Gregory Y. H. Lip
Leveraging Persistence Image to Enhance Robustness and Performance in Curvilinear Structure Segmentation
Jan 25, 2026Abstract:Segmenting curvilinear structures in medical images is essential for analyzing morphological patterns in clinical applications. Integrating topological properties, such as connectivity, improves segmentation accuracy and consistency. However, extracting and embedding such properties - especially from Persistence Diagrams (PD) - is challenging due to their non-differentiability and computational cost. Existing approaches mostly encode topology through handcrafted loss functions, which generalize poorly across tasks. In this paper, we propose PIs-Regressor, a simple yet effective module that learns persistence image (PI) - finite, differentiable representations of topological features - directly from data. Together with Topology SegNet, which fuses these features in both downsampling and upsampling stages, our framework integrates topology into the network architecture itself rather than auxiliary losses. Unlike existing methods that depend heavily on handcrafted loss functions, our approach directly incorporates topological information into the network structure, leading to more robust segmentation. Our design is flexible and can be seamlessly combined with other topology-based methods to further enhance segmentation performance. Experimental results show that integrating topological features enhances model robustness, effectively handling challenges like overexposure and blurring in medical imaging. Our approach on three curvilinear benchmarks demonstrate state-of-the-art performance in both pixel-level accuracy and topological fidelity.
StealthMark: Harmless and Stealthy Ownership Verification for Medical Segmentation via Uncertainty-Guided Backdoors
Jan 23, 2026Abstract:Annotating medical data for training AI models is often costly and limited due to the shortage of specialists with relevant clinical expertise. This challenge is further compounded by privacy and ethical concerns associated with sensitive patient information. As a result, well-trained medical segmentation models on private datasets constitute valuable intellectual property requiring robust protection mechanisms. Existing model protection techniques primarily focus on classification and generative tasks, while segmentation models-crucial to medical image analysis-remain largely underexplored. In this paper, we propose a novel, stealthy, and harmless method, StealthMark, for verifying the ownership of medical segmentation models under black-box conditions. Our approach subtly modulates model uncertainty without altering the final segmentation outputs, thereby preserving the model's performance. To enable ownership verification, we incorporate model-agnostic explanation methods, e.g. LIME, to extract feature attributions from the model outputs. Under specific triggering conditions, these explanations reveal a distinct and verifiable watermark. We further design the watermark as a QR code to facilitate robust and recognizable ownership claims. We conducted extensive experiments across four medical imaging datasets and five mainstream segmentation models. The results demonstrate the effectiveness, stealthiness, and harmlessness of our method on the original model's segmentation performance. For example, when applied to the SAM model, StealthMark consistently achieved ASR above 95% across various datasets while maintaining less than a 1% drop in Dice and AUC scores, significantly outperforming backdoor-based watermarking methods and highlighting its strong potential for practical deployment. Our implementation code is made available at: https://github.com/Qinkaiyu/StealthMark.
GROK: From Quantitative Biomarkers to Qualitative Diagnosis via a Grounded MLLM with Knowledge-Guided Instruction
Oct 05, 2025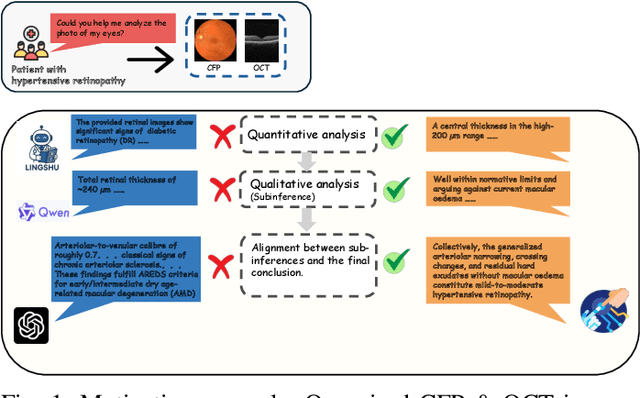
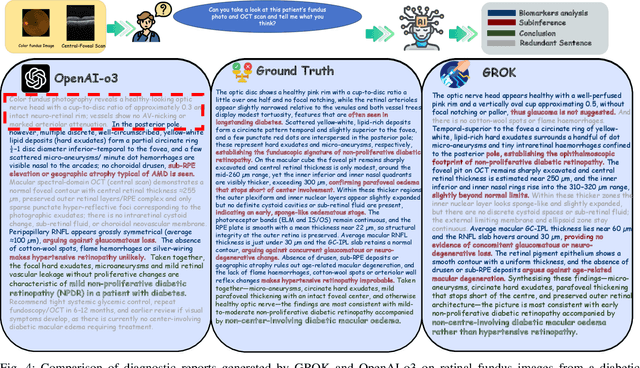
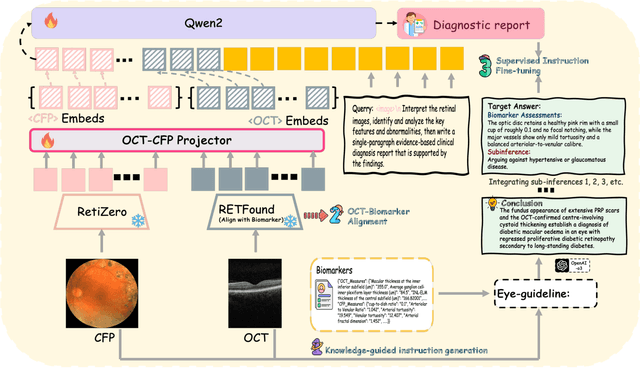
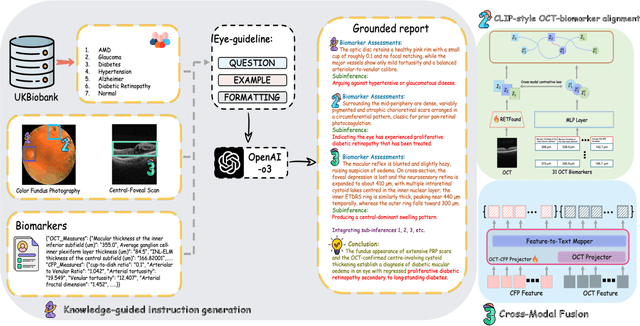

Abstract:Multimodal large language models (MLLMs) hold promise for integrating diverse data modalities, but current medical adaptations such as LLaVA-Med often fail to fully exploit the synergy between color fundus photography (CFP) and optical coherence tomography (OCT), and offer limited interpretability of quantitative biomarkers. We introduce GROK, a grounded multimodal large language model that jointly processes CFP, OCT, and text to deliver clinician-grade diagnoses of ocular and systemic disease. GROK comprises three core modules: Knowledge-Guided Instruction Generation, CLIP-Style OCT-Biomarker Alignment, and Supervised Instruction Fine-Tuning, which together establish a quantitative-to-qualitative diagnostic chain of thought, mirroring real clinical reasoning when producing detailed lesion annotations. To evaluate our approach, we introduce the Grounded Ophthalmic Understanding benchmark, which covers six disease categories and three tasks: macro-level diagnostic classification, report generation quality, and fine-grained clinical assessment of the generated chain of thought. Experiments show that, with only LoRA (Low-Rank Adaptation) fine-tuning of a 7B-parameter Qwen2 backbone, GROK outperforms comparable 7B and 32B baselines on both report quality and fine-grained clinical metrics, and even exceeds OpenAI o3. Code and data are publicly available in the GROK repository.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge