Jiawen Yao
A deep local attention network for pre-operative lymph node metastasis prediction in pancreatic cancer via multiphase CT imaging
Jan 04, 2023



Abstract:Lymph node (LN) metastasis status is one of the most critical prognostic and cancer staging factors for patients with resectable pancreatic ductal adenocarcinoma (PDAC), or in general, for any types of solid malignant tumors. Preoperative prediction of LN metastasis from non-invasive CT imaging is highly desired, as it might be straightforwardly used to guide the following neoadjuvant treatment decision and surgical planning. Most studies only capture the tumor characteristics in CT imaging to implicitly infer LN metastasis and very few work exploit direct LN's CT imaging information. To the best of our knowledge, this is the first work to propose a fully-automated LN segmentation and identification network to directly facilitate the LN metastasis status prediction task. Nevertheless LN segmentation/detection is very challenging since LN can be easily confused with other hard negative anatomic structures (e.g., vessels) from radiological images. We explore the anatomical spatial context priors of pancreatic LN locations by generating a guiding attention map from related organs and vessels to assist segmentation and infer LN status. As such, LN segmentation is impelled to focus on regions that are anatomically adjacent or plausible with respect to the specific organs and vessels. The metastasized LN identification network is trained to classify the segmented LN instances into positives or negatives by reusing the segmentation network as a pre-trained backbone and padding a new classification head. More importantly, we develop a LN metastasis status prediction network that combines the patient-wise aggregation results of LN segmentation/identification and deep imaging features extracted from the tumor region. Extensive quantitative nested five-fold cross-validation is conducted on a discovery dataset of 749 patients with PDAC.
Hierarchical Transformer for Survival Prediction Using Multimodality Whole Slide Images and Genomics
Nov 29, 2022Abstract:Learning good representation of giga-pixel level whole slide pathology images (WSI) for downstream tasks is critical. Previous studies employ multiple instance learning (MIL) to represent WSIs as bags of sampled patches because, for most occasions, only slide-level labels are available, and only a tiny region of the WSI is disease-positive area. However, WSI representation learning still remains an open problem due to: (1) patch sampling on a higher resolution may be incapable of depicting microenvironment information such as the relative position between the tumor cells and surrounding tissues, while patches at lower resolution lose the fine-grained detail; (2) extracting patches from giant WSI results in large bag size, which tremendously increases the computational cost. To solve the problems, this paper proposes a hierarchical-based multimodal transformer framework that learns a hierarchical mapping between pathology images and corresponding genes. Precisely, we randomly extract instant-level patch features from WSIs with different magnification. Then a co-attention mapping between imaging and genomics is learned to uncover the pairwise interaction and reduce the space complexity of imaging features. Such early fusion makes it computationally feasible to use MIL Transformer for the survival prediction task. Our architecture requires fewer GPU resources compared with benchmark methods while maintaining better WSI representation ability. We evaluate our approach on five cancer types from the Cancer Genome Atlas database and achieved an average c-index of $0.673$, outperforming the state-of-the-art multimodality methods.
3D Graph Anatomy Geometry-Integrated Network for Pancreatic Mass Segmentation, Diagnosis, and Quantitative Patient Management
Dec 08, 2020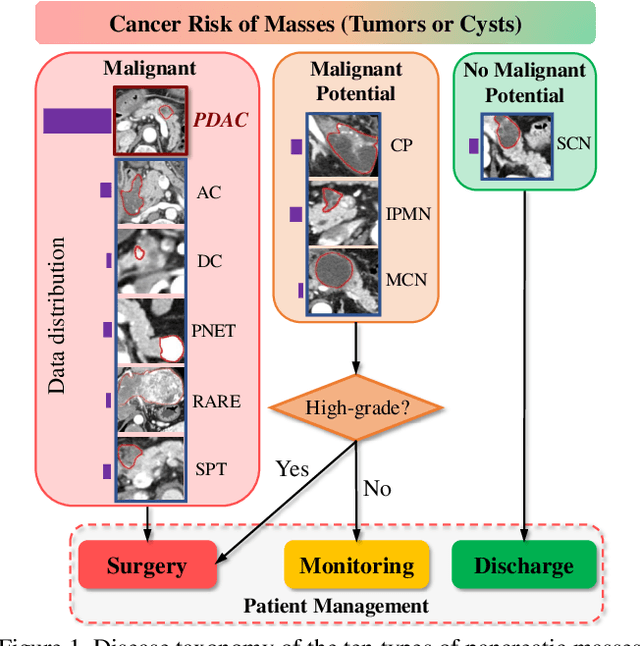
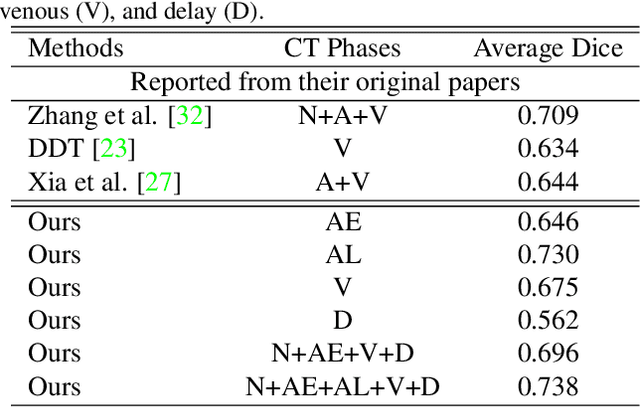
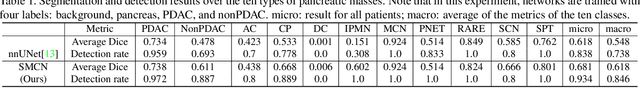
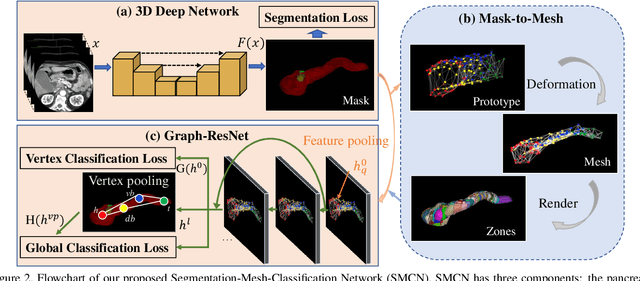

Abstract:The pancreatic disease taxonomy includes ten types of masses (tumors or cysts)[20,8]. Previous work focuses on developing segmentation or classification methods only for certain mass types. Differential diagnosis of all mass types is clinically highly desirable [20] but has not been investigated using an automated image understanding approach. We exploit the feasibility to distinguish pancreatic ductal adenocarcinoma (PDAC) from the nine other nonPDAC masses using multi-phase CT imaging. Both image appearance and the 3D organ-mass geometry relationship are critical. We propose a holistic segmentation-mesh-classification network (SMCN) to provide patient-level diagnosis, by fully utilizing the geometry and location information, which is accomplished by combining the anatomical structure and the semantic detection-by-segmentation network. SMCN learns the pancreas and mass segmentation task and builds an anatomical correspondence-aware organ mesh model by progressively deforming a pancreas prototype on the raw segmentation mask (i.e., mask-to-mesh). A new graph-based residual convolutional network (Graph-ResNet), whose nodes fuse the information of the mesh model and feature vectors extracted from the segmentation network, is developed to produce the patient-level differential classification results. Extensive experiments on 661 patients' CT scans (five phases per patient) show that SMCN can improve the mass segmentation and detection accuracy compared to the strong baseline method nnUNet (e.g., for nonPDAC, Dice: 0.611 vs. 0.478; detection rate: 89% vs. 70%), achieve similar sensitivity and specificity in differentiating PDAC and nonPDAC as expert radiologists (i.e., 94% and 90%), and obtain results comparable to a multimodality test [20] that combines clinical, imaging, and molecular testing for clinical management of patients.
Whole Slide Images based Cancer Survival Prediction using Attention Guided Deep Multiple Instance Learning Networks
Sep 23, 2020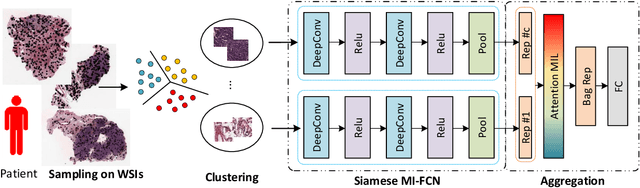
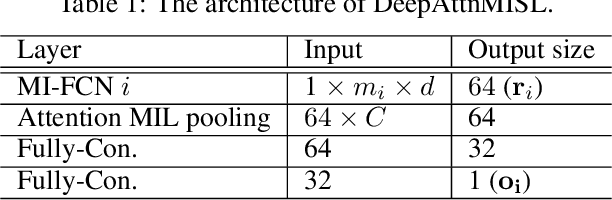

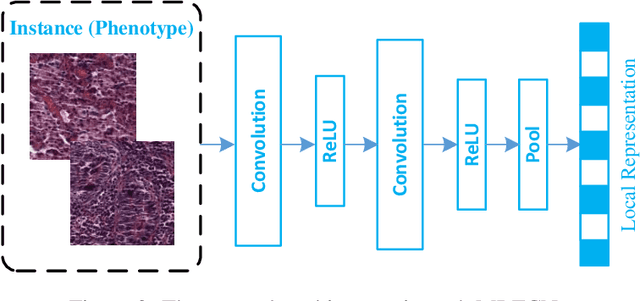
Abstract:Traditional image-based survival prediction models rely on discriminative patch labeling which make those methods not scalable to extend to large datasets. Recent studies have shown Multiple Instance Learning (MIL) framework is useful for histopathological images when no annotations are available in classification task. Different to the current image-based survival models that limit to key patches or clusters derived from Whole Slide Images (WSIs), we propose Deep Attention Multiple Instance Survival Learning (DeepAttnMISL) by introducing both siamese MI-FCN and attention-based MIL pooling to efficiently learn imaging features from the WSI and then aggregate WSI-level information to patient-level. Attention-based aggregation is more flexible and adaptive than aggregation techniques in recent survival models. We evaluated our methods on two large cancer whole slide images datasets and our results suggest that the proposed approach is more effective and suitable for large datasets and has better interpretability in locating important patterns and features that contribute to accurate cancer survival predictions. The proposed framework can also be used to assess individual patient's risk and thus assisting in delivering personalized medicine. Codes are available at https://github.com/uta-smile/DeepAttnMISL_MEDIA.
* 22 pages, 13 figures, published in Medical Image Analysis 65, 101789
DeepPrognosis: Preoperative Prediction of Pancreatic Cancer Survival and Surgical Margin via Contrast-Enhanced CT Imaging
Aug 26, 2020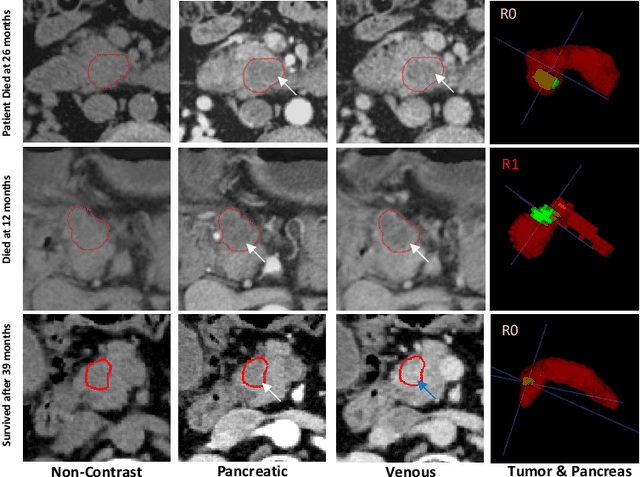
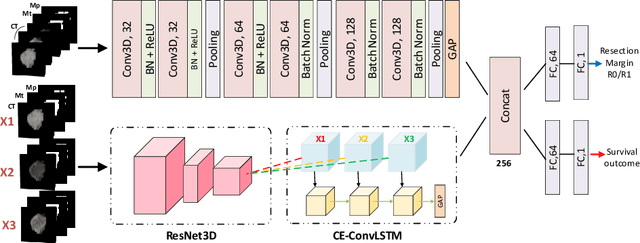
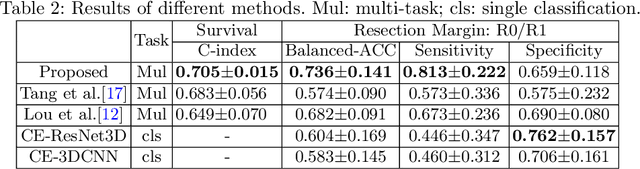
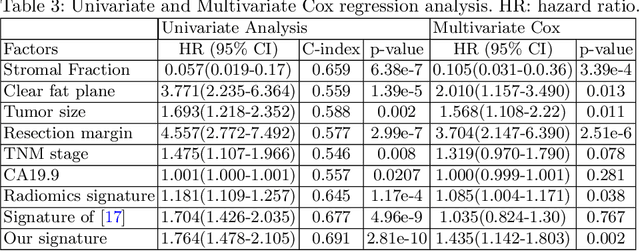
Abstract:Pancreatic ductal adenocarcinoma (PDAC) is one of the most lethal cancers and carries a dismal prognosis. Surgery remains the best chance of a potential cure for patients who are eligible for initial resection of PDAC. However, outcomes vary significantly even among the resected patients of the same stage and received similar treatments. Accurate preoperative prognosis of resectable PDACs for personalized treatment is thus highly desired. Nevertheless, there are no automated methods yet to fully exploit the contrast-enhanced computed tomography (CE-CT) imaging for PDAC. Tumor attenuation changes across different CT phases can reflect the tumor internal stromal fractions and vascularization of individual tumors that may impact the clinical outcomes. In this work, we propose a novel deep neural network for the survival prediction of resectable PDAC patients, named as 3D Contrast-Enhanced Convolutional Long Short-Term Memory network(CE-ConvLSTM), which can derive the tumor attenuation signatures or patterns from CE-CT imaging studies. We present a multi-task CNN to accomplish both tasks of outcome and margin prediction where the network benefits from learning the tumor resection margin related features to improve survival prediction. The proposed framework can improve the prediction performances compared with existing state-of-the-art survival analysis approaches. The tumor signature built from our model has evidently added values to be combined with the existing clinical staging system.
Robust Pancreatic Ductal Adenocarcinoma Segmentation with Multi-Institutional Multi-Phase Partially-Annotated CT Scans
Aug 24, 2020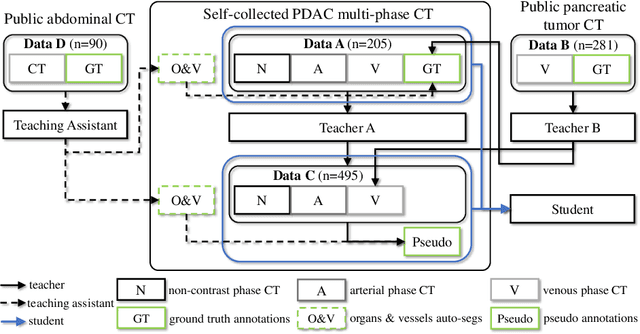
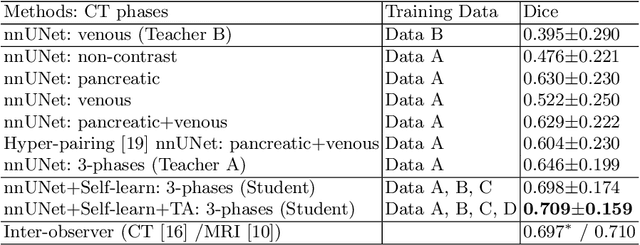
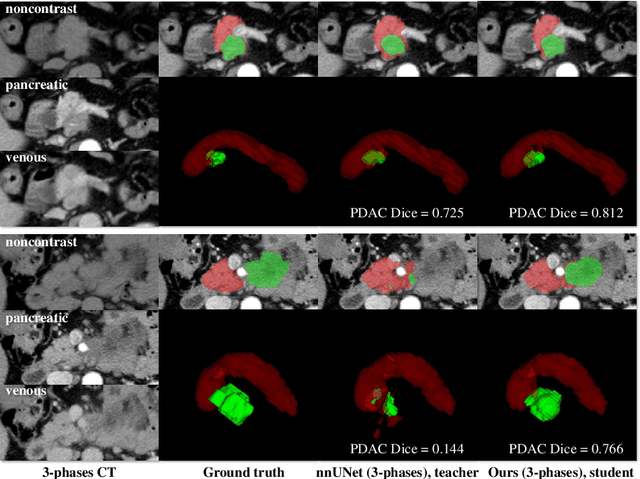
Abstract:Accurate and automated tumor segmentation is highly desired since it has the great potential to increase the efficiency and reproducibility of computing more complete tumor measurements and imaging biomarkers, comparing to (often partial) human measurements. This is probably the only viable means to enable the large-scale clinical oncology patient studies that utilize medical imaging. Deep learning approaches have shown robust segmentation performances for certain types of tumors, e.g., brain tumors in MRI imaging, when a training dataset with plenty of pixel-level fully-annotated tumor images is available. However, more than often, we are facing the challenge that only (very) limited annotations are feasible to acquire, especially for hard tumors. Pancreatic ductal adenocarcinoma (PDAC) segmentation is one of the most challenging tumor segmentation tasks, yet critically important for clinical needs. Previous work on PDAC segmentation is limited to the moderate amounts of annotated patient images (n<300) from venous or venous+arterial phase CT scans. Based on a new self-learning framework, we propose to train the PDAC segmentation model using a much larger quantity of patients (n~=1,000), with a mix of annotated and un-annotated venous or multi-phase CT images. Pseudo annotations are generated by combining two teacher models with different PDAC segmentation specialties on unannotated images, and can be further refined by a teaching assistant model that identifies associated vessels around the pancreas. A student model is trained on both manual and pseudo annotated multi-phase images. Experiment results show that our proposed method provides an absolute improvement of 6.3% Dice score over the strong baseline of nnUNet trained on annotated images, achieving the performance (Dice = 0.71) similar to the inter-observer variability between radiologists.
CT Data Curation for Liver Patients: Phase Recognition in Dynamic Contrast-Enhanced CT
Sep 27, 2019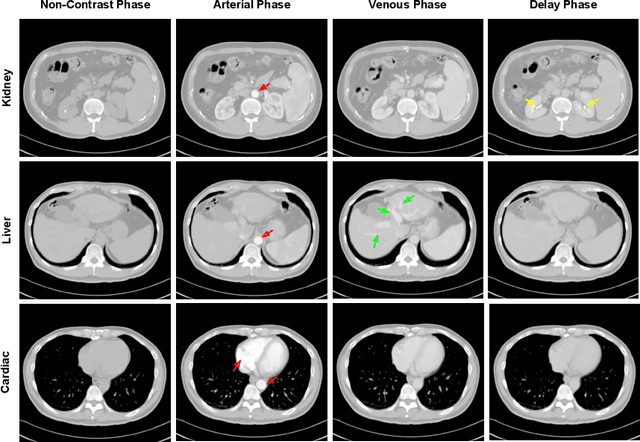
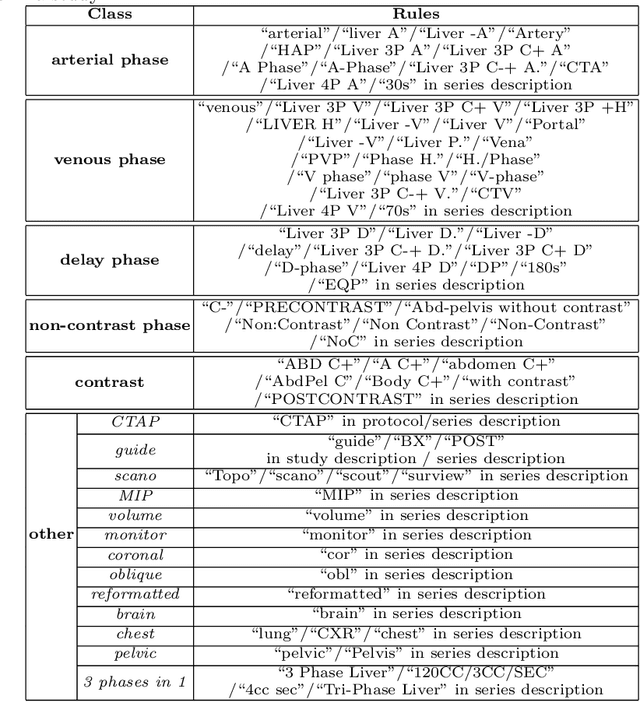
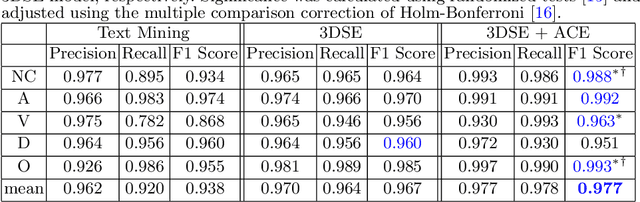
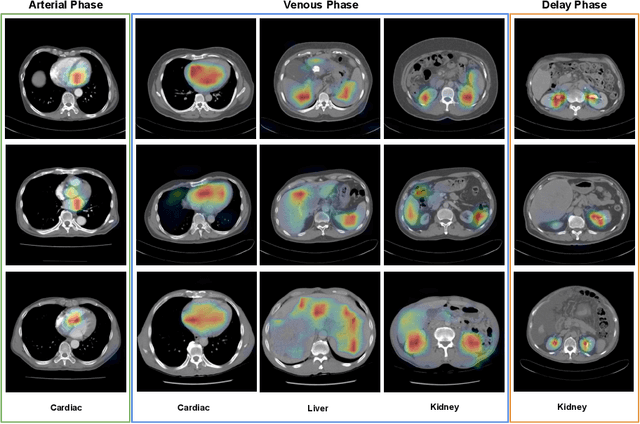

Abstract:As the demand for more descriptive machine learning models grows within medical imaging, bottlenecks due to data paucity will exacerbate. Thus, collecting enough large-scale data will require automated tools to harvest data/label pairs from messy and real-world datasets, such as hospital PACS. This is the focus of our work, where we present a principled data curation tool to extract multi-phase CT liver studies and identify each scan's phase from a real-world and heterogenous hospital PACS dataset. Emulating a typical deployment scenario, we first obtain a set of noisy labels from our institutional partners that are text mined using simple rules from DICOM tags. We train a deep learning system, using a customized and streamlined 3D SE architecture, to identify non-contrast, arterial, venous, and delay phase dynamic CT liver scans, filtering out anything else, including other types of liver contrast studies. To exploit as much training data as possible, we also introduce an aggregated cross entropy loss that can learn from scans only identified as "contrast". Extensive experiments on a dataset of 43K scans of 7680 patient imaging studies demonstrate that our 3DSE architecture, armed with our aggregated loss, can achieve a mean F1 of 0.977 and can correctly harvest up to 92.7% of studies, which significantly outperforms the text-mined and standard-loss approach, and also outperforms other, and more complex, model architectures.
Weakly Supervised Deep Learning for Thoracic Disease Classification and Localization on Chest X-rays
Jul 16, 2018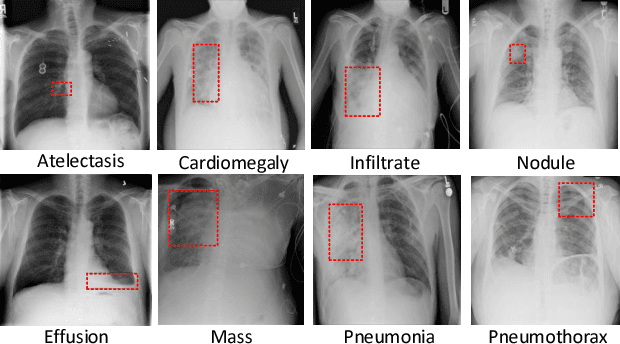
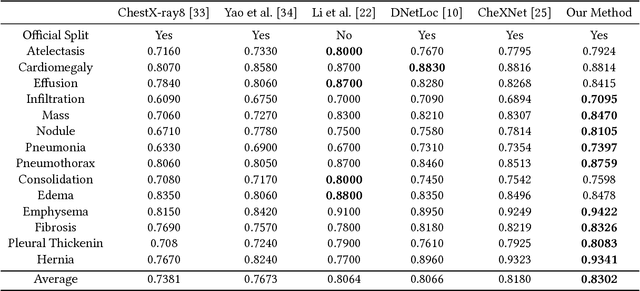
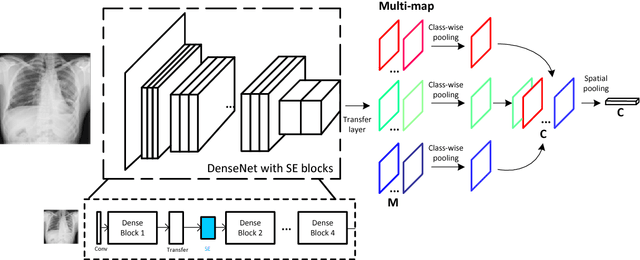
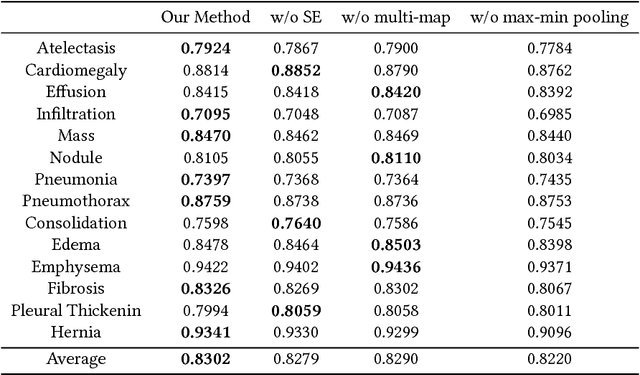
Abstract:Chest X-rays is one of the most commonly available and affordable radiological examinations in clinical practice. While detecting thoracic diseases on chest X-rays is still a challenging task for machine intelligence, due to 1) the highly varied appearance of lesion areas on X-rays from patients of different thoracic disease and 2) the shortage of accurate pixel-level annotations by radiologists for model training. Existing machine learning methods are unable to deal with the challenge that thoracic diseases usually happen in localized disease-specific areas. In this article, we propose a weakly supervised deep learning framework equipped with squeeze-and-excitation blocks, multi-map transfer, and max-min pooling for classifying thoracic diseases as well as localizing suspicious lesion regions. The comprehensive experiments and discussions are performed on the ChestX-ray14 dataset. Both numerical and visual results have demonstrated the effectiveness of the proposed model and its better performance against the state-of-the-art pipelines.
Robust Contextual Bandit via the Capped-$\ell_{2}$ norm
Aug 17, 2017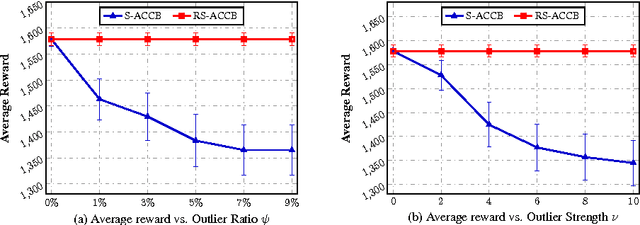
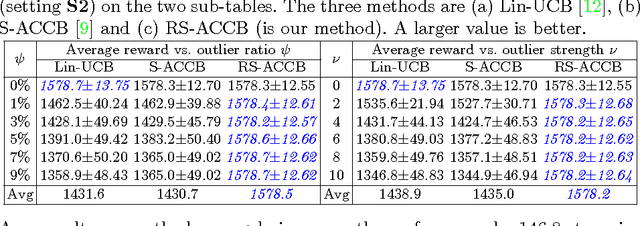
Abstract:This paper considers the actor-critic contextual bandit for the mobile health (mHealth) intervention. The state-of-the-art decision-making methods in mHealth generally assume that the noise in the dynamic system follows the Gaussian distribution. Those methods use the least-square-based algorithm to estimate the expected reward, which is prone to the existence of outliers. To deal with the issue of outliers, we propose a novel robust actor-critic contextual bandit method for the mHealth intervention. In the critic updating, the capped-$\ell_{2}$ norm is used to measure the approximation error, which prevents outliers from dominating our objective. A set of weights could be achieved from the critic updating. Considering them gives a weighted objective for the actor updating. It provides the badly noised sample in the critic updating with zero weights for the actor updating. As a result, the robustness of both actor-critic updating is enhanced. There is a key parameter in the capped-$\ell_{2}$ norm. We provide a reliable method to properly set it by making use of one of the most fundamental definitions of outliers in statistics. Extensive experiment results demonstrate that our method can achieve almost identical results compared with the state-of-the-art methods on the dataset without outliers and dramatically outperform them on the datasets noised by outliers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge