Gregory D. Hager
SANTIAGO: Spine Association for Neuron Topology Improvement and Graph Optimization
Aug 08, 2016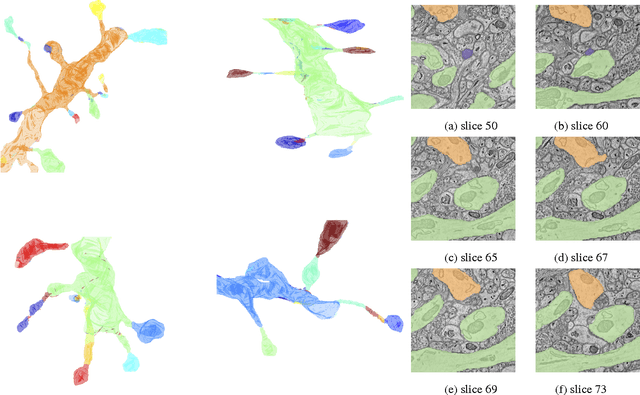
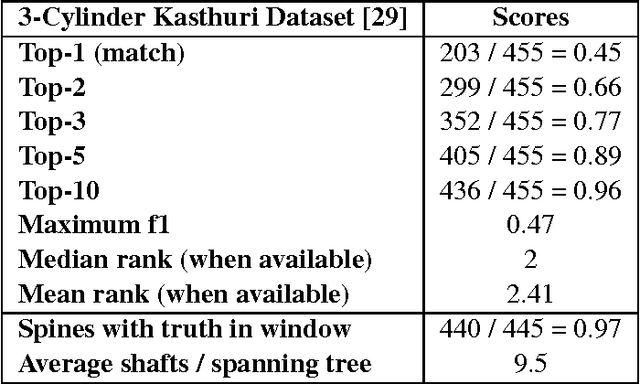
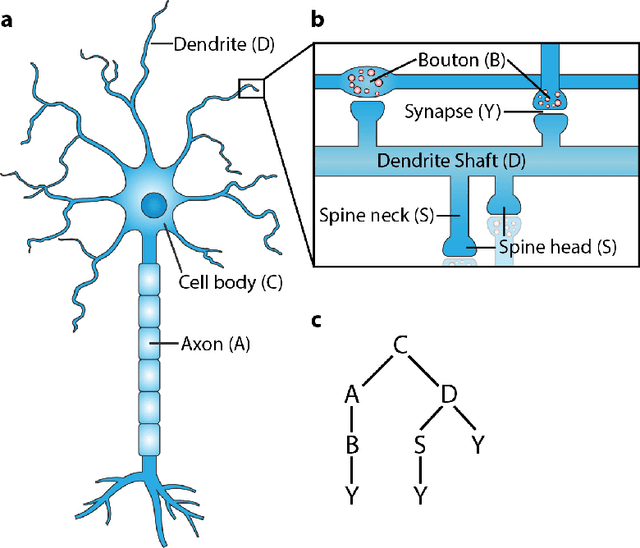
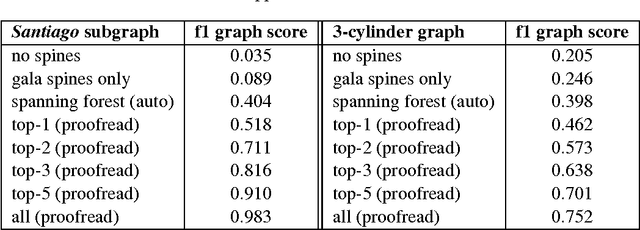
Abstract:Developing automated and semi-automated solutions for reconstructing wiring diagrams of the brain from electron micrographs is important for advancing the field of connectomics. While the ultimate goal is to generate a graph of neuron connectivity, most prior automated methods have focused on volume segmentation rather than explicit graph estimation. In these approaches, one of the key, commonly occurring error modes is dendritic shaft-spine fragmentation. We posit that directly addressing this problem of connection identification may provide critical insight into estimating more accurate brain graphs. To this end, we develop a network-centric approach motivated by biological priors image grammars. We build a computer vision pipeline to reconnect fragmented spines to their parent dendrites using both fully-automated and semi-automated approaches. Our experiments show we can learn valid connections despite uncertain segmentation paths. We curate the first known reference dataset for analyzing the performance of various spine-shaft algorithms and demonstrate promising results that recover many previously lost connections. Our automated approach improves the local subgraph score by more than four times and the full graph score by 60 percent. These data, results, and evaluation tools are all available to the broader scientific community. This reframing of the connectomics problem illustrates a semantic, biologically inspired solution to remedy a major problem with neuron tracking.
Recognizing Surgical Activities with Recurrent Neural Networks
Jun 22, 2016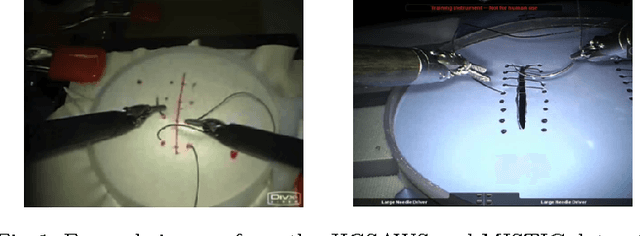
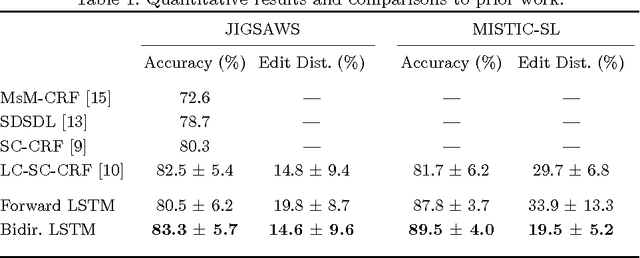

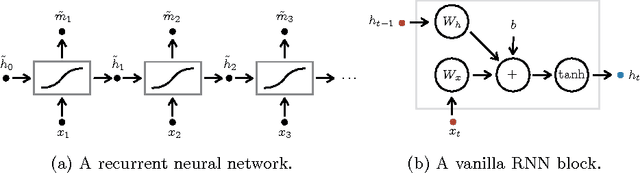
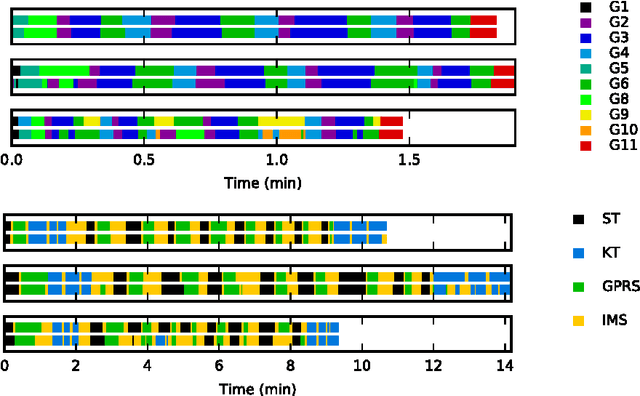
Abstract:We apply recurrent neural networks to the task of recognizing surgical activities from robot kinematics. Prior work in this area focuses on recognizing short, low-level activities, or gestures, and has been based on variants of hidden Markov models and conditional random fields. In contrast, we work on recognizing both gestures and longer, higher-level activites, or maneuvers, and we model the mapping from kinematics to gestures/maneuvers with recurrent neural networks. To our knowledge, we are the first to apply recurrent neural networks to this task. Using a single model and a single set of hyperparameters, we match state-of-the-art performance for gesture recognition and advance state-of-the-art performance for maneuver recognition, in terms of both accuracy and edit distance. Code is available at https://github.com/rdipietro/miccai-2016-surgical-activity-rec .
Towards Robot Task Planning From Probabilistic Models of Human Skills
Feb 15, 2016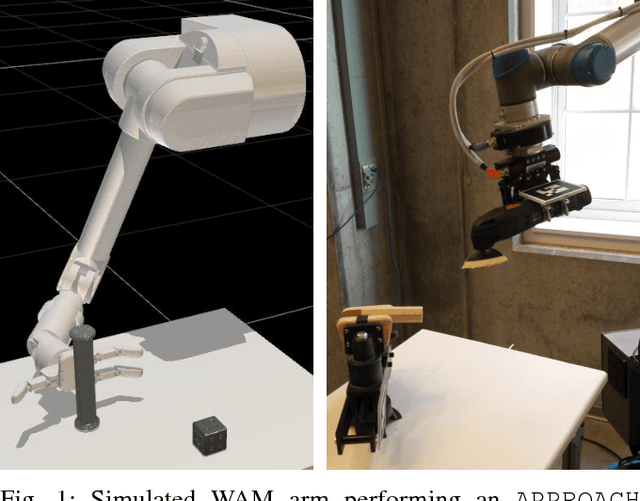
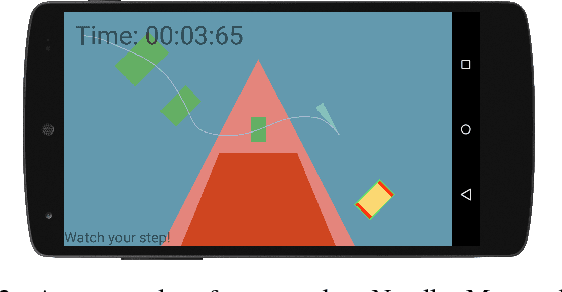
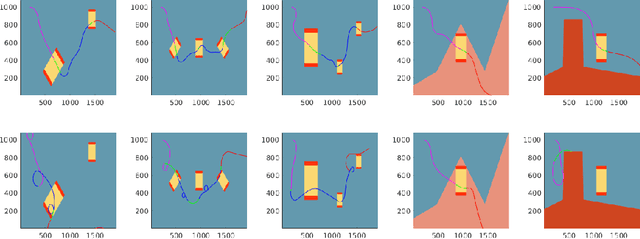
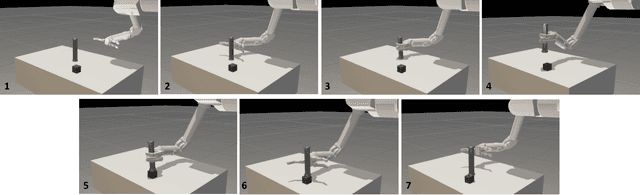
Abstract:We describe an algorithm for motion planning based on expert demonstrations of a skill. In order to teach robots to perform complex object manipulation tasks that can generalize robustly to new environments, we must (1) learn a representation of the effects of a task and (2) find an optimal trajectory that will reproduce these effects in a new environment. We represent robot skills in terms of a probability distribution over features learned from multiple expert demonstrations. When utilizing a skill in a new environment, we compute feature expectations over trajectory samples in order to stochastically optimize the likelihood of a trajectory in the new environment. The purpose of this method is to enable execution of complex tasks based on a library of probabilistic skill models. Motions can be combined to accomplish complex tasks in hybrid domains. Our approach is validated in a variety of case studies, including an Android game, simulated assembly task, and real robot experiment with a UR5.
VESICLE: Volumetric Evaluation of Synaptic Interfaces using Computer vision at Large Scale
Sep 07, 2015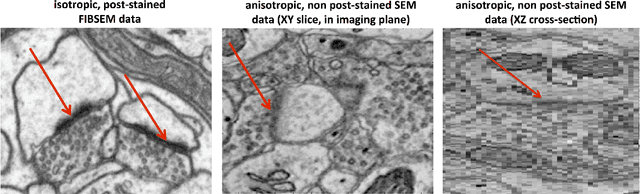
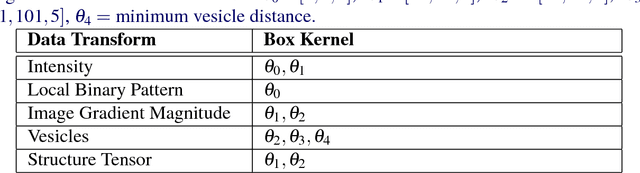

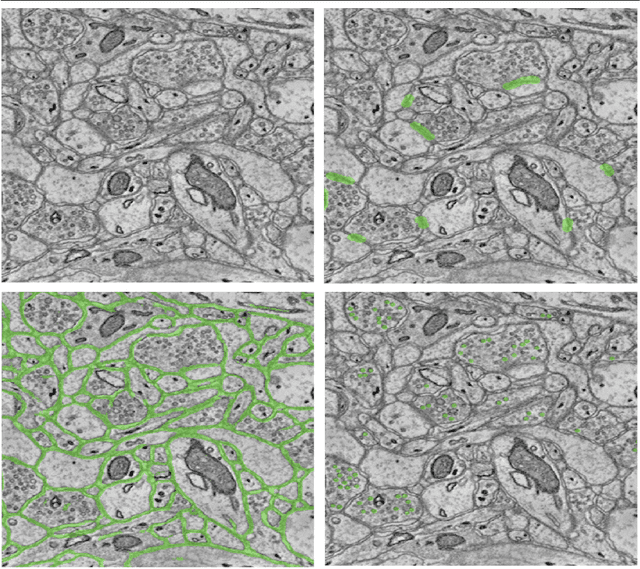
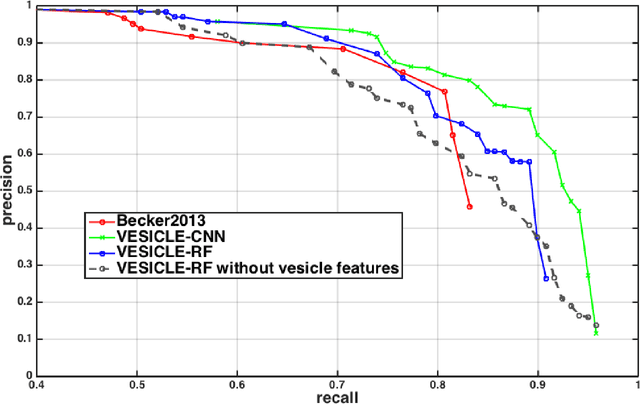
Abstract:An open challenge problem at the forefront of modern neuroscience is to obtain a comprehensive mapping of the neural pathways that underlie human brain function; an enhanced understanding of the wiring diagram of the brain promises to lead to new breakthroughs in diagnosing and treating neurological disorders. Inferring brain structure from image data, such as that obtained via electron microscopy (EM), entails solving the problem of identifying biological structures in large data volumes. Synapses, which are a key communication structure in the brain, are particularly difficult to detect due to their small size and limited contrast. Prior work in automated synapse detection has relied upon time-intensive biological preparations (post-staining, isotropic slice thicknesses) in order to simplify the problem. This paper presents VESICLE, the first known approach designed for mammalian synapse detection in anisotropic, non-post-stained data. Our methods explicitly leverage biological context, and the results exceed existing synapse detection methods in terms of accuracy and scalability. We provide two different approaches - one a deep learning classifier (VESICLE-CNN) and one a lightweight Random Forest approach (VESICLE-RF) to offer alternatives in the performance-scalability space. Addressing this synapse detection challenge enables the analysis of high-throughput imaging data soon expected to reach petabytes of data, and provide tools for more rapid estimation of brain-graphs. Finally, to facilitate community efforts, we developed tools for large-scale object detection, and demonstrated this framework to find $\approx$ 50,000 synapses in 60,000 $\mu m ^3$ (220 GB on disk) of electron microscopy data.
* v4: added clarifying figures and updates for readability. v3: fixed metadata. 11 pp v2: Added CNN classifier, significant changes to improve performance and generalization
An Automated Images-to-Graphs Framework for High Resolution Connectomics
Apr 30, 2015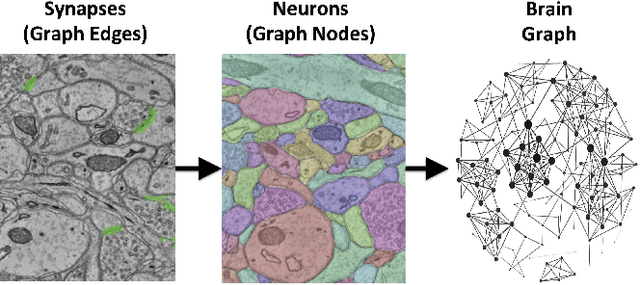
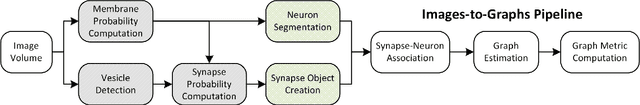
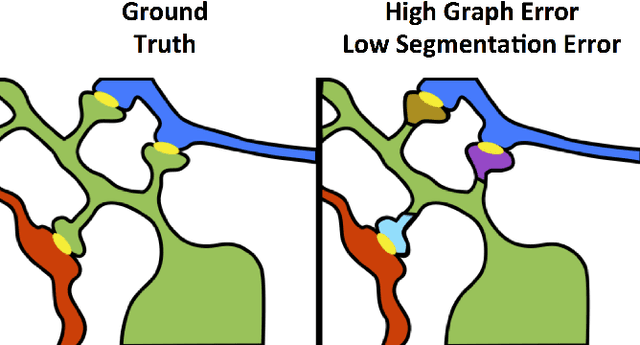
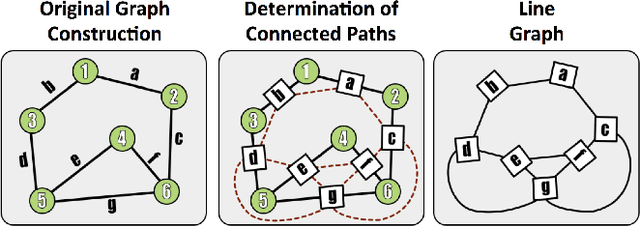
Abstract:Reconstructing a map of neuronal connectivity is a critical challenge in contemporary neuroscience. Recent advances in high-throughput serial section electron microscopy (EM) have produced massive 3D image volumes of nanoscale brain tissue for the first time. The resolution of EM allows for individual neurons and their synaptic connections to be directly observed. Recovering neuronal networks by manually tracing each neuronal process at this scale is unmanageable, and therefore researchers are developing automated image processing modules. Thus far, state-of-the-art algorithms focus only on the solution to a particular task (e.g., neuron segmentation or synapse identification). In this manuscript we present the first fully automated images-to-graphs pipeline (i.e., a pipeline that begins with an imaged volume of neural tissue and produces a brain graph without any human interaction). To evaluate overall performance and select the best parameters and methods, we also develop a metric to assess the quality of the output graphs. We evaluate a set of algorithms and parameters, searching possible operating points to identify the best available brain graph for our assessment metric. Finally, we deploy a reference end-to-end version of the pipeline on a large, publicly available data set. This provides a baseline result and framework for community analysis and future algorithm development and testing. All code and data derivatives have been made publicly available toward eventually unlocking new biofidelic computational primitives and understanding of neuropathologies.
Hierarchical Sparse and Collaborative Low-Rank Representation for Emotion Recognition
Apr 01, 2015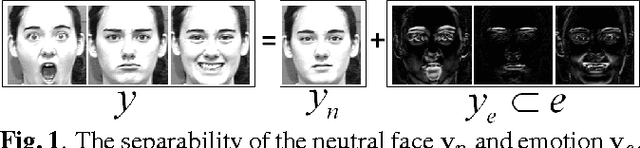
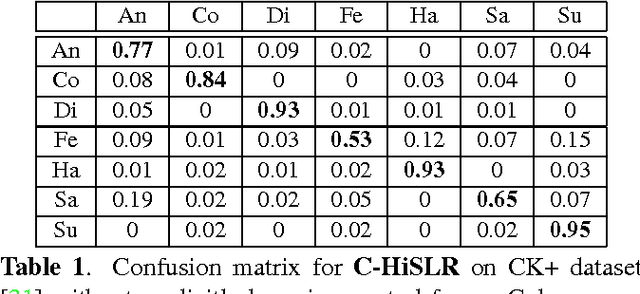

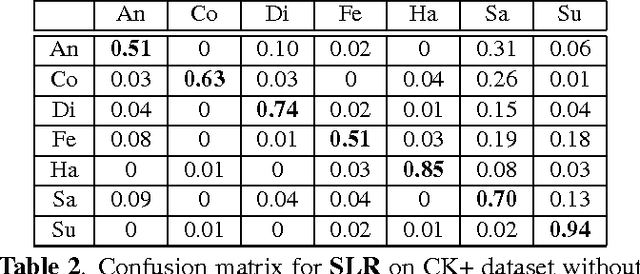
Abstract:In this paper, we design a Collaborative-Hierarchical Sparse and Low-Rank (C-HiSLR) model that is natural for recognizing human emotion in visual data. Previous attempts require explicit expression components, which are often unavailable and difficult to recover. Instead, our model exploits the lowrank property over expressive facial frames and rescue inexact sparse representations by incorporating group sparsity. For the CK+ dataset, C-HiSLR on raw expressive faces performs as competitive as the Sparse Representation based Classification (SRC) applied on manually prepared emotions. C-HiSLR performs even better than SRC in terms of true positive rate.
Automated Objective Surgical Skill Assessment in the Operating Room Using Unstructured Tool Motion
Dec 18, 2014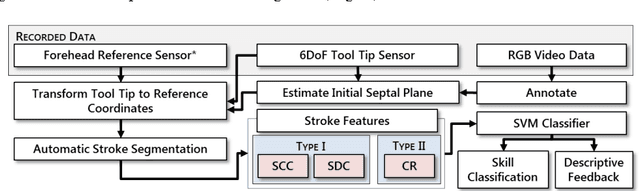
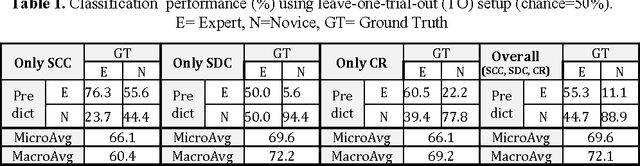
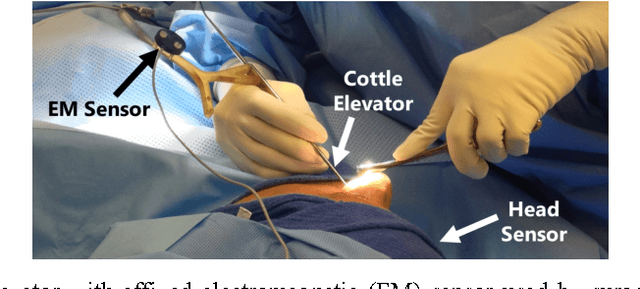
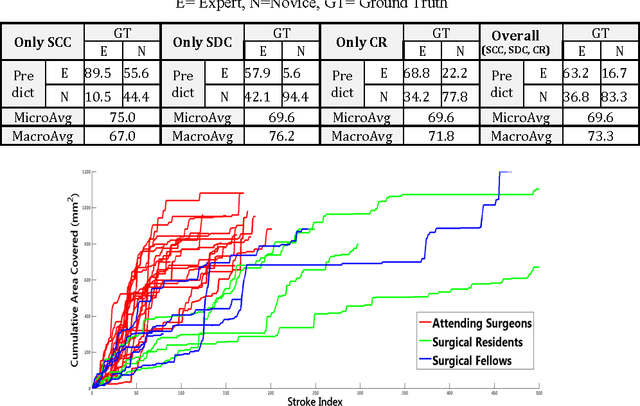
Abstract:Previous work on surgical skill assessment using intraoperative tool motion in the operating room (OR) has focused on highly-structured surgical tasks such as cholecystectomy. Further, these methods only considered generic motion metrics such as time and number of movements, which are of limited instructive value. In this paper, we developed and evaluated an automated approach to the surgical skill assessment of nasal septoplasty in the OR. The obstructed field of view and highly unstructured nature of septoplasty precludes trainees from efficiently learning the procedure. We propose a descriptive structure of septoplasty consisting of two types of activity: (1) brushing activity directed away from the septum plane characterizing the consistency of the surgeon's wrist motion and (2) activity along the septal plane characterizing the surgeon's coverage pattern. We derived features related to these two activity types that classify a surgeon's level of training with an average accuracy of about 72%. The features we developed provide surgeons with personalized, actionable feedback regarding their tool motion.
Multi-environment model estimation for motility analysis of Caenorhabditis Elegans
Jul 08, 2010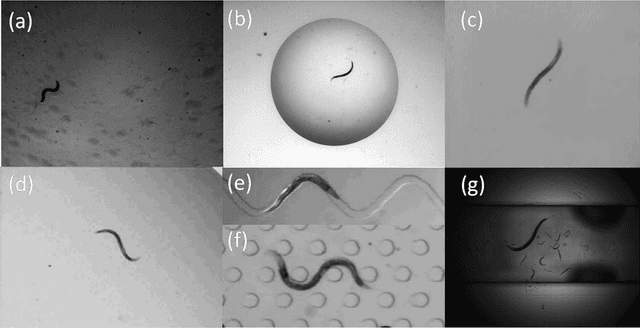
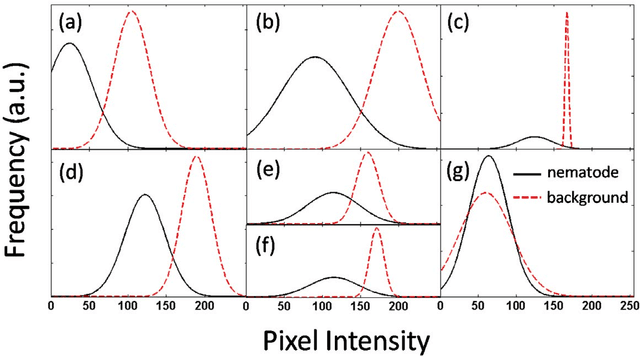
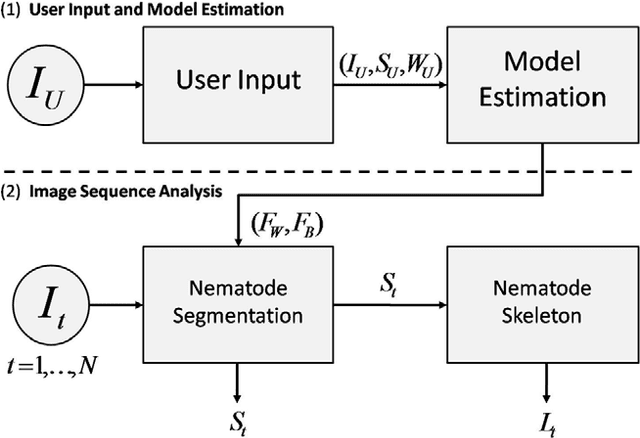
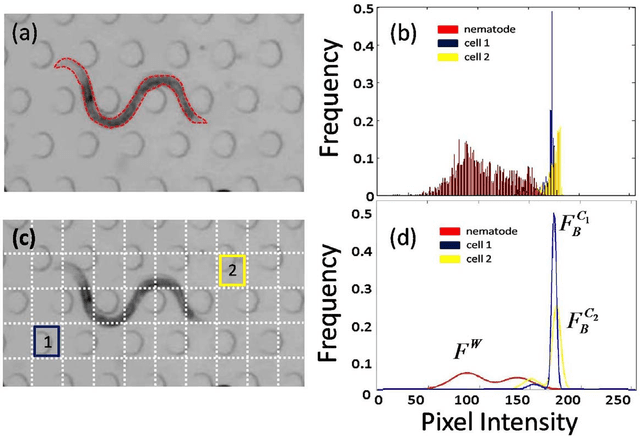
Abstract:The nematode Caenorhabditis elegans is a well-known model organism used to investigate fundamental questions in biology. Motility assays of this small roundworm are designed to study the relationships between genes and behavior. Commonly, motility analysis is used to classify nematode movements and characterize them quantitatively. Over the past years, C. elegans' motility has been studied across a wide range of environments, including crawling on substrates, swimming in fluids, and locomoting through microfluidic substrates. However, each environment often requires customized image processing tools relying on heuristic parameter tuning. In the present study, we propose a novel Multi-Environment Model Estimation (MEME) framework for automated image segmentation that is versatile across various environments. The MEME platform is constructed around the concept of Mixture of Gaussian (MOG) models, where statistical models for both the background environment and the nematode appearance are explicitly learned and used to accurately segment a target nematode. Our method is designed to simplify the burden often imposed on users; here, only a single image which includes a nematode in its environment must be provided for model learning. In addition, our platform enables the extraction of nematode `skeletons' for straightforward motility quantification. We test our algorithm on various locomotive environments and compare performances with an intensity-based thresholding method. Overall, MEME outperforms the threshold-based approach for the overwhelming majority of cases examined. Ultimately, MEME provides researchers with an attractive platform for C. elegans' segmentation and `skeletonizing' across a wide range of motility assays.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge