Huangjing Lin
HMIL: Hierarchical Multi-Instance Learning for Fine-Grained Whole Slide Image Classification
Nov 12, 2024



Abstract:Fine-grained classification of whole slide images (WSIs) is essential in precision oncology, enabling precise cancer diagnosis and personalized treatment strategies. The core of this task involves distinguishing subtle morphological variations within the same broad category of gigapixel-resolution images, which presents a significant challenge. While the multi-instance learning (MIL) paradigm alleviates the computational burden of WSIs, existing MIL methods often overlook hierarchical label correlations, treating fine-grained classification as a flat multi-class classification task. To overcome these limitations, we introduce a novel hierarchical multi-instance learning (HMIL) framework. By facilitating on the hierarchical alignment of inherent relationships between different hierarchy of labels at instance and bag level, our approach provides a more structured and informative learning process. Specifically, HMIL incorporates a class-wise attention mechanism that aligns hierarchical information at both the instance and bag levels. Furthermore, we introduce supervised contrastive learning to enhance the discriminative capability for fine-grained classification and a curriculum-based dynamic weighting module to adaptively balance the hierarchical feature during training. Extensive experiments on our large-scale cytology cervical cancer (CCC) dataset and two public histology datasets, BRACS and PANDA, demonstrate the state-of-the-art class-wise and overall performance of our HMIL framework. Our source code is available at https://github.com/ChengJin-git/HMIL.
Holistic and Historical Instance Comparison for Cervical Cell Detection
Sep 21, 2024



Abstract:Cytology screening from Papanicolaou (Pap) smears is a common and effective tool for the preventive clinical management of cervical cancer, where abnormal cell detection from whole slide images serves as the foundation for reporting cervical cytology. However, cervical cell detection remains challenging due to 1) hazily-defined cell types (e.g., ASC-US) with subtle morphological discrepancies caused by the dynamic cancerization process, i.e., cell class ambiguity, and 2) imbalanced class distributions of clinical data may cause missed detection, especially for minor categories, i.e., cell class imbalance. To this end, we propose a holistic and historical instance comparison approach for cervical cell detection. Specifically, we first develop a holistic instance comparison scheme enforcing both RoI-level and class-level cell discrimination. This coarse-to-fine cell comparison encourages the model to learn foreground-distinguishable and class-wise representations. To emphatically improve the distinguishability of minor classes, we then introduce a historical instance comparison scheme with a confident sample selection-based memory bank, which involves comparing current embeddings with historical embeddings for better cell instance discrimination. Extensive experiments and analysis on two large-scale cytology datasets including 42,592 and 114,513 cervical cells demonstrate the effectiveness of our method. The code is available at https://github.com/hjiangaz/HERO.
GAInS: Gradient Anomaly-aware Biomedical Instance Segmentation
Sep 21, 2024



Abstract:Instance segmentation plays a vital role in the morphological quantification of biomedical entities such as tissues and cells, enabling precise identification and delineation of different structures. Current methods often address the challenges of touching, overlapping or crossing instances through individual modeling, while neglecting the intrinsic interrelation between these conditions. In this work, we propose a Gradient Anomaly-aware Biomedical Instance Segmentation approach (GAInS), which leverages instance gradient information to perceive local gradient anomaly regions, thus modeling the spatial relationship between instances and refining local region segmentation. Specifically, GAInS is firstly built on a Gradient Anomaly Mapping Module (GAMM), which encodes the radial fields of instances through window sliding to obtain instance gradient anomaly maps. To efficiently refine boundaries and regions with gradient anomaly attention, we propose an Adaptive Local Refinement Module (ALRM) with a gradient anomaly-aware loss function. Extensive comparisons and ablation experiments in three biomedical scenarios demonstrate that our proposed GAInS outperforms other state-of-the-art (SOTA) instance segmentation methods. The code is available at https://github.com/DeepGAInS/GAInS.
A Multimodal Knowledge-enhanced Whole-slide Pathology Foundation Model
Jul 22, 2024



Abstract:Remarkable strides in computational pathology have been made in the task-agnostic foundation model that advances the performance of a wide array of downstream clinical tasks. Despite the promising performance, there are still several challenges. First, prior works have resorted to either vision-only or vision-captions data, disregarding invaluable pathology reports and gene expression profiles which respectively offer distinct knowledge for versatile clinical applications. Second, the current progress in pathology FMs predominantly concentrates on the patch level, where the restricted context of patch-level pretraining fails to capture whole-slide patterns. Here we curated the largest multimodal dataset consisting of H\&E diagnostic whole slide images and their associated pathology reports and RNA-Seq data, resulting in 26,169 slide-level modality pairs from 10,275 patients across 32 cancer types. To leverage these data for CPath, we propose a novel whole-slide pretraining paradigm which injects multimodal knowledge at the whole-slide context into the pathology FM, called Multimodal Self-TAught PRetraining (mSTAR). The proposed paradigm revolutionizes the workflow of pretraining for CPath, which enables the pathology FM to acquire the whole-slide context. To our knowledge, this is the first attempt to incorporate multimodal knowledge at the slide level for enhancing pathology FMs, expanding the modelling context from unimodal to multimodal knowledge and from patch-level to slide-level. To systematically evaluate the capabilities of mSTAR, extensive experiments including slide-level unimodal and multimodal applications, are conducted across 7 diverse types of tasks on 43 subtasks, resulting in the largest spectrum of downstream tasks. The average performance in various slide-level applications consistently demonstrates significant performance enhancements for mSTAR compared to SOTA FMs.
Deep Omni-supervised Learning for Rib Fracture Detection from Chest Radiology Images
Jun 23, 2023



Abstract:Deep learning (DL)-based rib fracture detection has shown promise of playing an important role in preventing mortality and improving patient outcome. Normally, developing DL-based object detection models requires huge amount of bounding box annotation. However, annotating medical data is time-consuming and expertise-demanding, making obtaining a large amount of fine-grained annotations extremely infeasible. This poses pressing need of developing label-efficient detection models to alleviate radiologists' labeling burden. To tackle this challenge, the literature of object detection has witnessed an increase of weakly-supervised and semi-supervised approaches, yet still lacks a unified framework that leverages various forms of fully-labeled, weakly-labeled, and unlabeled data. In this paper, we present a novel omni-supervised object detection network, ORF-Netv2, to leverage as much available supervision as possible. Specifically, a multi-branch omni-supervised detection head is introduced with each branch trained with a specific type of supervision. A co-training-based dynamic label assignment strategy is then proposed to enable flexibly and robustly learning from the weakly-labeled and unlabeled data. Extensively evaluation was conducted for the proposed framework with three rib fracture datasets on both chest CT and X-ray. By leveraging all forms of supervision, ORF-Netv2 achieves mAPs of 34.7, 44.7, and 19.4 on the three datasets, respectively, surpassing the baseline detector which uses only box annotations by mAP gains of 3.8, 4.8, and 5.0, respectively. Furthermore, ORF-Netv2 consistently outperforms other competitive label-efficient methods over various scenarios, showing a promising framework for label-efficient fracture detection.
Scale-aware Super-resolution Network with Dual Affinity Learning for Lesion Segmentation from Medical Images
May 30, 2023Abstract:Convolutional Neural Networks (CNNs) have shown remarkable progress in medical image segmentation. However, lesion segmentation remains a challenge to state-of-the-art CNN-based algorithms due to the variance in scales and shapes. On the one hand, tiny lesions are hard to be delineated precisely from the medical images which are often of low resolutions. On the other hand, segmenting large-size lesions requires large receptive fields, which exacerbates the first challenge. In this paper, we present a scale-aware super-resolution network to adaptively segment lesions of various sizes from the low-resolution medical images. Our proposed network contains dual branches to simultaneously conduct lesion mask super-resolution and lesion image super-resolution. The image super-resolution branch will provide more detailed features for the segmentation branch, i.e., the mask super-resolution branch, for fine-grained segmentation. Meanwhile, we introduce scale-aware dilated convolution blocks into the multi-task decoders to adaptively adjust the receptive fields of the convolutional kernels according to the lesion sizes. To guide the segmentation branch to learn from richer high-resolution features, we propose a feature affinity module and a scale affinity module to enhance the multi-task learning of the dual branches. On multiple challenging lesion segmentation datasets, our proposed network achieved consistent improvements compared to other state-of-the-art methods.
ORF-Net: Deep Omni-supervised Rib Fracture Detection from Chest CT Scans
Jul 05, 2022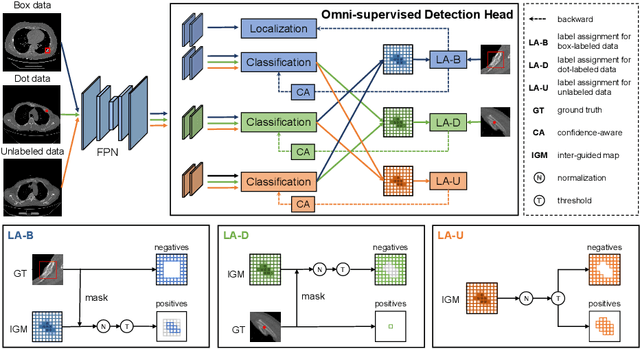
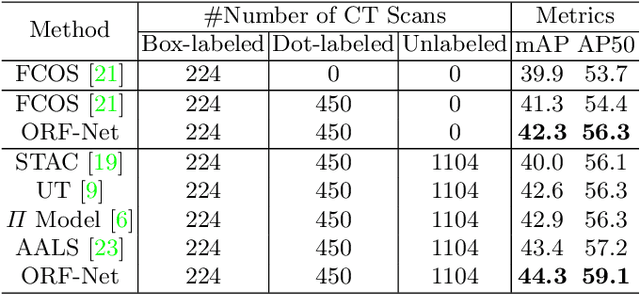
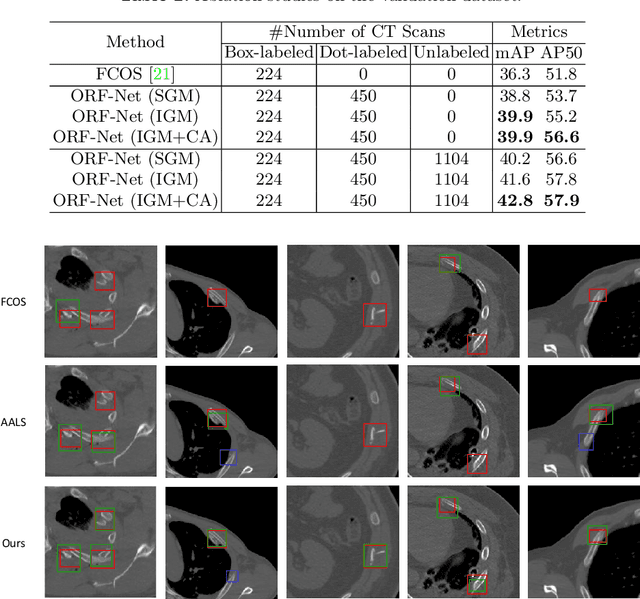
Abstract:Most of the existing object detection works are based on the bounding box annotation: each object has a precise annotated box. However, for rib fractures, the bounding box annotation is very labor-intensive and time-consuming because radiologists need to investigate and annotate the rib fractures on a slice-by-slice basis. Although a few studies have proposed weakly-supervised methods or semi-supervised methods, they could not handle different forms of supervision simultaneously. In this paper, we proposed a novel omni-supervised object detection network, which can exploit multiple different forms of annotated data to further improve the detection performance. Specifically, the proposed network contains an omni-supervised detection head, in which each form of annotation data corresponds to a unique classification branch. Furthermore, we proposed a dynamic label assignment strategy for different annotated forms of data to facilitate better learning for each branch. Moreover, we also design a confidence-aware classification loss to emphasize the samples with high confidence and further improve the model's performance. Extensive experiments conducted on the testing dataset show our proposed method outperforms other state-of-the-art approaches consistently, demonstrating the efficacy of deep omni-supervised learning on improving rib fracture detection performance.
Deep Semi-supervised Metric Learning with Dual Alignment for Cervical Cancer Cell Detection
Apr 07, 2021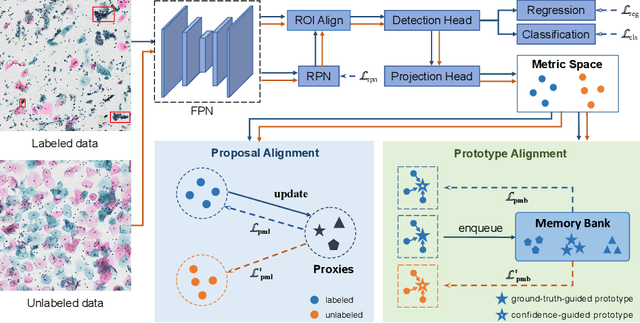
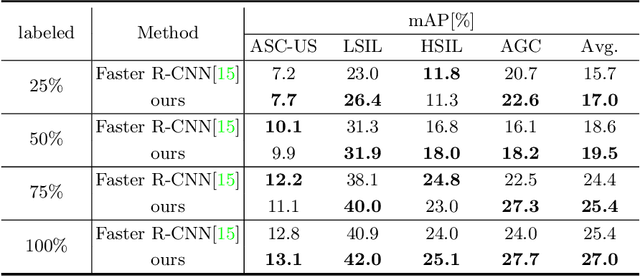
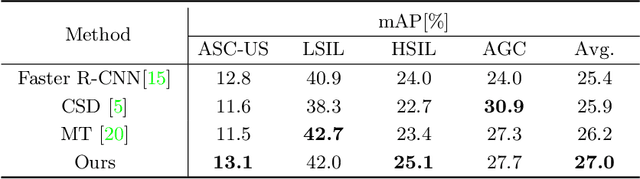
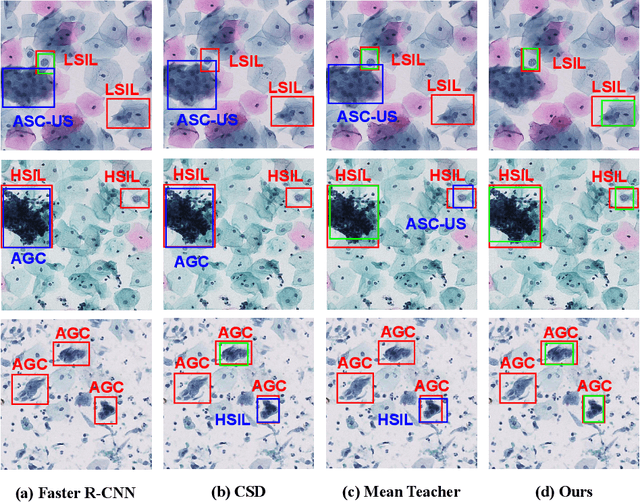
Abstract:With availability of huge amounts of labeled data, deep learning has achieved unprecedented success in various object detection tasks. However, large-scale annotations for medical images are extremely challenging to be acquired due to the high demand of labour and expertise. To address this difficult issue, in this paper we propose a novel semi-supervised deep metric learning method to effectively leverage both labeled and unlabeled data with application to cervical cancer cell detection. Different from previous methods, our model learns an embedding metric space and conducts dual alignment of semantic features on both the proposal and prototype levels. First, on the proposal level, we generate pseudo labels for the unlabeled data to align the proposal features with learnable class proxies derived from the labeled data. Furthermore, we align the prototypes generated from each mini-batch of labeled and unlabeled data to alleviate the influence of possibly noisy pseudo labels. Moreover, we adopt a memory bank to store the labeled prototypes and hence significantly enrich the metric learning information from larger batches. To comprehensively validate the method, we construct a large-scale dataset for semi-supervised cervical cancer cell detection for the first time, consisting of 240,860 cervical cell images in total. Extensive experiments show our proposed method outperforms other state-of-the-art semi-supervised approaches consistently, demonstrating efficacy of deep semi-supervised metric learning with dual alignment on improving cervical cancer cell detection performance.
Dual-Consistency Semi-Supervised Learning with Uncertainty Quantification for COVID-19 Lesion Segmentation from CT Images
Apr 07, 2021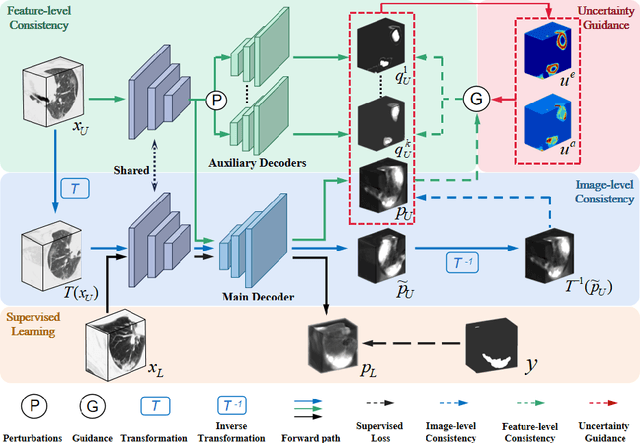
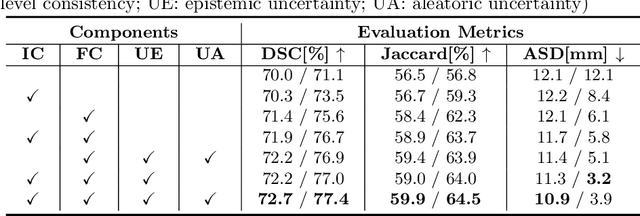
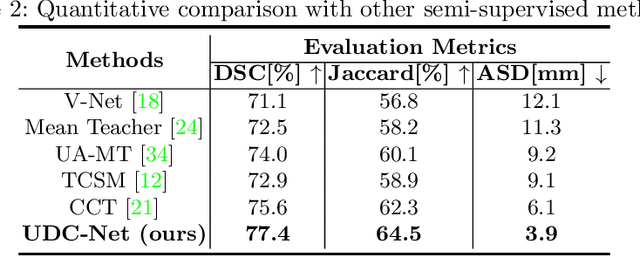
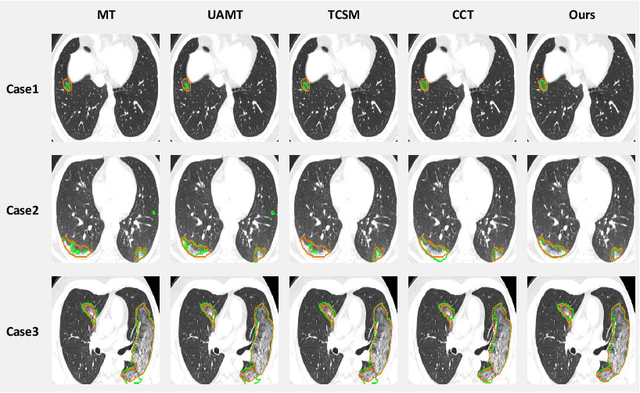
Abstract:The novel coronavirus disease 2019 (COVID-19) characterized by atypical pneumonia has caused millions of deaths worldwide. Automatically segmenting lesions from chest Computed Tomography (CT) is a promising way to assist doctors in COVID-19 screening, treatment planning, and follow-up monitoring. However, voxel-wise annotations are extremely expert-demanding and scarce, especially when it comes to novel diseases, while an abundance of unlabeled data could be available. To tackle the challenge of limited annotations, in this paper, we propose an uncertainty-guided dual-consistency learning network (UDC-Net) for semi-supervised COVID-19 lesion segmentation from CT images. Specifically, we present a dual-consistency learning scheme that simultaneously imposes image transformation equivalence and feature perturbation invariance to effectively harness the knowledge from unlabeled data. We then quantify both the epistemic uncertainty and the aleatoric uncertainty and employ them together to guide the consistency regularization for more reliable unsupervised learning. Extensive experiments showed that our proposed UDC-Net improves the fully supervised method by 6.3% in Dice and outperforms other competitive semi-supervised approaches by significant margins, demonstrating high potential in real-world clinical practice.
OXnet: Omni-supervised Thoracic Disease Detection from Chest X-rays
Apr 07, 2021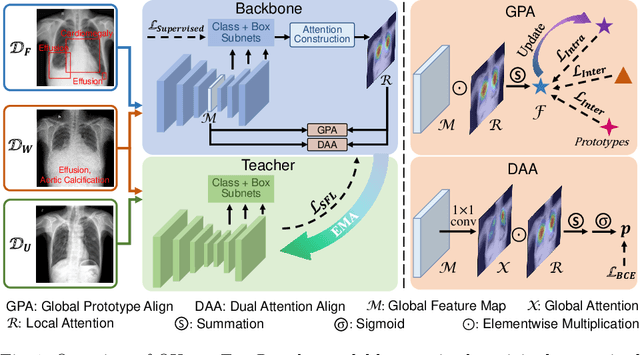
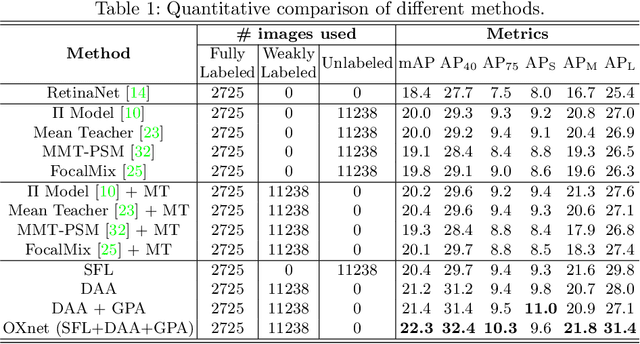
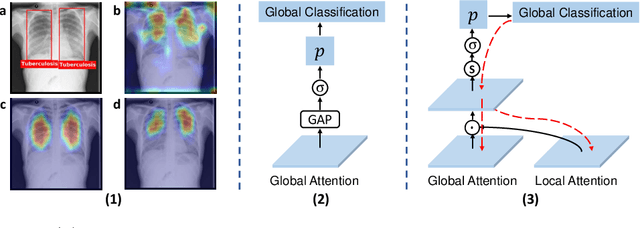
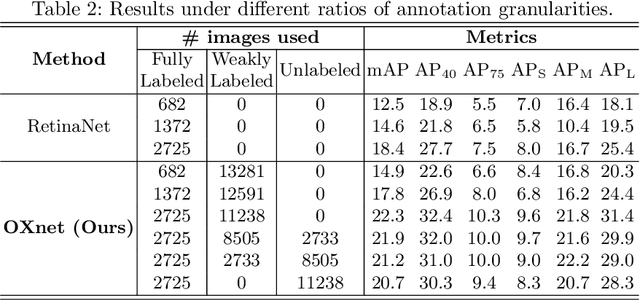
Abstract:Chest X-ray (CXR) is the most typical medical image worldwide to examine various thoracic diseases. Automatically localizing lesions from CXR is a promising way to alleviate radiologists' daily reading burden. However, CXR datasets often have numerous image-level annotations and scarce lesion-level annotations, and more often, without annotations. Thus far, unifying different supervision granularities to develop thoracic disease detection algorithms has not been comprehensively addressed. In this paper, we present OXnet, the first deep omni-supervised thoracic disease detection network to our best knowledge that uses as much available supervision as possible for CXR diagnosis. Besides fully supervised learning, to enable learning from weakly-annotated data, we guide the information from a global classification branch to the lesion localization branch by a dual attention alignment module. To further enhance global information learning, we impose intra-class compactness and inter-class separability with a global prototype alignment module. For unsupervised data learning, we extend the focal loss to be its soft form to distill knowledge from a teacher model. Extensive experiments show the proposed OXnet outperforms competitive methods with significant margins. Further, we investigate omni-supervision under various annotation granularities and corroborate OXnet is a promising choice to mitigate the plight of annotation shortage for medical image diagnosis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge