Uazman Alam
A WDLoRA-Based Multimodal Generative Framework for Clinically Guided Corneal Confocal Microscopy Image Synthesis in Diabetic Neuropathy
Feb 14, 2026Abstract:Corneal Confocal Microscopy (CCM) is a sensitive tool for assessing small-fiber damage in Diabetic Peripheral Neuropathy (DPN), yet the development of robust, automated deep learning-based diagnostic models is limited by scarce labelled data and fine-grained variability in corneal nerve morphology. Although Artificial Intelligence (AI)-driven foundation generative models excel at natural image synthesis, they often struggle in medical imaging due to limited domain-specific training, compromising the anatomical fidelity required for clinical analysis. To overcome these limitations, we propose a Weight-Decomposed Low-Rank Adaptation (WDLoRA)-based multimodal generative framework for clinically guided CCM image synthesis. WDLoRA is a parameter-efficient fine-tuning (PEFT) mechanism that decouples magnitude and directional weight updates, enabling foundation generative models to independently learn the orientation (nerve topology) and intensity (stromal contrast) required for medical realism. By jointly conditioning on nerve segmentation masks and disease-specific clinical prompts, the model synthesises anatomically coherent images across the DPN spectrum (Control, T1NoDPN, T1DPN). A comprehensive three-pillar evaluation demonstrates that the proposed framework achieves state-of-the-art visual fidelity (Fréchet Inception Distance (FID): 5.18) and structural integrity (Structural Similarity Index Measure (SSIM): 0.630), significantly outperforming GAN and standard diffusion baselines. Crucially, the synthetic images preserve gold-standard clinical biomarkers and are statistically equivalent to real patient data. When used to train automated diagnostic models, the synthetic dataset improves downstream diagnostic accuracy by 2.1% and segmentation performance by 2.2%, validating the framework's potential to alleviate data bottlenecks in medical AI.
GROK: From Quantitative Biomarkers to Qualitative Diagnosis via a Grounded MLLM with Knowledge-Guided Instruction
Oct 05, 2025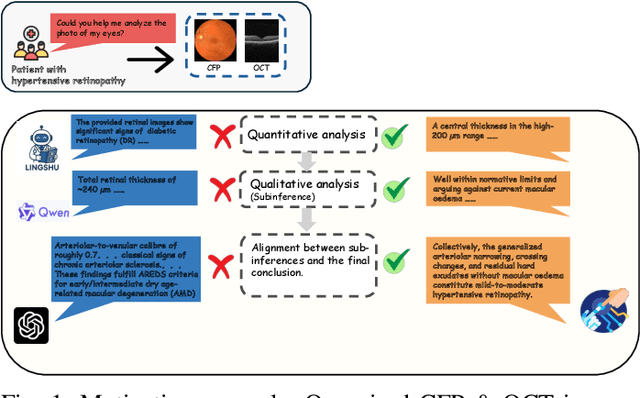
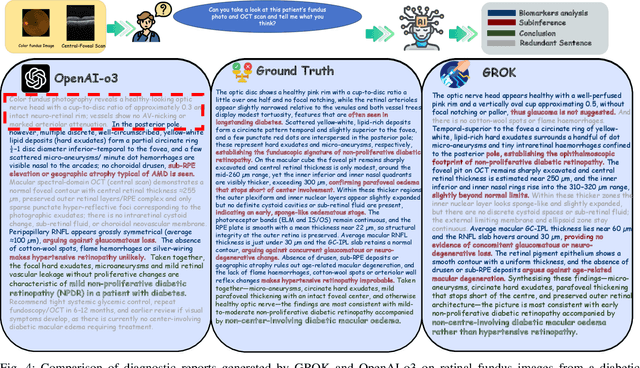
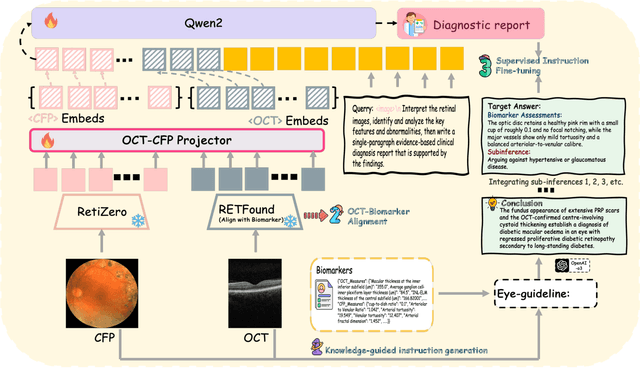
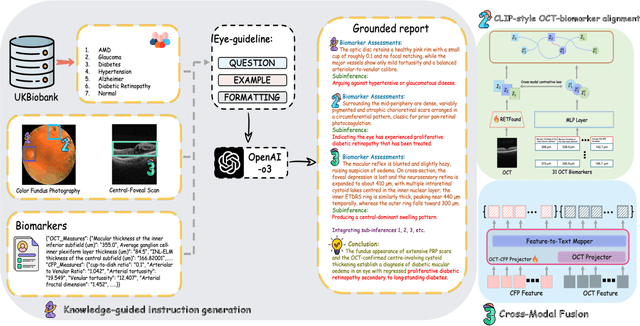

Abstract:Multimodal large language models (MLLMs) hold promise for integrating diverse data modalities, but current medical adaptations such as LLaVA-Med often fail to fully exploit the synergy between color fundus photography (CFP) and optical coherence tomography (OCT), and offer limited interpretability of quantitative biomarkers. We introduce GROK, a grounded multimodal large language model that jointly processes CFP, OCT, and text to deliver clinician-grade diagnoses of ocular and systemic disease. GROK comprises three core modules: Knowledge-Guided Instruction Generation, CLIP-Style OCT-Biomarker Alignment, and Supervised Instruction Fine-Tuning, which together establish a quantitative-to-qualitative diagnostic chain of thought, mirroring real clinical reasoning when producing detailed lesion annotations. To evaluate our approach, we introduce the Grounded Ophthalmic Understanding benchmark, which covers six disease categories and three tasks: macro-level diagnostic classification, report generation quality, and fine-grained clinical assessment of the generated chain of thought. Experiments show that, with only LoRA (Low-Rank Adaptation) fine-tuning of a 7B-parameter Qwen2 backbone, GROK outperforms comparable 7B and 32B baselines on both report quality and fine-grained clinical metrics, and even exceeds OpenAI o3. Code and data are publicly available in the GROK repository.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge