J. Alison Noble
Principled Ultrasound Data Augmentation for Classification of Standard Planes
Mar 14, 2021
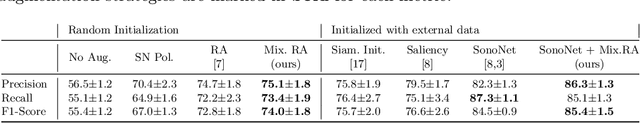
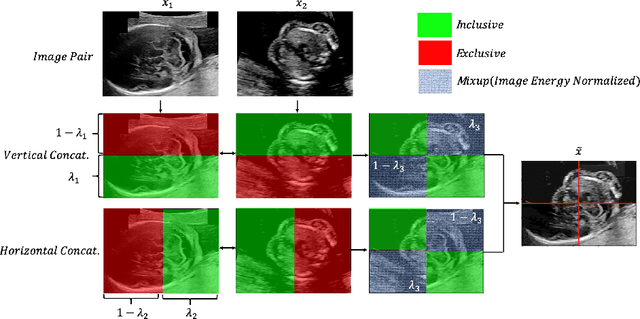


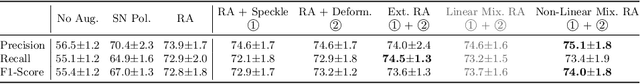
Abstract:Deep learning models with large learning capacities often overfit to medical imaging datasets. This is because training sets are often relatively small due to the significant time and financial costs incurred in medical data acquisition and labelling. Data augmentation is therefore often used to expand the availability of training data and to increase generalization. However, augmentation strategies are often chosen on an ad-hoc basis without justification. In this paper, we present an augmentation policy search method with the goal of improving model classification performance. We include in the augmentation policy search additional transformations that are often used in medical image analysis and evaluate their performance. In addition, we extend the augmentation policy search to include non-linear mixed-example data augmentation strategies. Using these learned policies, we show that principled data augmentation for medical image model training can lead to significant improvements in ultrasound standard plane detection, with an an average F1-score improvement of 7.0% overall over naive data augmentation strategies in ultrasound fetal standard plane classification. We find that the learned representations of ultrasound images are better clustered and defined with optimized data augmentation.
Cross-Task Representation Learning for Anatomical Landmark Detection
Sep 28, 2020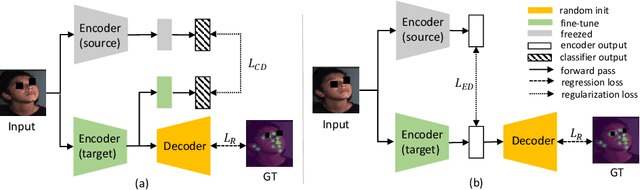
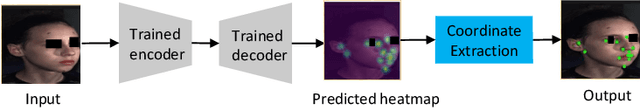

Abstract:Recently, there is an increasing demand for automatically detecting anatomical landmarks which provide rich structural information to facilitate subsequent medical image analysis. Current methods related to this task often leverage the power of deep neural networks, while a major challenge in fine tuning such models in medical applications arises from insufficient number of labeled samples. To address this, we propose to regularize the knowledge transfer across source and target tasks through cross-task representation learning. The proposed method is demonstrated for extracting facial anatomical landmarks which facilitate the diagnosis of fetal alcohol syndrome. The source and target tasks in this work are face recognition and landmark detection, respectively. The main idea of the proposed method is to retain the feature representations of the source model on the target task data, and to leverage them as an additional source of supervisory signals for regularizing the target model learning, thereby improving its performance under limited training samples. Concretely, we present two approaches for the proposed representation learning by constraining either final or intermediate model features on the target model. Experimental results on a clinical face image dataset demonstrate that the proposed approach works well with few labeled data, and outperforms other compared approaches.
Longitudinal Image Registration with Temporal-order and Subject-specificity Discrimination
Aug 29, 2020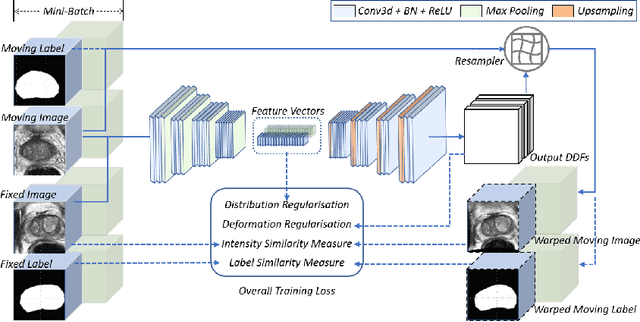

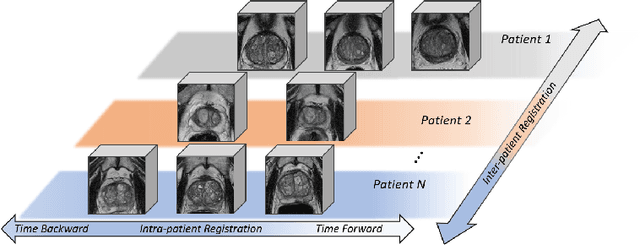
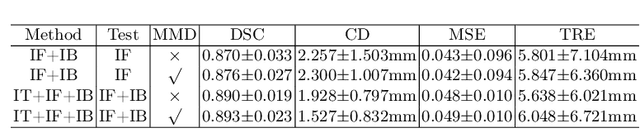
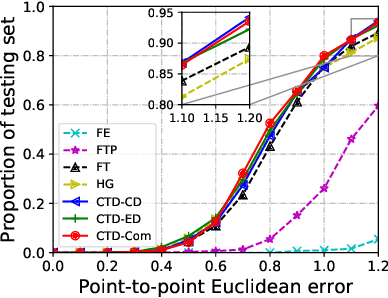
Abstract:Morphological analysis of longitudinal MR images plays a key role in monitoring disease progression for prostate cancer patients, who are placed under an active surveillance program. In this paper, we describe a learning-based image registration algorithm to quantify changes on regions of interest between a pair of images from the same patient, acquired at two different time points. Combining intensity-based similarity and gland segmentation as weak supervision, the population-data-trained registration networks significantly lowered the target registration errors (TREs) on holdout patient data, compared with those before registration and those from an iterative registration algorithm. Furthermore, this work provides a quantitative analysis on several longitudinal-data-sampling strategies and, in turn, we propose a novel regularisation method based on maximum mean discrepancy, between differently-sampled training image pairs. Based on 216 3D MR images from 86 patients, we report a mean TRE of 5.6 mm and show statistically significant differences between the different training data sampling strategies.
Self-Supervised Ultrasound to MRI Fetal Brain Image Synthesis
Aug 19, 2020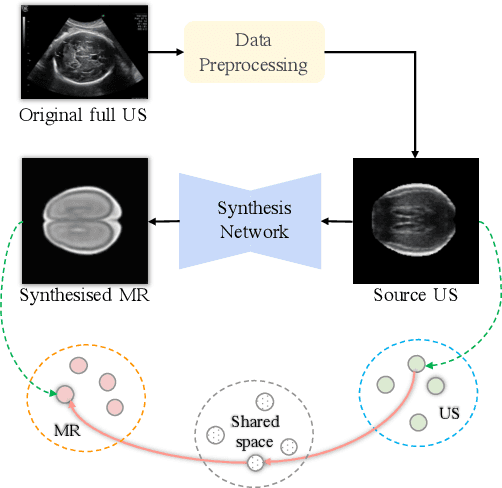
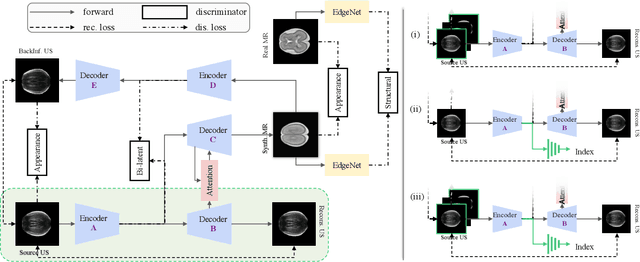
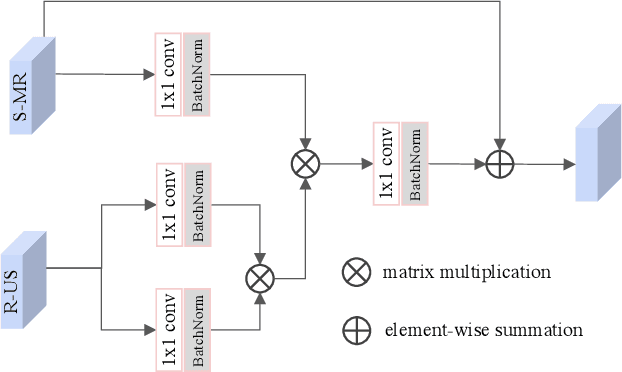
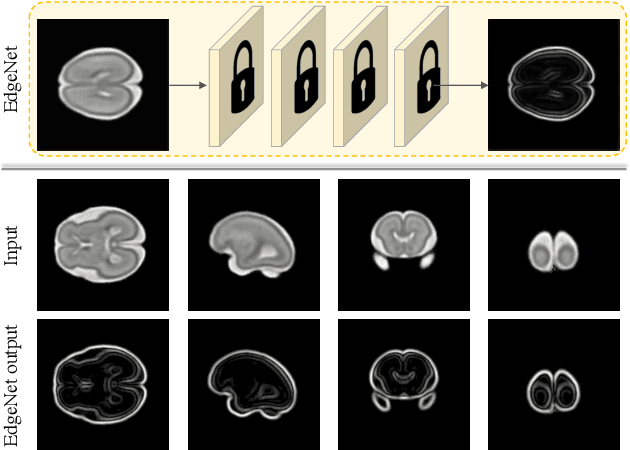
Abstract:Fetal brain magnetic resonance imaging (MRI) offers exquisite images of the developing brain but is not suitable for second-trimester anomaly screening, for which ultrasound (US) is employed. Although expert sonographers are adept at reading US images, MR images which closely resemble anatomical images are much easier for non-experts to interpret. Thus in this paper we propose to generate MR-like images directly from clinical US images. In medical image analysis such a capability is potentially useful as well, for instance for automatic US-MRI registration and fusion. The proposed model is end-to-end trainable and self-supervised without any external annotations. Specifically, based on an assumption that the US and MRI data share a similar anatomical latent space, we first utilise a network to extract the shared latent features, which are then used for MRI synthesis. Since paired data is unavailable for our study (and rare in practice), pixel-level constraints are infeasible to apply. We instead propose to enforce the distributions to be statistically indistinguishable, by adversarial learning in both the image domain and feature space. To regularise the anatomical structures between US and MRI during synthesis, we further propose an adversarial structural constraint. A new cross-modal attention technique is proposed to utilise non-local spatial information, by encouraging multi-modal knowledge fusion and propagation. We extend the approach to consider the case where 3D auxiliary information (e.g., 3D neighbours and a 3D location index) from volumetric data is also available, and show that this improves image synthesis. The proposed approach is evaluated quantitatively and qualitatively with comparison to real fetal MR images and other approaches to synthesis, demonstrating its feasibility of synthesising realistic MR images.
Self-supervised Contrastive Video-Speech Representation Learning for Ultrasound
Aug 14, 2020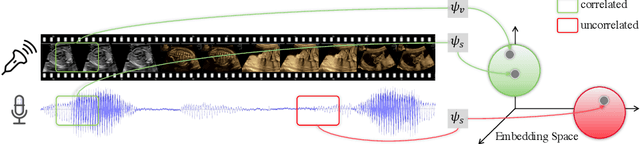
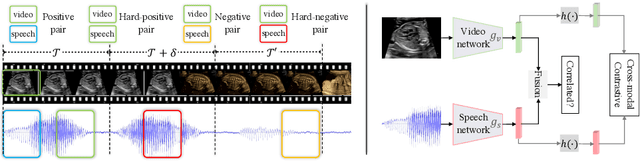


Abstract:In medical imaging, manual annotations can be expensive to acquire and sometimes infeasible to access, making conventional deep learning-based models difficult to scale. As a result, it would be beneficial if useful representations could be derived from raw data without the need for manual annotations. In this paper, we propose to address the problem of self-supervised representation learning with multi-modal ultrasound video-speech raw data. For this case, we assume that there is a high correlation between the ultrasound video and the corresponding narrative speech audio of the sonographer. In order to learn meaningful representations, the model needs to identify such correlation and at the same time understand the underlying anatomical features. We designed a framework to model the correspondence between video and audio without any kind of human annotations. Within this framework, we introduce cross-modal contrastive learning and an affinity-aware self-paced learning scheme to enhance correlation modelling. Experimental evaluations on multi-modal fetal ultrasound video and audio show that the proposed approach is able to learn strong representations and transfers well to downstream tasks of standard plane detection and eye-gaze prediction.
Automatic Probe Movement Guidance for Freehand Obstetric Ultrasound
Jul 08, 2020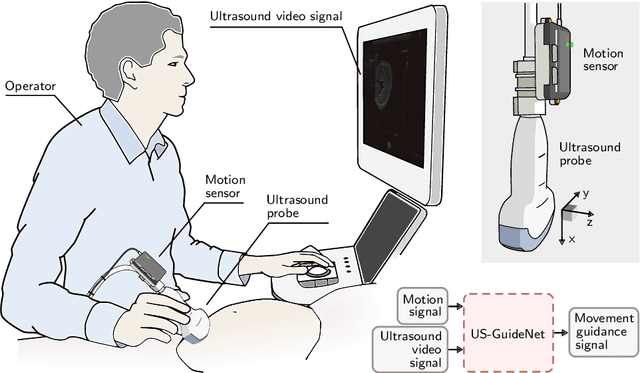
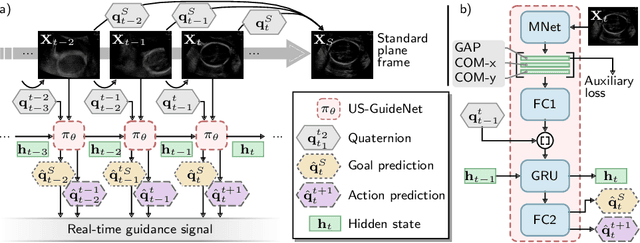
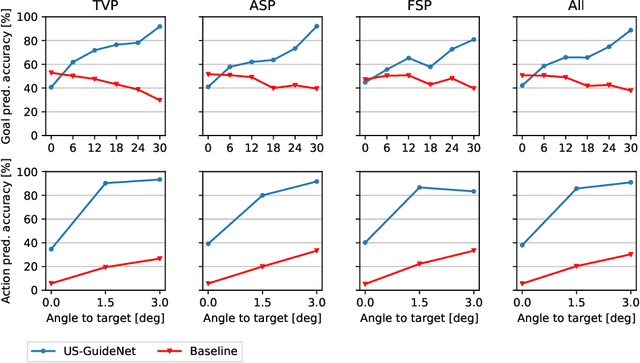
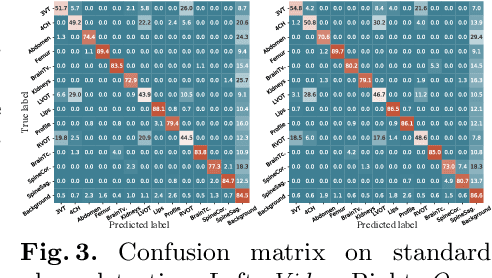
Abstract:We present the first system that provides real-time probe movement guidance for acquiring standard planes in routine freehand obstetric ultrasound scanning. Such a system can contribute to the worldwide deployment of obstetric ultrasound scanning by lowering the required level of operator expertise. The system employs an artificial neural network that receives the ultrasound video signal and the motion signal of an inertial measurement unit (IMU) that is attached to the probe, and predicts a guidance signal. The network termed US-GuideNet predicts either the movement towards the standard plane position (goal prediction), or the next movement that an expert sonographer would perform (action prediction). While existing models for other ultrasound applications are trained with simulations or phantoms, we train our model with real-world ultrasound video and probe motion data from 464 routine clinical scans by 17 accredited sonographers. Evaluations for 3 standard plane types show that the model provides a useful guidance signal with an accuracy of 88.8% for goal prediction and 90.9% for action prediction.
Unified Image and Video Saliency Modeling
Mar 11, 2020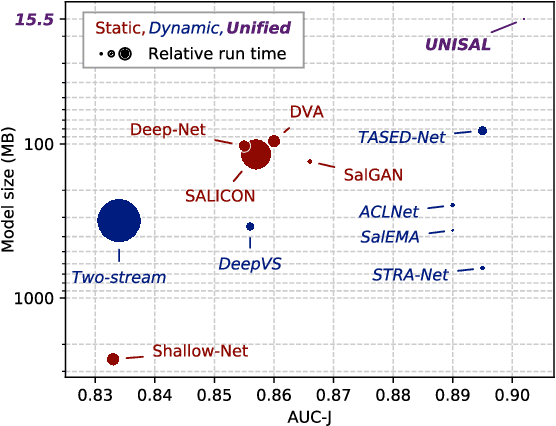

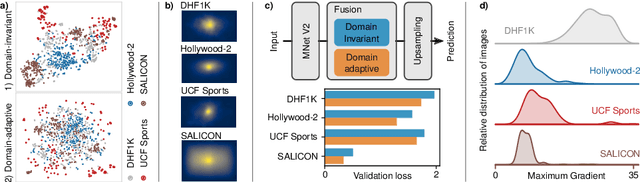
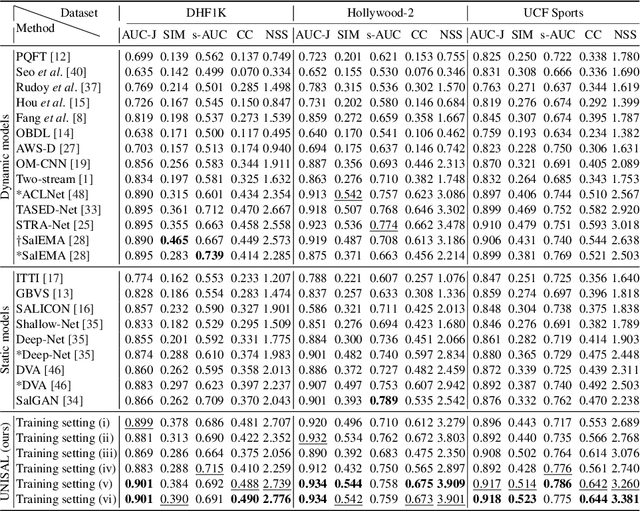
Abstract:Visual saliency modeling for images and videos is treated as two independent tasks in recent computer vision literature. On the one hand, image saliency modeling is a well-studied problem and progress on benchmarks like \mbox{SALICON} and MIT300 is slowing. For video saliency prediction on the other hand, rapid gains have been achieved on the recent DHF1K benchmark through network architectures that are optimized for this task. Here, we take a step back and ask: Can image and video saliency modeling be approached via a unified model, with mutual benefit? We find that it is crucial to model the domain shift between image and video saliency data and between different video saliency datasets for effective joint modeling. We identify different sources of domain shift and address them through four novel domain adaptation techniques - Domain-Adaptive Priors, Domain-Adaptive Fusion, Domain-Adaptive Smoothing and Bypass-RNN - in addition to an improved formulation of learned Gaussian priors. We integrate these techniques into a simple and lightweight encoder-RNN-decoder-style network, UNISAL, and train the entire network simultaneously with image and video saliency data. We evaluate our method on the video saliency datasets DHF1K, Hollywood-2 and UCF-Sports, as well as the image saliency datasets SALICON and MIT300. With one set of parameters, our method achieves state-of-the-art performance on all video saliency datasets and is on par with the state-of-the-art for image saliency prediction, despite a 5 to 20-fold reduction in model size and the fastest runtime among all competing deep models. We provide retrospective analyses and ablation studies which demonstrate the importance of the domain shift modeling. The code is available at https://github.com/rdroste/unisal.
Self-supervised Representation Learning for Ultrasound Video
Feb 28, 2020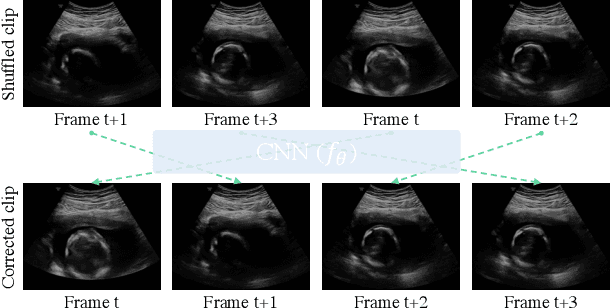
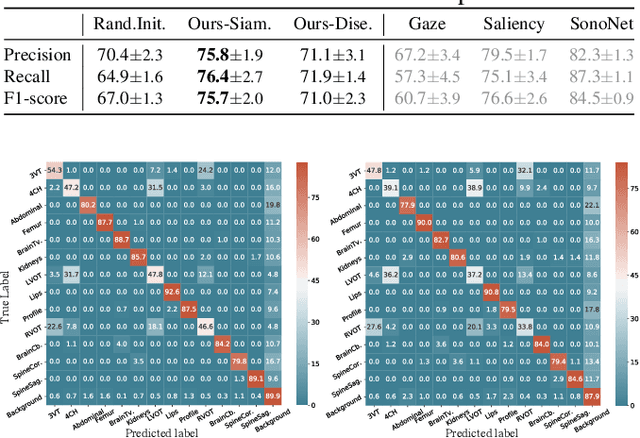
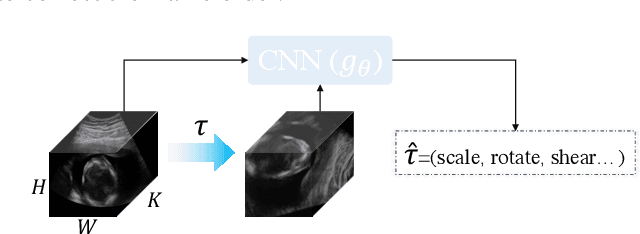
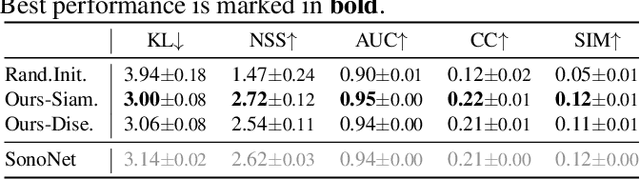
Abstract:Recent advances in deep learning have achieved promising performance for medical image analysis, while in most cases ground-truth annotations from human experts are necessary to train the deep model. In practice, such annotations are expensive to collect and can be scarce for medical imaging applications. Therefore, there is significant interest in learning representations from unlabelled raw data. In this paper, we propose a self-supervised learning approach to learn meaningful and transferable representations from medical imaging video without any type of human annotation. We assume that in order to learn such a representation, the model should identify anatomical structures from the unlabelled data. Therefore we force the model to address anatomy-aware tasks with free supervision from the data itself. Specifically, the model is designed to correct the order of a reshuffled video clip and at the same time predict the geometric transformation applied to the video clip. Experiments on fetal ultrasound video show that the proposed approach can effectively learn meaningful and strong representations, which transfer well to downstream tasks like standard plane detection and saliency prediction.
Discovering Salient Anatomical Landmarks by Predicting Human Gaze
Jan 22, 2020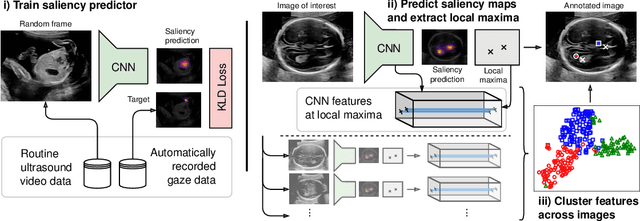
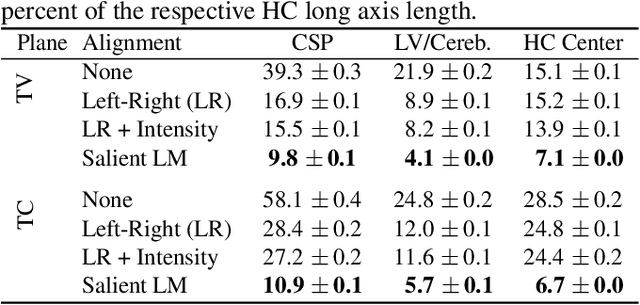
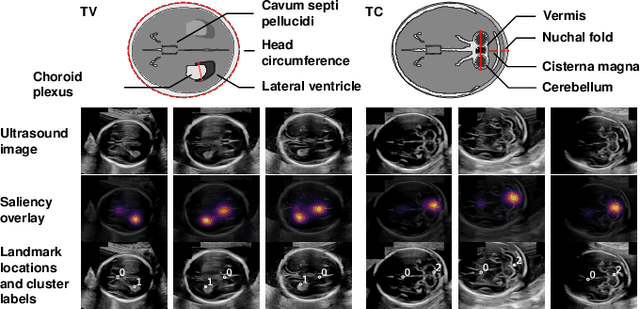
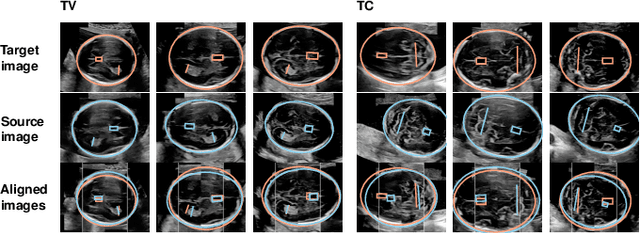
Abstract:Anatomical landmarks are a crucial prerequisite for many medical imaging tasks. Usually, the set of landmarks for a given task is predefined by experts. The landmark locations for a given image are then annotated manually or via machine learning methods trained on manual annotations. In this paper, in contrast, we present a method to automatically discover and localize anatomical landmarks in medical images. Specifically, we consider landmarks that attract the visual attention of humans, which we term visually salient landmarks. We illustrate the method for fetal neurosonographic images. First, full-length clinical fetal ultrasound scans are recorded with live sonographer gaze-tracking. Next, a convolutional neural network (CNN) is trained to predict the gaze point distribution (saliency map) of the sonographers on scan video frames. The CNN is then used to predict saliency maps of unseen fetal neurosonographic images, and the landmarks are extracted as the local maxima of these saliency maps. Finally, the landmarks are matched across images by clustering the landmark CNN features. We show that the discovered landmarks can be used within affine image registration, with average landmark alignment errors between 4.1% and 10.9% of the fetal head long axis length.
Heterogeneous tissue characterization using ultrasound: a comparison of fractal analysis backscatter models on liver tumors
Dec 20, 2019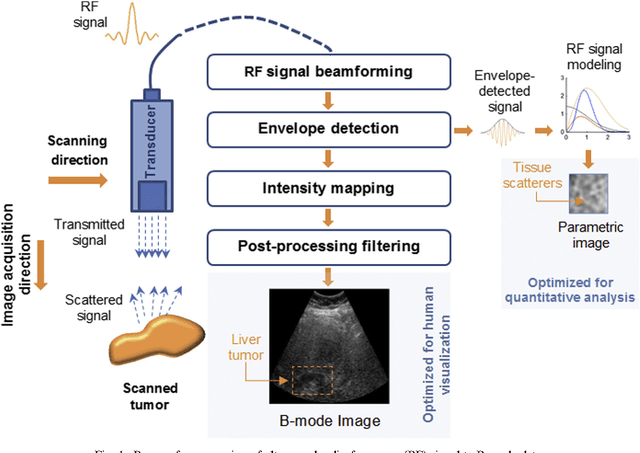
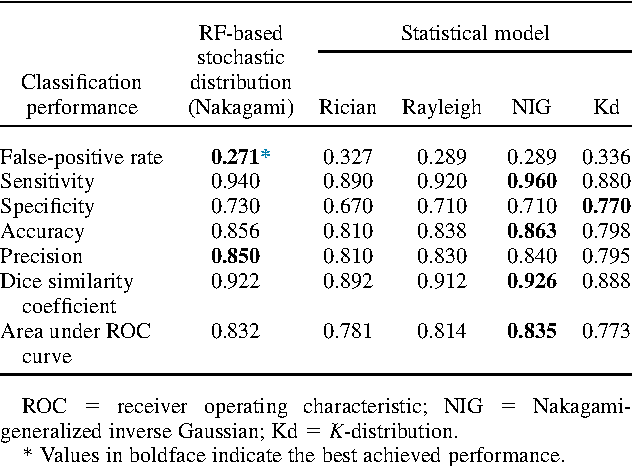
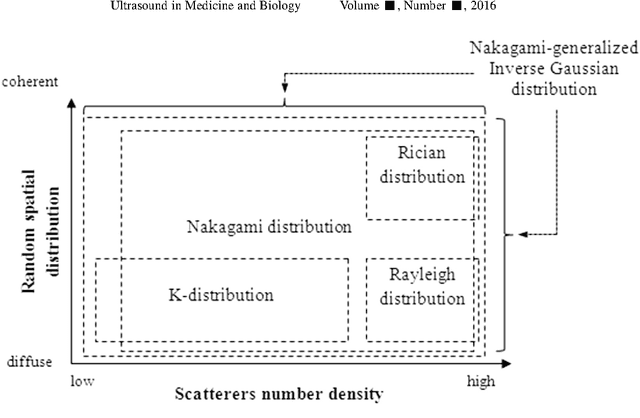
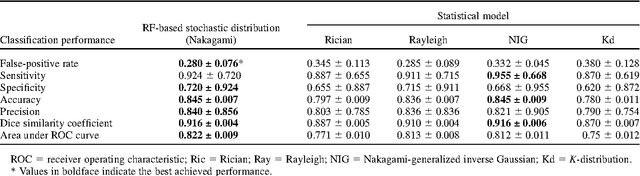
Abstract:Assessing tumor tissue heterogeneity via ultrasound has recently been suggested for predicting early response to treatment. The ultrasound backscattering characteristics can assist in better understanding the tumor texture by highlighting local concentration and spatial arrangement of tissue scatterers. However, it is challenging to quantify the various tissue heterogeneities ranging from fine-to-coarse of the echo envelope peaks in tumor texture. Local parametric fractal features extracted via maximum likelihood estimation from five well-known statistical model families are evaluated for the purpose of ultrasound tissue characterization. The fractal dimension (self-similarity measure) was used to characterize the spatial distribution of scatterers, while the Lacunarity (sparsity measure) was applied to determine scatterer number density. Performance was assessed based on 608 cross-sectional clinical ultrasound RF images of liver tumors (230 and 378 demonstrating respondent and non-respondent cases, respectively). Crossvalidation via leave-one-tumor-out and with different k-folds methodologies using a Bayesian classifier were employed for validation. The fractal properties of the backscattered echoes based on the Nakagami model (Nkg) and its extend four-parameter Nakagami-generalized inverse Gaussian (NIG) distribution achieved best results - with nearly similar performance - for characterizing liver tumor tissue. Accuracy, sensitivity and specificity for the Nkg/NIG were: 85.6%/86.3%, 94.0%/96.0%, and 73.0%/71.0%, respectively. Other statistical models, such as the Rician, Rayleigh, and K-distribution were found to not be as effective in characterizing the subtle changes in tissue texture as an indication of response to treatment. Employing the most relevant and practical statistical model could have potential consequences for the design of an early and effective clinical therapy.
* 31 pages, 7 figures, 3 tables, journal article
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge