Jin Tian
Iowa State University
Finding and Listing Front-door Adjustment Sets
Oct 14, 2022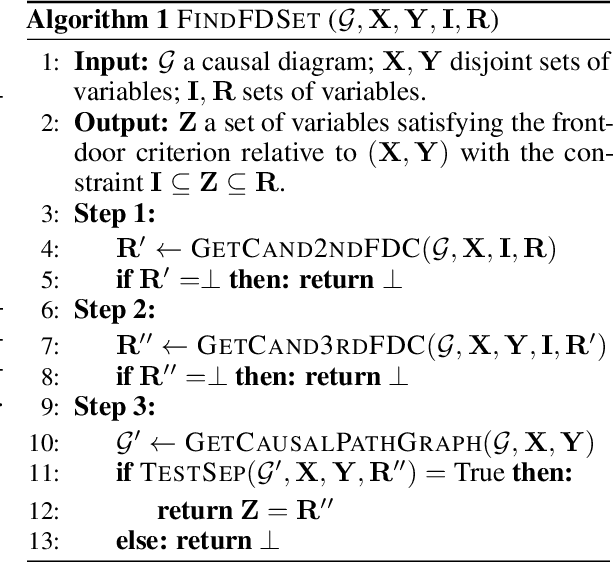

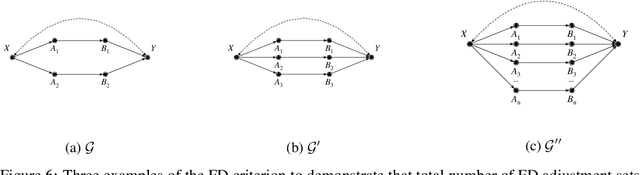
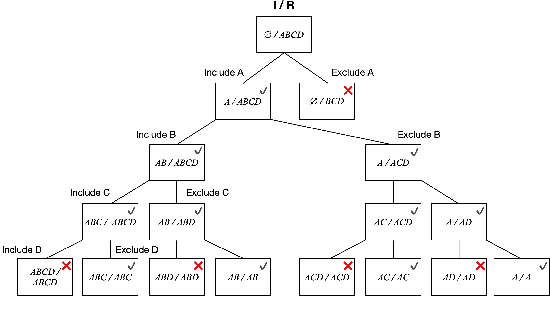
Abstract:Identifying the effects of new interventions from data is a significant challenge found across a wide range of the empirical sciences. A well-known strategy for identifying such effects is Pearl's front-door (FD) criterion (Pearl, 1995). The definition of the FD criterion is declarative, only allowing one to decide whether a specific set satisfies the criterion. In this paper, we present algorithms for finding and enumerating possible sets satisfying the FD criterion in a given causal diagram. These results are useful in facilitating the practical applications of the FD criterion for causal effects estimation and helping scientists to select estimands with desired properties, e.g., based on cost, feasibility of measurement, or statistical power.
Effect Identification in Cluster Causal Diagrams
Feb 22, 2022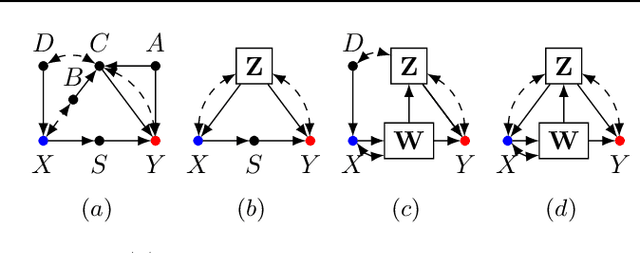
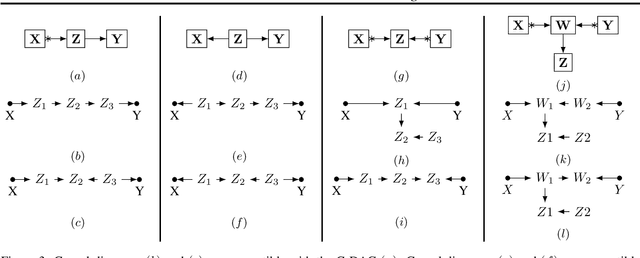
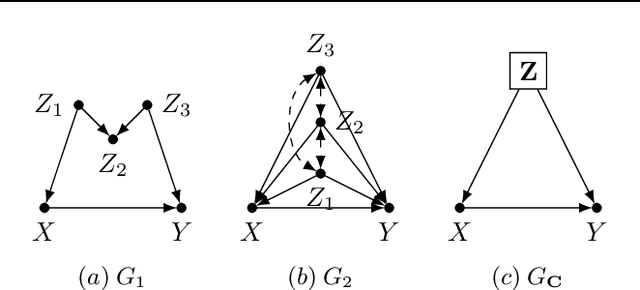

Abstract:One pervasive task found throughout the empirical sciences is to determine the effect of interventions from non-experimental data. It is well-understood that assumptions are necessary to perform causal inferences, which are commonly articulated through causal diagrams (Pearl, 2000). Despite the power of this approach, there are settings where the knowledge necessary to specify a causal diagram over all observed variables may not be available, particularly in complex, high-dimensional domains. In this paper, we introduce a new type of graphical model called cluster causal diagrams (for short, C-DAGs) that allows for the partial specification of relationships among variables based on limited prior knowledge, alleviating the stringent requirement of specifying a full causal diagram. A C-DAG specifies relationships between clusters of variables, while the relationships between the variables within a cluster are left unspecified. We develop the foundations and machinery for valid causal inferences over C-DAGs. In particular, we first define a new version of the d-separation criterion and prove its soundness and completeness. Secondly, we extend these new separation rules and prove the validity of the corresponding do-calculus. Lastly, we show that a standard identification algorithm is sound and complete to systematically compute causal effects from observational data given a C-DAG.
Partial Counterfactual Identification from Observational and Experimental Data
Oct 12, 2021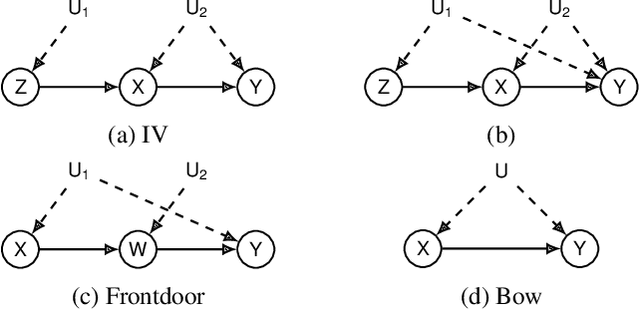
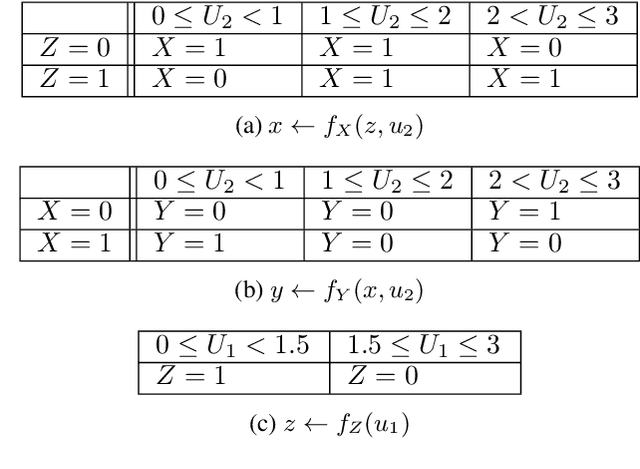

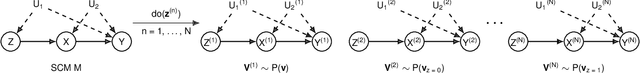
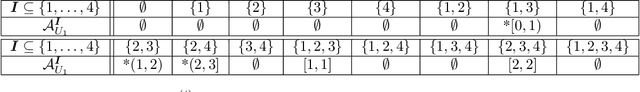
Abstract:This paper investigates the problem of bounding counterfactual queries from an arbitrary collection of observational and experimental distributions and qualitative knowledge about the underlying data-generating model represented in the form of a causal diagram. We show that all counterfactual distributions in an arbitrary structural causal model (SCM) could be generated by a canonical family of SCMs with the same causal diagram where unobserved (exogenous) variables are discrete with a finite domain. Utilizing the canonical SCMs, we translate the problem of bounding counterfactuals into that of polynomial programming whose solution provides optimal bounds for the counterfactual query. Solving such polynomial programs is in general computationally expensive. We therefore develop effective Monte Carlo algorithms to approximate the optimal bounds from an arbitrary combination of observational and experimental data. Our algorithms are validated extensively on synthetic and real-world datasets.
Beyond Discriminant Patterns: On the Robustness of Decision Rule Ensembles
Sep 21, 2021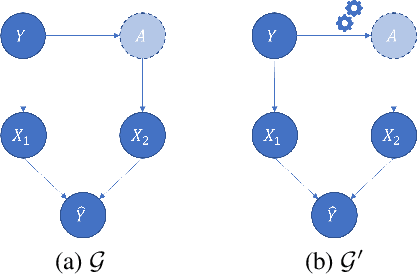
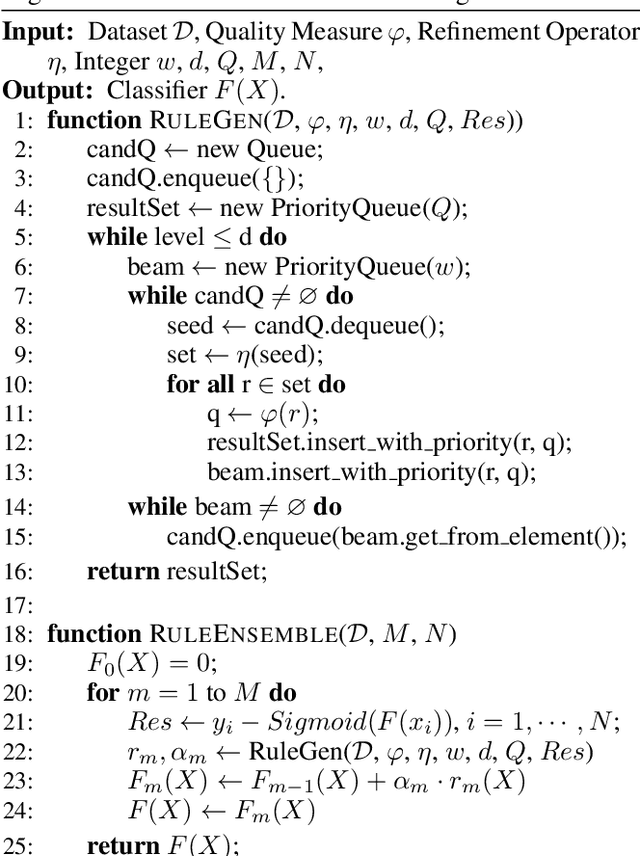
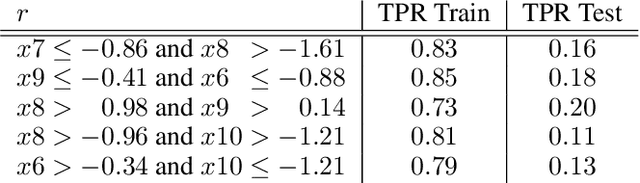

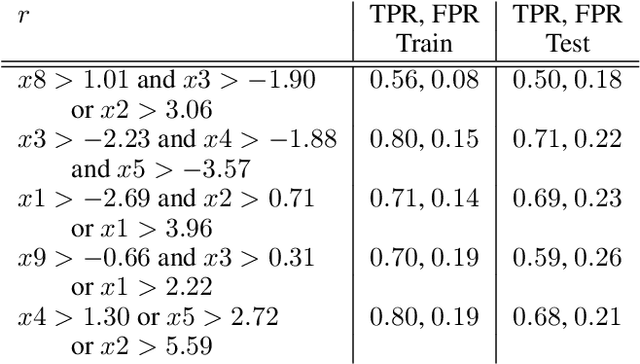
Abstract:Local decision rules are commonly understood to be more explainable, due to the local nature of the patterns involved. With numerical optimization methods such as gradient boosting, ensembles of local decision rules can gain good predictive performance on data involving global structure. Meanwhile, machine learning models are being increasingly used to solve problems in high-stake domains including healthcare and finance. Here, there is an emerging consensus regarding the need for practitioners to understand whether and how those models could perform robustly in the deployment environments, in the presence of distributional shifts. Past research on local decision rules has focused mainly on maximizing discriminant patterns, without due consideration of robustness against distributional shifts. In order to fill this gap, we propose a new method to learn and ensemble local decision rules, that are robust both in the training and deployment environments. Specifically, we propose to leverage causal knowledge by regarding the distributional shifts in subpopulations and deployment environments as the results of interventions on the underlying system. We propose two regularization terms based on causal knowledge to search for optimal and stable rules. Experiments on both synthetic and benchmark datasets show that our method is effective and robust against distributional shifts in multiple environments.
Data Poisoning Attacks and Defenses to Crowdsourcing Systems
Feb 24, 2021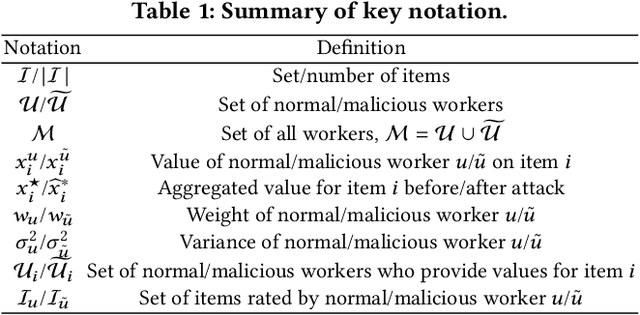
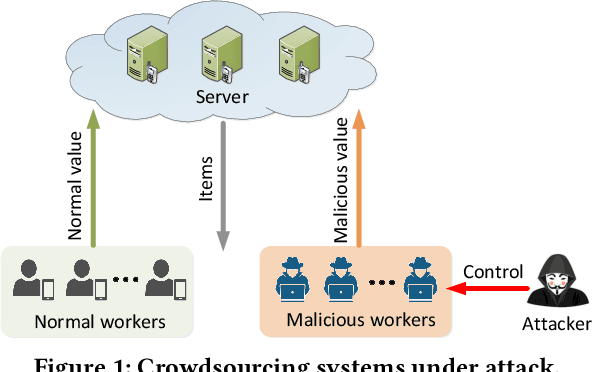
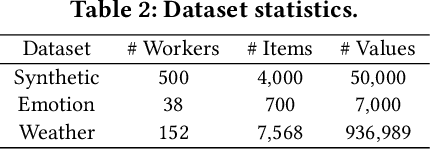
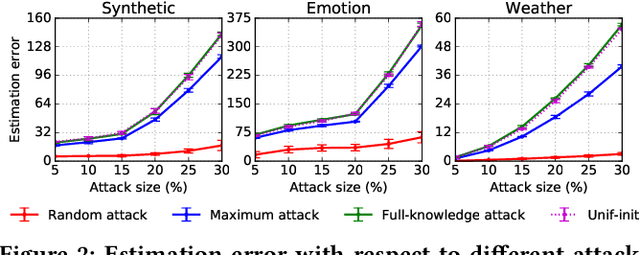
Abstract:A key challenge of big data analytics is how to collect a large volume of (labeled) data. Crowdsourcing aims to address this challenge via aggregating and estimating high-quality data (e.g., sentiment label for text) from pervasive clients/users. Existing studies on crowdsourcing focus on designing new methods to improve the aggregated data quality from unreliable/noisy clients. However, the security aspects of such crowdsourcing systems remain under-explored to date. We aim to bridge this gap in this work. Specifically, we show that crowdsourcing is vulnerable to data poisoning attacks, in which malicious clients provide carefully crafted data to corrupt the aggregated data. We formulate our proposed data poisoning attacks as an optimization problem that maximizes the error of the aggregated data. Our evaluation results on one synthetic and two real-world benchmark datasets demonstrate that the proposed attacks can substantially increase the estimation errors of the aggregated data. We also propose two defenses to reduce the impact of malicious clients. Our empirical results show that the proposed defenses can substantially reduce the estimation errors of the data poisoning attacks.
Supervised Whole DAG Causal Discovery
Jun 08, 2020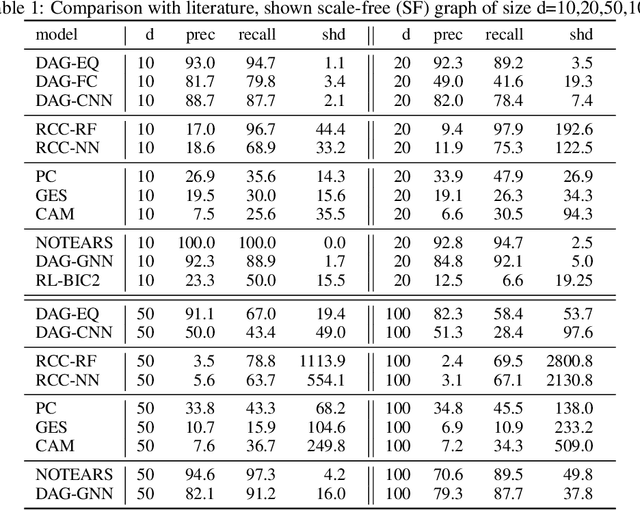
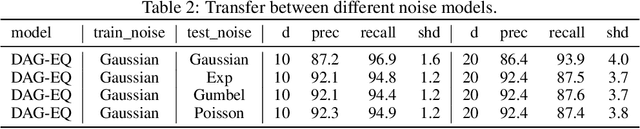
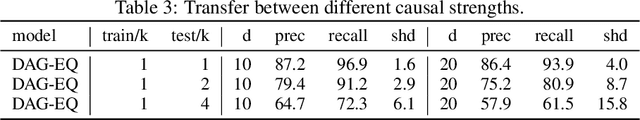
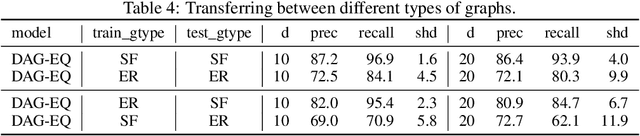
Abstract:We propose to address the task of causal structure learning from data in a supervised manner. Existing work on learning causal directions by supervised learning is restricted to learning pairwise relation, and not well suited for whole DAG discovery. We propose a novel approach of modeling the whole DAG structure discovery as a supervised learning. To fit the problem in hand, we propose to use permutation equivariant models that align well with the problem domain. We evaluate the proposed approach extensively on synthetic graphs of size 10,20,50,100 and real data, and show promising results compared with a variety of previous approaches.
Adjustment Criteria for Recovering Causal Effects from Missing Data
Aug 13, 2019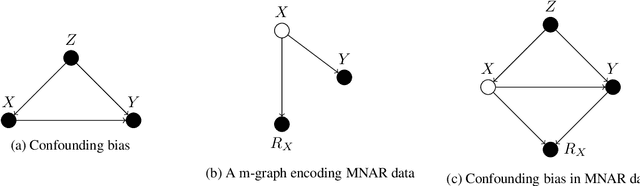
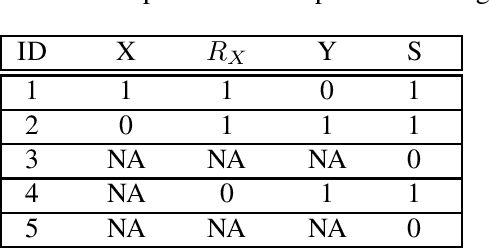

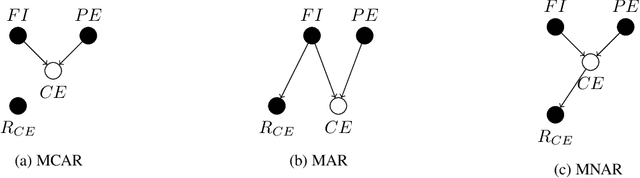
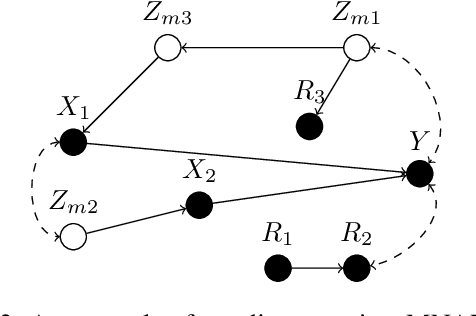
Abstract:Confounding bias, missing data, and selection bias are three common obstacles to valid causal inference in the data sciences. Covariate adjustment is the most pervasive technique for recovering casual effects from confounding bias. In this paper, we introduce a covariate adjustment formulation for controlling confounding bias in the presence of missing-not-at-random data and develop a necessary and sufficient condition for recovering causal effects using the adjustment. We also introduce an adjustment formulation for controlling both confounding and selection biases in the presence of missing data and develop a necessary and sufficient condition for valid adjustment. Furthermore, we present an algorithm that lists all valid adjustment sets and an algorithm that finds a valid adjustment set containing the minimum number of variables, which are useful for researchers interested in selecting adjustment sets with desired properties.
Purifying Adversarial Perturbation with Adversarially Trained Auto-encoders
May 26, 2019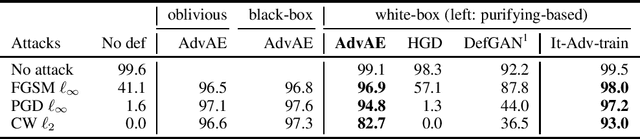
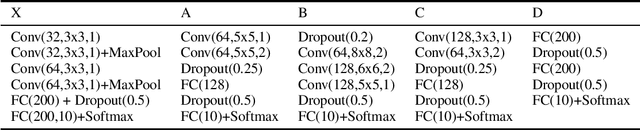

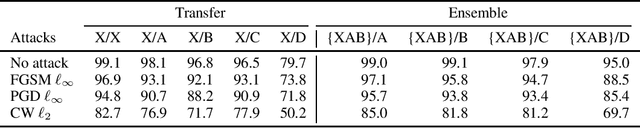
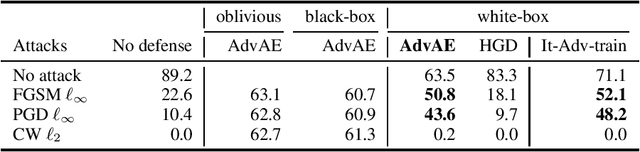
Abstract:Machine learning models are vulnerable to adversarial examples. Iterative adversarial training has shown promising results against strong white-box attacks. However, adversarial training is very expensive, and every time a model needs to be protected, such expensive training scheme needs to be performed. In this paper, we propose to apply iterative adversarial training scheme to an external auto-encoder, which once trained can be used to protect other models directly. We empirically show that our model outperforms other purifying-based methods against white-box attacks, and transfers well to directly protect other base models with different architectures.
Recoverability of Joint Distribution from Missing Data
Nov 15, 2016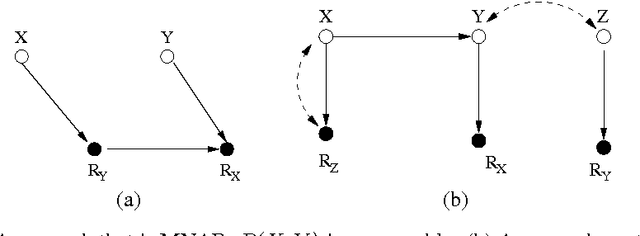
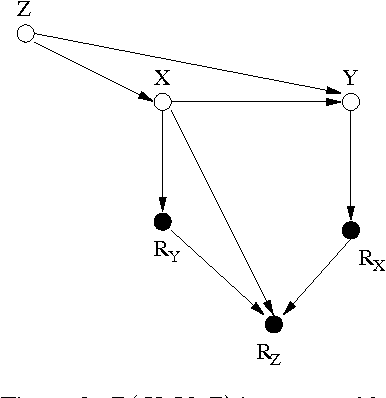
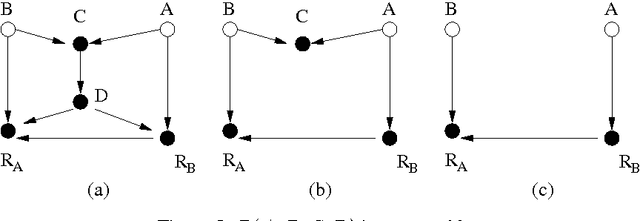
Abstract:A probabilistic query may not be estimable from observed data corrupted by missing values if the data are not missing at random (MAR). It is therefore of theoretical interest and practical importance to determine in principle whether a probabilistic query is estimable from missing data or not when the data are not MAR. We present an algorithm that systematically determines whether the joint probability is estimable from observed data with missing values, assuming that the data-generation model is represented as a Bayesian network containing unobserved latent variables that not only encodes the dependencies among the variables but also explicitly portrays the mechanisms responsible for the missingness process. The result significantly advances the existing work.
A Parallel Algorithm for Exact Bayesian Structure Discovery in Bayesian Networks
Aug 13, 2016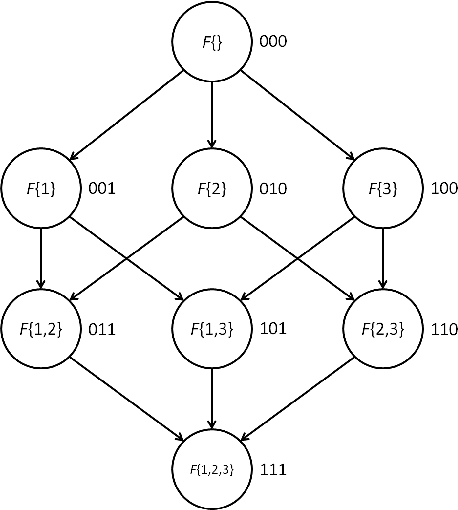
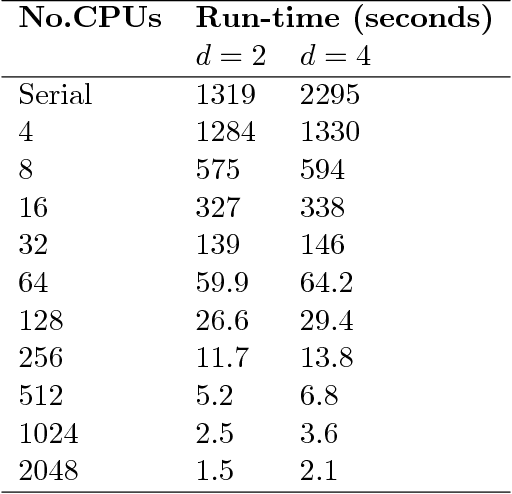
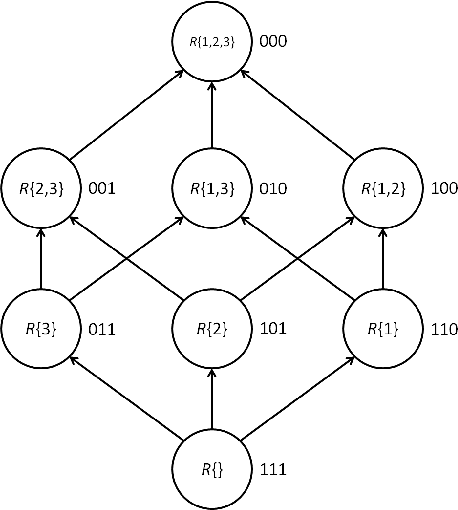
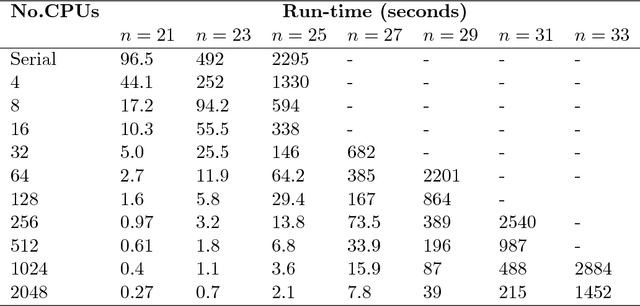
Abstract:Exact Bayesian structure discovery in Bayesian networks requires exponential time and space. Using dynamic programming (DP), the fastest known sequential algorithm computes the exact posterior probabilities of structural features in $O(2(d+1)n2^n)$ time and space, if the number of nodes (variables) in the Bayesian network is $n$ and the in-degree (the number of parents) per node is bounded by a constant $d$. Here we present a parallel algorithm capable of computing the exact posterior probabilities for all $n(n-1)$ edges with optimal parallel space efficiency and nearly optimal parallel time efficiency. That is, if $p=2^k$ processors are used, the run-time reduces to $O(5(d+1)n2^{n-k}+k(n-k)^d)$ and the space usage becomes $O(n2^{n-k})$ per processor. Our algorithm is based the observation that the subproblems in the sequential DP algorithm constitute a $n$-$D$ hypercube. We take a delicate way to coordinate the computation of correlated DP procedures such that large amount of data exchange is suppressed. Further, we develop parallel techniques for two variants of the well-known \emph{zeta transform}, which have applications outside the context of Bayesian networks. We demonstrate the capability of our algorithm on datasets with up to 33 variables and its scalability on up to 2048 processors. We apply our algorithm to a biological data set for discovering the yeast pheromone response pathways.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge