Daniel Y. F. Chung
Heterogeneous tissue characterization using ultrasound: a comparison of fractal analysis backscatter models on liver tumors
Dec 20, 2019
Abstract:Assessing tumor tissue heterogeneity via ultrasound has recently been suggested for predicting early response to treatment. The ultrasound backscattering characteristics can assist in better understanding the tumor texture by highlighting local concentration and spatial arrangement of tissue scatterers. However, it is challenging to quantify the various tissue heterogeneities ranging from fine-to-coarse of the echo envelope peaks in tumor texture. Local parametric fractal features extracted via maximum likelihood estimation from five well-known statistical model families are evaluated for the purpose of ultrasound tissue characterization. The fractal dimension (self-similarity measure) was used to characterize the spatial distribution of scatterers, while the Lacunarity (sparsity measure) was applied to determine scatterer number density. Performance was assessed based on 608 cross-sectional clinical ultrasound RF images of liver tumors (230 and 378 demonstrating respondent and non-respondent cases, respectively). Crossvalidation via leave-one-tumor-out and with different k-folds methodologies using a Bayesian classifier were employed for validation. The fractal properties of the backscattered echoes based on the Nakagami model (Nkg) and its extend four-parameter Nakagami-generalized inverse Gaussian (NIG) distribution achieved best results - with nearly similar performance - for characterizing liver tumor tissue. Accuracy, sensitivity and specificity for the Nkg/NIG were: 85.6%/86.3%, 94.0%/96.0%, and 73.0%/71.0%, respectively. Other statistical models, such as the Rician, Rayleigh, and K-distribution were found to not be as effective in characterizing the subtle changes in tissue texture as an indication of response to treatment. Employing the most relevant and practical statistical model could have potential consequences for the design of an early and effective clinical therapy.
* 31 pages, 7 figures, 3 tables, journal article
Quantification of Ultrasonic Texture heterogeneity via Volumetric Stochastic Modeling for Tissue Characterization
Jan 14, 2016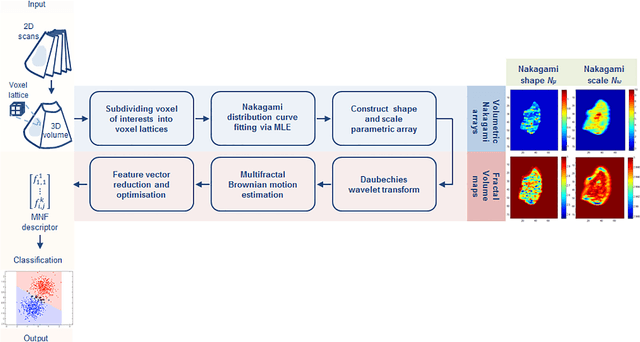


Abstract:Intensity variations in image texture can provide powerful quantitative information about physical properties of biological tissue. However, tissue patterns can vary according to the utilized imaging system and are intrinsically correlated to the scale of analysis. In the case of ultrasound, the Nakagami distribution is a general model of the ultrasonic backscattering envelope under various scattering conditions and densities where it can be employed for characterizing image texture, but the subtle intra-heterogeneities within a given mass are difficult to capture via this model as it works at a single spatial scale. This paper proposes a locally adaptive 3D multi-resolution Nakagami-based fractal feature descriptor that extends Nakagami-based texture analysis to accommodate subtle speckle spatial frequency tissue intensity variability in volumetric scans. Local textural fractal descriptors - which are invariant to affine intensity changes - are extracted from volumetric patches at different spatial resolutions from voxel lattice-based generated shape and scale Nakagami parameters. Using ultrasound radio-frequency datasets we found that after applying an adaptive fractal decomposition label transfer approach on top of the generated Nakagami voxels, tissue characterization results were superior to the state of art. Experimental results on real 3D ultrasonic pre-clinical and clinical datasets suggest that describing tumor intra-heterogeneity via this descriptor may facilitate improved prediction of therapy response and disease characterization.
* Supplementary data associated with this article can be found, in the online version, at http://dx.doi.org/10.1016/j.media.2014.12. 004
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge