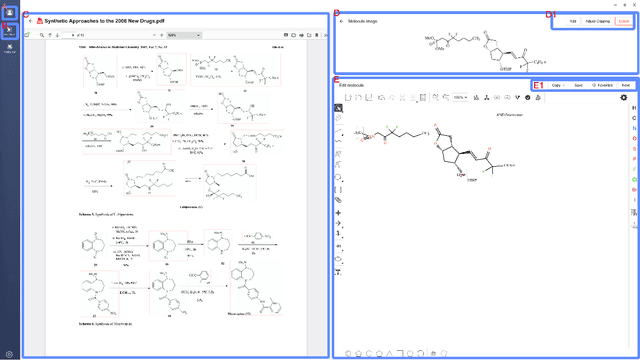
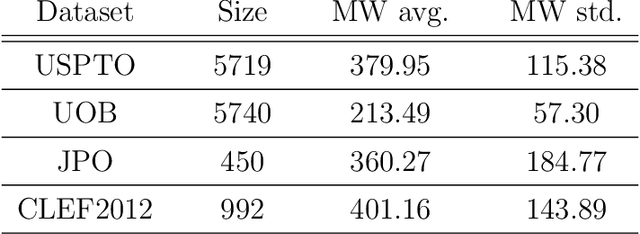
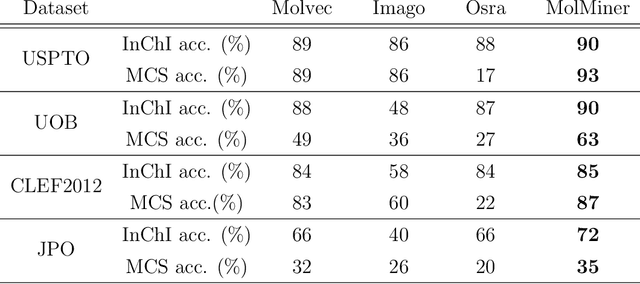
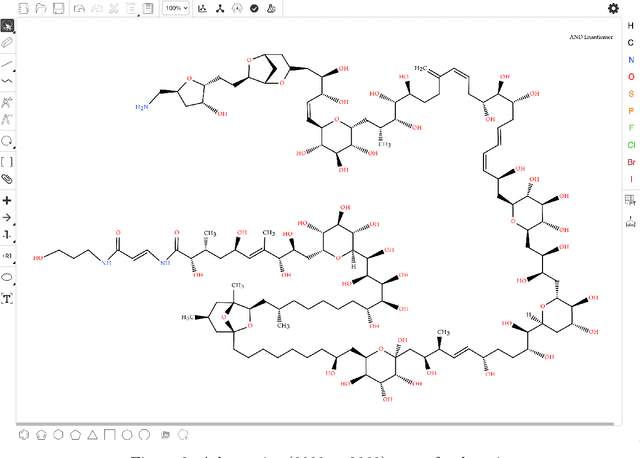
Abstract:Molecular structures are always depicted as 2D printed form in scientific documents like journal papers and patents. However, these 2D depictions are not machine-readable. Due to a backlog of decades and an increasing amount of these printed literature, there is a high demand for the translation of printed depictions into machine-readable formats, which is known as Optical Chemical Structure Recognition (OCSR). Most OCSR systems developed over the last three decades follow a rule-based approach where the key step of vectorization of the depiction is based on the interpretation of vectors and nodes as bonds and atoms. Here, we present a practical software MolMiner, which is primarily built up using deep neural networks originally developed for semantic segmentation and object detection to recognize atom and bond elements from documents. These recognized elements can be easily connected as a molecular graph with distance-based construction algorithm. We carefully evaluate our software on four benchmark datasets with the state-of-the-art performance. Various real application scenarios are also tested, yielding satisfactory outcomes. The free download links of Mac and Windows versions are available: Mac: https://molminer-cdn.iipharma.cn/pharma-mind/artifact/latest/mac/PharmaMind-mac-latest-setup.dmg and Windows: https://molminer-cdn.iipharma.cn/pharma-mind/artifact/latest/win/PharmaMind-win-latest-setup.exe
Improving Face Anti-Spoofing by 3D Virtual Synthesis
Jan 02, 2019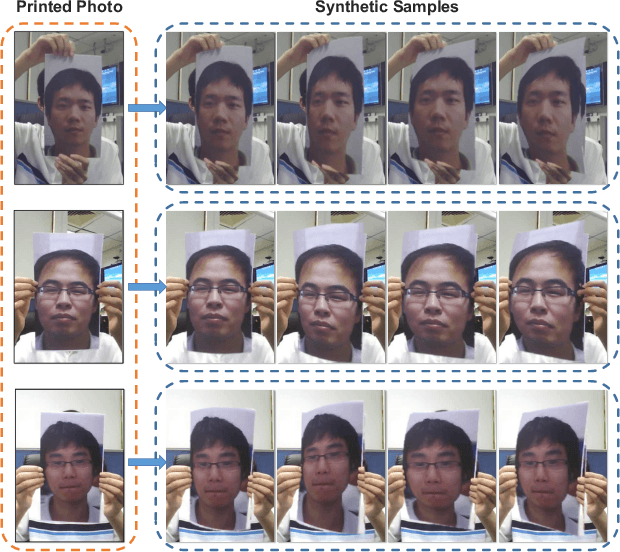
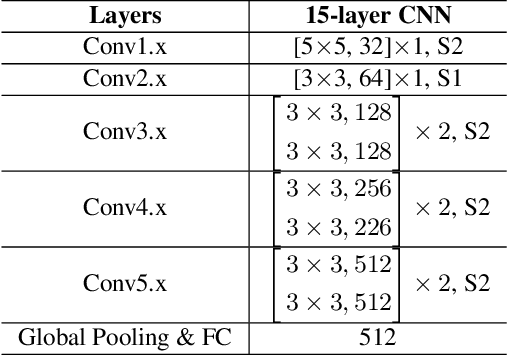

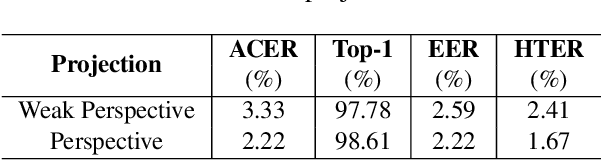
Abstract:Face anti-spoofing is crucial for the security of face recognition systems. Learning based methods especially deep learning based methods need large-scale training samples to reduce overfitting. However, acquiring spoof data is very expensive since the live faces should be re-printed and re-captured in many views. In this paper, we present a method to synthesize virtual spoof data in 3D space to alleviate this problem. Specifically, we consider a printed photo as a flat surface and mesh it into a 3D object, which is then randomly bent and rotated in 3D space. Afterward, the transformed 3D photo is rendered through perspective projection as a virtual sample. The synthetic virtual samples can significantly boost the anti-spoofing performance when combined with a proposed data balancing strategy. Our promising results open up new possibilities for advancing face anti-spoofing using cheap and large-scale synthetic data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge